Abstract
Rising temperatures and associated heat stress threaten soybean [Glycine max (L.) Merr.] productivity. With few mitigation options, improving genetic tolerance is key. However, the genetics of heat tolerance in soybean remain understudied. We evaluated 450 accessions (maturity groups 0–IV) under optimal and heat stress conditions to characterize physiological and growth traits, explore trait relationships, and assess genomic prediction. We identified 37 significant marker–trait associations (MTAs): 20 unique MTAs in optimal conditions, 16 under heat stress, and one MTA for a heat tolerance index. Only one MTA was consistent across temperatures, indicating genetic divergence in responses. Genomic prediction had moderate predictive ability for biomass traits but less accurate for physiological traits related to photosynthetic parameters. Several heat-tolerant accessions were identified for use in breeding. These results provide insights into the genetic architecture of heat tolerance, supporting marker-assisted and genomic selection to develop heat-tolerant soybean varieties.
Similar content being viewed by others
Introduction
Soybean [Glycine max L. (Merr.)] is a major oilseed crop1 belonging to the Leguminosae family, with origins in the temperate regions of China ~5000 years ago2. Soybean production has rapidly expanded globally and spans a wide latitude distribution3, demonstrating its adaptability to different environments. Soybean, similar to other row crops, is adversely affected by abiotic stresses, which, individually or in combination, cause changes that negatively affect crop growth and productivity. Soybean is a warm-season legume with a C3 system of photosynthesis4 and is considered a relatively heat-tolerant crop in comparison to cool-season legume crops such as field pea (Pisum sativum L.). However, heat, drought, cold, and salinity are the primary climatic abiotic factors that cause severe crop damage and associated yield losses5.
Climate change, including elevated CO2 levels, will impact soybean production, though any yield increases from CO2 fertilization are negated by drought and heat stresses6,7,8,9. The long-term forecast of changing climatic conditions, including an increase in temperatures, threatens all crops, including legume crops10,11,12,13,14. Plants have an inherent and evolved mechanism to respond to abiotic stresses, which are considered a complex trait since they involve numerous metabolic pathways, cellular components, and molecular components15. Plant response to abiotic stresses involves mechanisms that are interconnected, leading to cellular damage and secondary stresses, including osmotic and oxidative stresses. The inheritance of abiotic stress tolerance is multi-genic, generally with small individual effects controlling the trait16.
The effect of heat stress on crop morphology, physiology, and reproduction has been well-studied in various crop species17,18,19, although the responses differ across species and genotypes. Heat imposed in vegetative tissue has been found to negatively affect vital physiological processes, including chlorophyll content, photosynthesis, cellular respiration, and stomatal conductance20,21,22. Additionally, heat stress has been found to increase Reactive Oxygen Species (ROS) activity, which affects the cellular structure and the production of pigments and can deteriorate the thylakoid membranes23. These effects also occur in the plant during reproductive stages, which can be more at risk to heat due to the sensitivity of gametogenesis17, leading to crop yield losses due to reduced success of fertilization. Genetic variation of these traits has been used to distinguish sensitive and tolerant genotypes in various crops24,25, which can be utilized to breed for improved crop performance under stressful environments. Genetic studies aimed at exploring molecular mechanisms of heat stress response complement breeding efforts. These studies establish the mechanisms of heat stress tolerance and crop response; however, limited information is available for soybean, and most of the heat-related studies come from other crop species, particularly in cereals. Given the complexity of tolerance to abiotic stresses such as heat15, breeding for heat tolerance is one of the most economical approaches for protecting soybean yields. Due to the paucity of data on heat stress response in soybean, large accession panels are required to sufficiently sample the genetic variation and study the genetics of heat tolerance.
In other legumes, marker trait associations (MTAs) and candidate genes have been reported for field peas26, common bean (Phaseolus vulgaris L.)27 peanuts (Arachis hypogaea L.)28, faba beans (Vicia faba L.)29, and chickpeas (Cicer arietinum L.)30 grown under heat stress. Sequencing studies of soybeans grown in heat conditions have shown an overexpression of heat shock proteins and transcription factors31,32,33, as well as differentially expressed genes related to metabolic processes and molecular transports34. Heat stress has been found to induce overexpression of the DREB1 gene family, LEA genes, and dehydrins, which were also overexpressed for other abiotic stresses35.
The response of soybean to heat stress is less studied than heat stress in cool-season legume species, although soybean yield losses due to heat have been reported in numerous studies9,36,37,38,39,40. Additionally, simulations of soybean yield at end-of-century conditions have found yield losses of up to 22% due to the effects of high temperatures10, or a decline of 3.3% for each decade, with temperature rise being one of the main factors for the decline14. However, information on the impact of heat stress on early-season soybean growth is limited41, and the genetic architecture that may be underlying soybean tolerance to heat stress is poorly understood. To date, no study has reported on GWAS-detected markers associated with physiological traits of soybeans grown under heat stress conditions. Soybean has a wide genetic variation for multiple traits, as demonstrated in studies for yield and abiotic stress tolerance that identified candidate genes via GWAS for yield, seed composition42, drought43,44, flooding45, and iron deficiency chlorosis46,47.
Soybean growth and development stages include vegetative and reproductive phases, and previous research has identified that crop losses start when the average daily air temperature exceeds ~ 28 to 30°C during the vegetative stage48 and 22 to 24 °C during the reproductive stage6. However, these optimum temperatures for healthy plant development were proposed before the more dramatic shift in climate patterns and increasing temperatures. Such temperature increases invariably coincide with water-limited conditions, i.e., drought. Drought has long been known as a major limiting factor in soybean productivity and is expected to remain a major limiting factor. However, it is predicted that a 2 °C rise in global temperatures may cause heat to become the foremost cause of yield losses in soybean and maize in the US Corn Belt10. As temperatures rise and water availability to plants at critical times becomes a dual threat of exceeding proportions, global soybean production will be severely impacted.
Although difficult and often uneconomical or unsustainable, irrigation can be provided to counter the water deficit. However, against heat, improved genetic materials currently remain the best solution. For example, one tool to counteract the adverse effects of future climate change is to plant varieties with a genetic tolerance to various abiotic stresses, such as heat. A difficulty of developing soybean genotypes with superior tolerance to abiotic stress, such as heat, is that such tolerance genes are generally quantitative and are controlled by many small-effect QTL. Plant breeders can use genomic tools, including marker-assisted selection (MAS), genomic selection (GS), and genetic engineering, to breed for soybean heat stress tolerance more rapidly. With genomic prediction, breeders can evaluate the genetic potential of a large number of genotypes without needing to grow and evaluate each genotype, which saves time, resources, and labor49.
The objectives of this study were: (a) screen a diverse accession panel to examine the early season heat tolerance response in soybean, (b) understand the impact of heat on physiological traits with potential for high throughput phenotyping (HTP) applications, (c) conduct genome-wide association studies to identify genomic regions crucial for heat stress tolerance and identify candidate genes for heat stress tolerance, and (d) investigate the usefulness of genomic prediction of different heat stress related physiological traits, and project the GP model to screening the USDA core collection across all maturity groups.
Results
Descriptive statistics
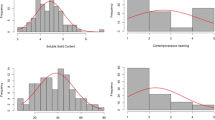
The distributions and descriptive statistics for all measured traits are summarized in Fig. 1 and Table 1. Phenotypic variation was observed for all traits across a single year, as well as temperature treatment. The effect of genotype was significant for all traits except canopy temperature (p < 0.05). The treatment effect was significant for SPAD, canopy temperature, fresh shoot biomass (FSB), dry shoot biomass (DRB), and stomatal conductance in 2022. The genotype by treatment effect was significant for all traits in 2022 except ΦPSII and canopy temperature (p < 0.05). The interaction effect of genotype and treatment was also significant for DSB and DRB in 2021. Comprehensive ANOVA results are included in Supplementary Table 1. The heritability estimate was higher in the optimal temperature treatments for most traits in both years (Table 1). The heritability estimates for fresh root and shoot biomass traits were similar across the temperature treatments. The genetic correlations between traits in the optimal and heat conditions were all significant (p < 0.01) and ranged from 0.5 for stomatal conductance to 0.99 for ΦPSII.
Data comes from 450 soybean accessions from the USDA-GRIN mini core collection. The red is the distribution in heat conditions, and the blue is the distribution in the optimal control conditions. Purple is the overlap of the two distributions. Panels A−D represent trait distributions for 2021 with A) SPAD, B) canopy temperature, C) dry shoot biomass, and D) dry root biomass. Panels E−L represent traits measured in 2022, including E) SPAD, F) canopy temperature, G) stomatal conductance, H) quantum efficiency in light, I) fresh shoot biomass, J) dry shoot biomass, K) fresh root biomass, and L) dry root biomass.
The chlorophyll content was not different between the heat and optimal conditions in 2021. However, in 2022, chlorophyll content significantly differed between the two temperature conditions, with a lower SPAD value in the heat than in the optimal. Canopy temperature was significantly higher in the heat treatment in both years, with a greater difference observed in 2022 (Table 1). The FSB, DSB, and DRB had significantly higher biomass in the heat treatment in 2022, while FRB was not significantly different between the heat and control. The dry biomass traits responded differently in 2021, with significantly lower biomass values in the heat treatment. Both photosynthetic traits, stomatal conductance (gsw) and ΦPSII, had significantly lower values in heat compared to the optimal treatment (p < 0.001). For many of the traits, the distribution was wider in the heat treatment as compared to the optimal treatment, indicating a greater diversity of genotypic responses to the higher temperature.
Due to a more comprehensive trait data availability in 2022, we focused on these data to identify five top heat-tolerant soybean accessions. The five heat-tolerant soybean accessions from the USDA-GRIN collection are listed in Table 2. These lines were selected based on the HtI values, with a lower HtI being better for all traits, excluding canopy temperature. A lower HtI indicates the cultivar’s phenotype was similar or better in the heat treatment as compared to the optimal treatment. The heat-tolerant lines are representative of two maturity groups tested in the panel, with the majority from MG II. These lines had phenotypes similar in both temperature treatments, or better in the heat, for some traits. All of the selected accessions had no significant changes in chlorophyll content, and the majority of the accessions had a significant increase in all biomass traits. The photosynthetic parameter traits had no change between the heat and optimal conditions, except for a higher stomatal conductance rate for PI 437840A under the heat treatment.
For comparison purposes, the difference in population means for the HT and OT treatments is provided to illustrate the trait value differences in these five tolerant accessions relative to the overall 423 accessions. We also investigated five heat-susceptible accessions and noted they generally had lower trait values when grown under heat conditions (Table 2). The chlorophyll content of these accessions was significantly lower in the heat treatment. These susceptible accessions had lower biomass, and lower photosynthetic parameters with a significant decrease in stomatal conductance for all five susceptible accessions when under heat stress (Table 2).
Genome-Wide Association Studies
Association studies detected 37 unique MTAs across different traits and in different temperature treatments (Table 3, Supplementary Table 2). No significant MTAs were detected in 2021; all further reported GWAS results are from 2022, and a comparison of GWAS results between SVEN and other GWAS models can be found in Supplementary Table 3. The lack of MTAs in 2021 could be due to lower heritability and greater environmental effects that were found in the data for that year. Table 3 summarizes the distribution of significant MTAs that were identified across traits. Four MTAs (ss715583256, ss715611949, ss715625497, and ss715627447) had a MAF less than 5% and are located on different chromosomes. The biomass traits had the most significant number of associated MTAs. More MTAs were associated with trait response in the optimal control temperature for most traits. Across all traits, none of the MTAs detected in the heat treatment were detected in the optimal treatment for the same trait. Five of the significant MTAs were associated with more than one trait, with four detected for DRB. A single SNP, ss715607454, was significant for DSB HtI, and was the only MTA detected for a heat tolerance index trait. This SNP is 3.5 kbp away from Glyma.10G219100 and 9.1 kbp away from Glyma.10G219200. Both genes are described as encoding for laccase proteins, which are glycoproteins involved in the production of lignin for plant growth and have been found to be involved with abiotic stress response in several species50. In addition to these nearby candidate genes, the gene Gylma.10G219600 is 37.4 kbp downstream and encodes for an NAC transcription factor, GmNAC074, a class of proteins previously reported to be involved in response to abiotic stress in soybean51,52. Further upstream, at 135.8 kbp away, Glyma.10G217900 has been reported to be upregulated in soybeans grown in heat-stress conditions34.
Table 4 lists a subset of potential candidate genes from the MTAs we detected. The SNP ss715588916 was detected in the heat treatment and associated with FSB and DRB. This SNP is 61.9 kbp away from Glyma.04G242300, which has been previously reported to be downregulated in soybean due to heat34 and encodes for a plantacyanin protein. The SNP ss715604755 was detected in the optimal conditions for the two fresh biomass traits. This SNP is within Glyma.09G249100, a glutamine-fructose-6-phopshate transaminase (GFAT). In optimal conditions, two SNPs, ss715613671 and ss715612610, were associated with FSB and DRB. The latter SNP is 5.4 kbp away from the candidate gene Glyma.12G197600, a xyloglucan endotransglucosylase/hydrolase (XTH) protein. A single MTA was detected across temperature treatments, although in different traits. This SNP, ss715627447, was significant for FSB in the heat treatment and for DRB in optimal. It is 49.8 kbp upstream from Glyma.17G228800, which has been reported to be upregulated in soybean exposed to heat stress (Wang et al., 2018a) and is a glycine decarboxylase P-protein.
The candidate gene Glyma.01G148800 is 9.9 kbp away from ss715579731, a SNP reported in association with FSB in heat. This gene encodes for an uncharacterized protein and has been previously found to be downregulated in soybeans exposed to heat stress34. This SNP is also within the candidate gene Glyma.01G148900, which is a Brevis radix-like (BRX) protein. BRX proteins have been found to promote biomass development in Arabidopsis53. Another potential candidate gene on Chromosome 2 is Glyma.02G238300, a mitochondrial FE/S cluster exporter protein previously reported as significant for total chlorophyll content54. This gene is 157.4 kbp away from ss715583256, which was associated with SPAD in heat. The SNP is also 25.6 kbp away from an additional potential candidate gene, Glyma.02G275800, a trigalactosyldiacylglycerol2 (TGD2) protein. The gene Glyma.06G194900 is 17.2 kbp away from a SNP, ss715593740, associated with stomatal conductance in the optimal treatment. This candidate gene is a light-harvesting chlorophyll-protein complex. Another SNP associated with stomatal conductance, ss715616040, is downstream of Glyma.13G285300, which is a cytochrome P450 (CYP450) family protein. The SNP ss715601977, which is associated with ΦPSII in heat, is 40.9 kbp away from the candidate gene Glyma.08G306200, which is a phytoene synthase protein. The SNP ss715613794 was associated with ΦPSII in heat and is 42.3 kbp away from the candidate gene Glyma.13G090600, which encodes for a glycine-rich protein (GRP) and has been found to be upregulated in heat34.
Genomic prediction
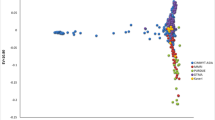
Population structure analyses indicated no clear structure due to maturity groups. The PCA showed no clear clustering, and genotypes from different MGs were dispersed amongst each other (Supplementary Fig. 3). The DA showed that groups from different MGs greatly overlapped with each other, and the prediction accuracy from the cross-validation was low (0.29). This indicates that any population structure in the panel is not likely due to MG. The SNMF analysis indicated that the lowest cross-entropy was a range of 35–40 ancestral clusters, which does not correspond with the five MGs that are in the panel. These analyses indicate a lack of clear population structure in the diversity panel, lending evidence that our results are not an artifact of the maturity group.
Genomic prediction ability (GPA) was calculated against the testing and unseen validation sets. Training was performed in 2021, 2022, and combined year data when available. Canopy temperature was dropped from genomic prediction analysis due to the non-significant genotypic effect. A summary of the GPAs is shown in Table 5. The GPA for most traits under heat conditions decreased when tested against the unseen validation set compared to the testing set. In three cases (2021 DSB, 2021 DRB, 2022 ΦPSII), the GPA against the validation set was higher than against the testing set. In traits with two years of data, there was a trend of higher GPA in the 2022 data, which follows a trend of higher heritability in the 2022 data. For SPAD, the highest GPA came from combining data from the two years, while this combined data resulted in a lower GPA for DSB and DRB. The same trend of higher GPA for the test set was true when trained on the traits grown in the optimal control. Only the 2022 SPAD and the 2022 ΦPSII had higher GPAs when tested against the validation set. For all traits except 2022 ΦPSII, the GPA for the test set was higher when trained and tested on data collected from plants grown in the control compared to the heat, although this increase was non-significant in most cases. The GPA for the validation set was higher in the control than heat, except in 2022 FSB, 2021 DSB, and 2022 ΦPSII. This trend of higher GPA in the control follows the higher heritabilities found in the control set. The GPAs for the trait indices were lower than those in a single temperature treatment and showed a poor ability to predict the stress tolerance index.
The number of marker-trait associated SNPs used for different trait and year combinations varied between 933 and 2289. The mean accuracy of the predictions based on trait-associated SNPs obtained through GWAS was lower than the mean accuracy when using the entire 50K SNP panel (Table 5). SPAD in the Heat 2021 environment was the only trait with a higher prediction accuracy for both the testing and validation sets when the trait-associated SNPs were used for training. Interestingly, when the SNPs detected in the control conditions were used to train to predict traits grown in heat, the prediction accuracy was not different from that of the heat-selected SNPs.
Based on data availability across years and GPAs, three traits were selected for use in germplasm projection: SPAD, DSB, and DRB. The average trait performance of the top 5% and 10% accessions in the USDA collection showed a higher average across the accessions evaluated in this study (Fig. 2), indicating that it is possible to find other heat-tolerant accessions in the USDA germplasm collection. The top 5% and top 10% of heat-tolerant accessions in the panel we screened overlapped with the top 5% accessions when projected to the entirety of the USDA collection. For the traits, there was an overlap of 100%, 80%, and 90% for SPAD, DSB, and DRB, respectively (Fig. 2). The country of origin of the top 5% heat tolerant accession from the germplasm collection varied, with China, United States, Japan, South Korea, and Russia as the five countries contributing the most (Supplementary Fig. 2). The MG of the top 5% heat tolerant accessions was mainly in the early maturity groups 0-II (Supplementary Fig. 2).
GEBVs for the 423 accessions included in this study and the projected GEBVs of selected varieties in the USDA germplasm collection. The y-axis represents the distribution of the predicted GEBVs and the x-axis represents the selected group displayed in each distribution. These groups include the 423 accessions screened in this study (Panel Mean), 10% highest for the 423 accessions evaluated (Panel Top 10%), 5% highest for the 423 accessions evaluated (Panel Top 5%), 10% highest for the entire USDA germplasm collection (USDA Top 10%), 5% highest for the entire USDA germplasm collection (USDA Top 5%) for SPAD, DSB, and DRB for two years (2021 and 2022). Distributions are shown for SPAD, dry shoot biomass, and dry root biomass in 2021 and 2022.
Discussion
Our study leveraged a soybean diversity panel of 450 accessions from the USDA-GRIN soybean mini core collection to obtain detailed trait data in response to heat stress. In this first step, we focused on the early growth stages stress response to aid in the identification of useful access for breeding applications, generate genetic insights, and enhance phenomics-assisted selection15. Our study revealed that heat stress adversely affected physiological traits such as chlorophyll content (SPAD), stomatal conductance, and Quantum Efficiency of PSII, similar to what was reported with previous studies22. Additionally, canopy temperatures increased under heat stress, consistent with previous reports22. The change in biomass to heat was inconsistent over two years; 2021 saw reduced biomass under heat stress, while 2022 exhibited increased biomass. The increase in biomass under heat in 2022 could possibly be due to accelerated growth stages and internode elongation, which has been previously reported in tomato55, although the varying biomass responses across years indicate the need for further study and for extended periods of stress into later growth stages. Further, we attribute the difference in biomass across years to temperature differences between 2021 and 2022. In 2021, the optimal temperature greenhouse had cooling issues due to a prolonged period of hot and humid weather in Ames. This caused temperatures in the greenhouse to rise above 30 °C. These elevated temperatures persisted for ten days and resulted in more rapid growth of the previously healthy plants grown in the optimal temperatures. In contrast, for 2022, we moved to greenhouses with improved climate control, and the control plants never experienced elevated temperatures. From 2021 to 2022, we had an improved temperature differential, with an average difference of 4.3 degrees in 2021 and 8.7 degrees in 2022. The differences we noted between years indicate the need for multiple years of testing, even when in a semi-controlled environment such as a greenhouse. We noted seven accessions had initiated flowering (R1 growth stage) within three weeks of planting, which is dramatically different from the normal expectation of planting to R1 duration of about 6-8 weeks. Generally, heritability was lower under heat stress, a trend also observed in other crops under marginal conditions56,57,58. The heritability of the biomass traits saw the least drop when comparing across the two temperature treatments, although the dry biomass traits saw a greater decline in 2022 than in 2021. The photosynthetic parameters saw the greatest drop in heritabilities when measured under heat stress.
Understanding the genetic architecture of physiological traits evaluated under heat stress helps identify markers and develop genomic selection pipelines to assist in the development of heat-tolerant soybean. To date, very limited studies have reported on GWAS in soybean tolerance to heat stress. To explore heat stress’s complex genetic architecture using GWAS involving 423 diverse soybean accessions, we measured eight traits related to growth and physiological parameters at an optimal control temperature and in a heat stress temperature. We identified 37 unique MTAs that were associated with different traits and distributed across 17 of the 20 soybean chromosomes. MTAs associated with traits in the optimal treatment made up 20 of the detected MTAs, with 15 of the additional MTAs detected in heat, one MTA detected across temperature treatments, and the remaining one MTA detected for an HtI. This lack of overlap indicates a different genetic control in soybean growth and development under optimal conditions and in stress conditions, which has been previously reported in corn58,59.
In the current study, several putative candidate genes were identified for the physiological traits in heat stress and without stress. The SNP detected for the HtI DSB, ss715607454, is of particular interest due to the four potential candidate genes in the region. These genes include two laccase proteins, a NAC transcription factor, and a sucrose-protein symporter. All of these proteins have been found to be involved in the response to abiotic stress in soybean and other plant species50,51,52,60. The SNP associated with ΦPSII in heat, ss715601977 is near a phytoene synthase protein gene. Phytoene synthase is essential for modulating the biosynthesis of the carotenoid pigment61, which are a light-harvesting pigment used in the photosystem complexes and are also essential for protection against reactive oxygen species produced in the chloroplast due to stress62.
The biomass traits have several candidate genes of particular interest. The SNP ss715579731 is within a gene that encodes for a BRX protein, which is a class of proteins that have been found to promote shoot and root growth in Arabidopsis53 and rice63. Two SNPs associated with DSB in heat have candidate genes of interest. The first SNP, ss715591790, is near the gene Glyma.05G233500, which encodes for an unknown protein that was downregulated in soybeans exposed to heat stress34. The second SNP, ss715610072, is near the candidate gene Glyma.11G206700. This gene is a cysteine-rich receptor-like kinase (CRK), which is a family of receptor-like kinases (RLKS) that are important in plant immunity, response to abiotic stress, and regulating growth and development64. CRKs are highly evolutionarily conserved across vascular plants and have been found to modulate the tradeoff between growth and stress response in many species64. The SNP on chromosome 13 associated with DRB in heat has two promising candidate genes nearby, Glyma.13G223000 and Glyma.13G223100. These two genes are the GmGELP103 and GmGELP104 genes65, which are GDSL-type esterase/lipase proteins. The GDSL gene family has been found to be involved in growth and development as well as response to stress in many plant species, including the identification of GmGELP genes as candidates for salinity and drought tolerance in soybean65. The numerous significant MTAs point to the complexity of heat tolerance response in soybean. In other crops, namely, common bean27, peanut28, maize58,66, and sorghum67, a large number of significant SNP were reported that control heat tolerance. While the multitude of SNPs and associated genes are useful to decipher the genetic control of heat tolerance, they have limitations in immediate application in heat-tolerant variety development due to their large number and smaller effects. Therefore, there is a need to explore genomic prediction methods to investigate their usefulness in genomic selection.
Genomic prediction is a powerful tool in plant breeding68, with the ability to estimate the breeding value of accessions without having to use the resources to phenotype in greenhouses or fields. Genomic prediction has been found in soybean to have a higher predictive ability than phenomic selection alone69. We evaluated the effectiveness of genomic selection for soybean biomass and physiological processes in heat versus optimal temperature conditions. We were also interested in using GP to evaluate the prediction in the USDA-GRIN collection that has been previously shown to be effective in soybean70 and maize71. In this study, we observed that GPA was higher for genomic selection applied at the optimal temperature compared to the heat treatment, and that the GPA for a heat index was poor. The GPA is largely dependent on several factors, including linkage disequilibrium (LD), number of markers, heritability, and population size. Given that the same population with the same number of markers was used for training, the differences in GPA can largely be attributed to the differences in trait heritability, which were lower in heat treatment and consistent with previous studies57,72. The GPA for the four biomass traits remained comparable across the heat and optimal treatments, indicating that genomic selection has the potential for selecting biomass traits under heat conditions, although validation in field studies and expansion to seed yield should be studied. In contrast, the physiological traits and the trait indices showed poor GPA, indicating that genomic selection may not be a suitable breeding schema for these traits. However, other GP models could be tested, such as Bayesian models, for their suitability for predicting these traits. While these traits may not be suitable for use as breeding targets via GP tools, these traits remain useful for screening germplasm in determining heat stress tolerance.
The moderate predictive ability of the biomass traits under heat makes them good candidates for use in GP methods, including in advancing breeding materials in stressful environments and searching for heat-tolerant germplasm in large collections such as the USDA soybean collection. Considering that the soybean collection has more than 20,000 accessions available, it is not feasible to phenotype all accessions for heat stress tolerance. By projecting the GP model, GEBVs can be estimated, which can be used by soybean breeders to select genotypes to be used in crossing for heat stress-tolerant varieties, enabling them to utilize accessions appropriate for their maturity group. While the use of the biomass traits in GP does not allow for direct measurement of heat tolerance like the indices allow, this application still allows for predicting which genotypes can perform well in the particular heat stress environment. The germplasm projection trended towards lower maturity groups for a large proportion of the heat-tolerant accession. This could be due to the model being trained primarily on these maturity groups or could indicate a differential response to heat stress between early and later maturity groups. Further efforts should be conducted to screen soybean accessions from additional maturity groups to further investigate the underlying trend of this projection towards lower maturity groups. The efforts to utilize germplasm collection need to be increased as they hold tremendous value in improving traits to combat the adverse effects of climate change.
The results from our study can benefit breeding efforts aimed at early season heat response, as the accessions we report are targeted at the early growth stage. Additional studies are needed to measure heat stress response in critical growth stages, such as flowering, pod and seed initiation. Large-scale screening for heat stress remains a bottleneck due to the difficulty of phenotyping accessions in the field with the option to compare them to optimal temperature treatment. Additional difficulty arises as the areas projected to experience temperature increase in the future do not presently provide fields with natural heat stress, while areas with higher current temperatures are in dissimilar maturity zone and soybean accessions will experience a confounding photoperiod response. Solutions for screening for soybean heat stress tolerance could include the utilization of large indoor spaces, the construction of artificial heat sources in specialized fields, and the development of phenotyping traits to be used in indirect selection of heat tolerance. Traits identified in this study should be explored further in near-field conditions and developed so that HTP methods can be applied for screening heat tolerance in large soybean breeding populations.
In conclusion, our study found that there was significant phenotypic and genotypic variability in soybean response to heat stress in the early season. We also identified phenotypes that could be adapted to field-scale HTP methods, which would allow soybean breeders in the future to screen large amounts of breeding materials for heat stress tolerance in the field. The existence of this natural genetic diversity and a preliminary genetic architecture for traits measured in a heat stress environment indicates that these traits can be improved through breeding strategies. Newer breeding strategies, such as genomic prediction, are valuable tools to overcome the difficulty of phenotyping large numbers of accessions for heat stress tolerance in a field setting. These techniques should be validated in the field in future studies to better understand the reaction of soybeans in a heat-stress setting in otherwise natural field conditions. However, this validation requires the development and optimization of methods to induce heat stress in soybean fields that are cost-effective and safe.
Methods
Plant materials
A panel of 450 diverse soybean accessions70,73 was used to study heat response in controlled environments (Supplementary Table 5). A subset of 418 plant introduction (PI) lines was obtained from the USDA-NPGS soybean mini-core collection74. These PI lines originated from 26 countries. An additional 22 SoyNAM parents75 and ten elite lines commonly used as checks in the Uniform Soybean Trials (UST) were selected for this study. A prior pre-screen process of the PIs was utilized to ensure that only accessions with good standability and non-shattering were included in this study. All accessions were selected for maturity suitable for production in Iowa and range in maturity groups 0-IV.
Experimental design and planting
The experimental design in each greenhouse was a randomized complete block with four replications in each temperature treatment. The replication was used as the blocking factor for placement within the greenhouse, with all four replications of a single treatment in the same greenhouse. To complete phenotypic measurements in a short amount of time to reduce for unaccounted variation from time lag, we planted each replication in the two treatments a day apart in a staggered manner, allowing complete phenotyping of a single replication of a single treatment in a day. For example, the first replication was planted in the control greenhouse on day 1. On day 2, the first replication was planted in the greenhouse with heat treatment. This sequence was followed for the entire experiment, allowing us to maintain consistency in days after planting for data collection. The complete details of the days of planting and data collection are provided in Supplementary Table 6.
Four seeds per accession were sown in a single Conetainer (Stuewe & Sons, Inc. Tangent, OR, USA) filled with sand. Each container had a drainage hole that was covered with landscape fabric (Sta-Green Premium Landscape Fabric, Lowes) to prevent sand loss and ensure well-drained conditions in the cones. A tablespoon of Osmocote slow-release fertilizer (14-14-14) was applied to each cone during planting. Cones were placed into their assigned temperature treatment at planting and remained in the designated treatment of the optimal control or the heat treatment until phenotyping. After emergence, plants were thinned to a single plant per cone. Plants were watered twice daily to ensure no drought stress would occur during the growing period.
Temperature treatments
The experiment was conducted in two greenhouse rooms, one each for the optimal (control with no stress) (29/18°C day/night) and heat (36/29°C day/night) treatment greenhouses. To minimize all other sources of variance except the temperature treatments, we carefully monitored air, light, and temperature through regular monitoring.
Phenotypic measurements
All phenotypic measurements were taken 28 days after planting. Measurements were taken on the center leaflet of the most recently fully developed trifoliate of each plant. Chlorophyll concentration (SPAD) was estimated non-destructively using a SPAD chlorophyll meter (Konica Minolta Sensing Americas Inc., USA). Chlorophyll concentration was measured as a SPAD value, which is unitless, and is calculated as the ratio of the intensity of light transmittance from 650-940 nm (red to infrared). Canopy temperature (CT) was measured using an infrared radiometer (Model SI-411-SS, Apogee Instruments, Logan, UT, USA) from 10 am to 2 pm to measure around the solar noon window. Biomass was collected from each plant and partitioned between shoot and root biomass. Samples were dried at 80°C for three days to remove all moisture and then weighed to get the dry biomass in grams.
During the 2022 data collection, additional traits were captured. An LI-600 (Li-Cor Bioscience, Lincoln, NE, USA) was used to measure the stomatal conductance and Photosystem II quantum efficiency (ΦPSII) during the solar noon window. Additionally, fresh shoot and root biomass weights were taken prior to drying in 2022. A summary of the measured traits is shown in Table 6.
Statistical analysis
Assumptions of analyses of variance (ANOVA) were tested using the Shapiro-Wilk test for normality and Barlett’s test. Additionally, residuals were visually checked via QQ plots and residual plots for heteroscedasticity to check for heterogeneous variance. Box-Cox transformations were used for traits with heteroscedasticity in their residuals. Outliers were removed by calculating studentized residuals for each observation of each trait and excluded from the analysis with values ± 3.476. The following mixed linear model was applied to the untransformed data:
Where, Yij is the phenotypic value of the ith genotype in the jth replication, µ is the overall mean, Gi is the fixed effect of the ith genotype, Rj is the random replication effect of the jth replication, TK is the fixed treatment effect of the kth treatment, Gi:Tk is the random interaction of the genotype and treatment effects, and ε(ij) is the residual error. A type III ANOVA was performed on the resulting mixed linear model in (1).
Broad sense heritability (H2Piepho) was calculated via the H2cal function in the inti package77 in R and was determined by the equation:
Where \({{\boldsymbol{\sigma }}}_{{\boldsymbol{g}}}^{{\boldsymbol{2}}}\) is the genotypic variance and \({\bar{{\boldsymbol{v}}}}_{{\boldsymbol{\triangle }}{\boldsymbol{.}}{\boldsymbol{.}}}^{{\boldsymbol{BLUE}}}\) is the mean variance of the difference between two treatment means (BLUEs)78.
Genetic correlations between traits in the different treatments were calculated using the Multi Environment Trial Analysis in R (META-R) suite79.
The best linear unbiased estimates (BLUEs) for each trait were calculated using the H2cal function. A mixed linear model for each condition (heat – 2021, 2022; optimal – 2021, 2022) was fit with the following equation:
Where Yij is the phenotypic value of the ith genotype in the jth replication, µ is the population mean, Gi is the fixed genotypic contribution for the ith genotype, Rj is the random replication effect of the jth replication, and ε(ij) is the residual error.
Using a completely random model, best linear unbiased predictors (BLUPs) for each trait were also calculated using (3), allowing for calculations of genotypic variance.
The estimated marginal means (emmeans) of each accession was calculated using the emmeans package in R and fit to the same equation as described in (3). Heat tolerance indices (HtI) for all the studied traits were estimated according to the following formula:
Where, YOT is the emmean of an accession under the optimal temperature conditions, and YHT is the emmean of the same accession under heat stress temperature conditions. The emmeans of accessions were used to select responses to heat (tolerance or susceptibility). Accessions were ranked on their HtI and the cumulative ranking of all traits was used to select the top five heat tolerant and top five heat-susceptible accessions.
Genome-wide association studies
Genome-wide association studies (GWAS) were performed using HtI and other traits BLUEs with the SVEN methodology80. Of the panel of 450 accessions, 423 had SoySNP50K publicly available81, and only these accessions were included in GWAS. The SoySNP50K data has 47K markers across the genome, which were imported into TASSEL and filtered on a minor allele frequency (MAF) of 1%. A MAF of 1% was selected so that rare variants were removed, but low-frequency SNP variants were still included. Genotypic data were imputed and then used to calculate a kinship matrix with Centered IBS in TASSEL. A principal component analysis (PCA) was performed with four components to determine familial or population structure, which was input in GWAS. The kinship matrix and PCA from TASSEL were utilized for GWAS in SVEN.
SVEN is a Bayesian method for GWAS based on a hierarchical multi-locus model. It can control for false discovery through prior regularization on the number of important markers80,82. The bravo package80 was implemented in R to run SVEN. Given the complexity of abiotic stress traits, we used a larger prior shrinkage lambda of 20. We have previously found this value to be optimal for quantitative traits in soybean83, and higher values are recommended for quantitative traits80. Markers with a marginal inclusion probability (MIP) greater than 0.5 were declared as significant and reported. Candidate gene search was done using the Genome Browser available on Soybase (soybase.org)84 with the Wm82.a2 assembly.
Genomic prediction (GP)
Underlying population structure due to maturity groups was tested for through a principal component and a discriminant analysis (DA) using the LEA package in R85. Using maturity groups as the known clusters, a ten-fold cross-validation of the resulting DA model was conducted to determine if the model could accurately classify validation genotypes into their proper maturity groups. An admixture analysis to identify ancestral clusters was conducted via the sparse nonnegative matrix factorization (SNMF) methodology. This analysis was run using a range of 1-100 ancestral populations and cross-entropy was plotted to determine the lowest values.
The genomic best linear unbiased prediction (GBLUP) methodology was utilized for genomic selection (GS) using the rrBLUP package86 in R. Phenotypic BLUP data were randomly divided into a training set (80%) and a validation set (20%). The full set of SNP markers from the 423 accessions was used for training, and a separate model was trained on a subset of GWAS significant markers that had a p ≤ 0.05 in TASSEL. A ten-fold cross-validation (CV) was performed to avoid inflated estimates of predictive ability. The list of training genotypes was divided into ten equally sized subgroups for the ten-fold CV. Nine of these subgroups were used for training the prediction model, and the remaining subgroup was used as the testing set. The testing set was used to assess the correlation between the genomic estimated breeding values (GEBVs) of the predicted trait value and the BLUPs of the observed trait value, using the Pearson correlation coefficient. This process was repeated ten times, with each of the subgroups being the testing population once. The process was repeated for 100 cycles by randomly reforming the folds with a different subset of genotypes in each cycle. Across all folds and cycles, the 20% validation set remained unobserved. The correlation accuracy between the GEBVs and BLUPs of the validation population was generated in each cycle. This process was repeated separately for training and predicting in the optimal environment and in the heat environment, and the HtI. The genomic prediction accuracies (GPAs) are reported as the means of the validation (rVS) and testing (rTS) correlation coefficients.
Using the developed GP model and the public SoySNP50K data available on SoyBase.org, the GEBV for 19,840 accessions in the USDA soybean germplasm collection was calculated for SPAD, DSB, and DRB for both years, and the top 5% (992 accessions) and 10% (1985 accessions) were selected. The top 5% (21 accessions) and 10% (42 accessions) most heat-tolerant lines were identified based on their GEBV for all traits for each year in the current panel of 423 accessions. In addition, aiming to understand the diversity between the selected accessions, the country of origin, and the maturity group were verified for the top 5% USDA germplasm. To verify the performance of the accessions tested in this research with the USDA top accessions, boxplots were generated to visually compare the 5% and 10% most tolerant accessions identified in the USDA soybean germplasm collection with the 5% and 10% most tolerant accessions identified in the current research.
Data availability
Data is available upon reasonable request to the corresponding author.
References
OECD & FAO - United Nations. in OECD-FAO Agricultural Outlook (OECD, 2023). https://doi.org/10.1787/308c1c56-en.
Sedivy, E. J., Wu, F. & Hanzawa, Y. Soybean domestication: the origin, genetic architecture and molecular bases. New Phytol 214, 539–553 (2017).
Leff, B., Ramankutty, N. & Foley, J. A. Geographic distribution of major crops across the world. Global Biogeochem. Cycles 18 (2004).
Vu, J. C. V., Allen, L. H., Jr Boote, K. J. & Bowes, G. Effects of elevated CO2 and temperature on photosynthesis and Rubisco in rice and soybean. Plant Cell Environ 20, 68–76 (1997).
Oerke, E.-C. Crop losses to pests. J. Agric. Sci. 144, 31–43 (2006).
Hatfield, J. L. et al. Climate impacts on agriculture: Implications for crop production. Agron. J. 103, 351–370 (2011).
Gray, S. B. et al. Intensifying drought eliminates the expected benefits of elevated carbon dioxide for soybean. Nat Plants 2, 16132 (2016).
Jin, Z., Ainsworth, E. A., Leakey, A. D. B. & Lobell, D. B. Increasing drought and diminishing benefits of elevated carbon dioxide for soybean yields across the US Midwest. Glob. Chang. Biol. 24, e522–e533 (2018).
Thomey, M. L., Slattery, R. A., Köhler, I. H., Bernacchi, C. J. & Ort, D. R. Yield response of field-grown soybean exposed to heat waves under current and elevated [CO2]. Glob. Chang. Biol. 25, 4352–4368 (2019).
Yang, M. & Wang, G. Heat stress to jeopardize crop production in the US Corn Belt based on downscaled CMIP5 projections. Agric. Syst. 211, 103746 (2023).
Sun, Q. et al. Global heat stress on health, wildfires, and agricultural crops under different levels of climate warming. Environ. Int. 128, 125–136 (2019).
Teixeira, E. I., Fischer, G., van Velthuizen, H., Walter, C. & Ewert, F. Global hot-spots of heat stress on agricultural crops due to climate change. Agric. For. Meteorol. 170, 206–215 (2013).
Mousavi-Derazmahalleh, M. et al. Adapting legume crops to climate change using genomic approaches. Plant Cell Environ 42, 6–19 (2019).
IPCC. in Climate Change 2022—Impacts, Adaptation and Vulnerability: Working Group II Contribution to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change 713–906 (Cambridge University Press, 2023). https://doi.org/10.1017/9781009325844.007.
Singh, D. P., Singh, A. K. & Singh, A. Plant Breeding and Cultivar Development (Elsevier Science, 2021).
Deshmukh, R. et al. Integrating omic approaches for abiotic stress tolerance in soybean. Front. Plant Sci. 5, 244 (2014).
Bita, C. E. & Gerats, T. Plant tolerance to high temperature in a changing environment: scientific fundamentals and production of heat stress-tolerant crops. Front. Plant Sci. 4, 273 (2013).
Prasad, P. V. V., Bheemanahalli, R. & Jagadish, S. V. K. Field crops and the fear of heat stress—Opportunities, challenges and future directions. Field Crops Res 200, 114–121 (2017).
Jagadish, S. V. K., Way, D. A. & Sharkey, T. D. Plant heat stress: Concepts directing future research. Plant Cell Environ 44, 1992–2005 (2021).
Almeselmani, M., Deshmukh, P. S. & Chinnusamy, V. Effects of prolonged high temperature stress on respiration, photosynthesis and gene expression in wheat (Triticum aestivum L.) varieties differing in their thermotolerance. Plant stress 6, 25–32 (2012).
Moore, C. E. et al. The effect of increasing temperature on crop photosynthesis: from enzymes to ecosystems. J. Exp. Bot. 72, 2822–2844 (2021).
Poudel, S. et al. Quantifying the physiological, yield, and quality plasticity of Southern USA soybeans under heat stress. Plant Stress 9, 100195 (2023).
Jumrani, K. & Bhatia, V. S. Interactive effect of temperature and water stress on physiological and biochemical processes in soybean. Physiol. Mol. Biol. Plants 25, 667–681 (2019).
Tesfaye, K. et al. Climate change impacts and potential benefits of heat-tolerant maize in South Asia. Theor. Appl. Climatol. 130, 959–970 (2017).
Balla, K. et al. Heat stress responses in a large set of winter wheat cultivars (Triticum aestivum L.) depend on the timing and duration of stress. PLoS One 14, e0222639 (2019).
Tafesse, E. G., Gali, K. K., Lachagari, V. B. R., Bueckert, R. & Warkentin, T. D. Genome-wide association mapping for heat stress responsive traits in field pea. Int. J. Mol. Sci. 21 (2020).
López-Hernández, F. & Cortés, A. J. Last-Generation Genome–Environment Associations Reveal the Genetic Basis of Heat Tolerance in Common Bean (Phaseolus vulgaris L.). Front. Genet. 10 (2019).
Sharma, V. et al. Genetic mapping identified three hotspot genomic regions and candidate genes controlling heat tolerance-related traits in groundnut. Front. Plant Sci. 14, 1182867 (2023).
Maalouf, F. et al. Genetic Dissection of Heat Stress Tolerance in Faba Bean (Vicia faba L.) Using GWAS. Plants 11 (2022).
Paul, P. J. et al. Molecular mapping of QTLs for heat tolerance in Chickpea. Int. J. Mol. Sci. 19 (2018).
Li, P.-S. et al. Genome-wide analysis of the Hsf family in soybean and functional identification of GmHsf-34 involvement in drought and heat stresses. BMC Genomics 15, 1009 (2014).
Song, K., Yim, W. C. & Lee, B.-M. Expression of heat shock proteins by heat stress in soybean. Plant Breed. Biotechnol. 5, 344–353 (2017).
Valdés-López, O. et al. Soybean roots grown under heat stress show global changes in their transcriptional and proteomic profiles. Front. Plant Sci. 7, 517 (2016).
Wang, L. et al. Transcriptome profilling analysis characterized the gene expression patterns responded to combined drought and heat stresses in soybean. Comput. Biol. Chem. 77, 413–429 (2018).
Kidokoro, S. et al. Soybean DREB1/CBF-type transcription factors function in heat and drought as well as cold stress-responsive gene expression. Plant J 81, 505–518 (2015).
Ruiz-Vera, U. M. et al. Global warming can negate the expected CO2 stimulation in photosynthesis and productivity for soybean grown in the Midwestern United States. Plant Physiol 162, 410–423 (2013).
Tacarindua, C. R. P., Shiraiwa, T., Homma, K., Kumagai, E. & Sameshima, R. The effects of increased temperature on crop growth and yield of soybean grown in a temperature gradient chamber. Field Crops Res 154, 74–81 (2013).
Siebers, M. H. et al. Heat waves imposed during early pod development in soybean (Glycine max) cause significant yield loss despite a rapid recovery from oxidative stress. Glob. Chang. Biol. 21, 3114–3125 (2015).
Alsajri, F. A. et al. Developing functional relationships between temperature and soybean yield and seed quality. Agron. J. 112, 194–204 (2020).
Cohen, I., Zandalinas, S. I., Huck, C., Fritschi, F. B. & Mittler, R. Meta-analysis of drought and heat stress combination impact on crop yield and yield components. Physiol. Plant. 171, 66–76 (2021).
Alsajri, F. A. et al. Evaluating Soybean Cultivars for Low- and High-Temperature Tolerance During the Seedling Growth Stage. Agronomy 9, 13 (2019).
Zhang, J. et al. Genome-wide Scan for Seed Composition Provides Insights into Soybean Quality Improvement and the Impacts of Domestication and Breeding. Mol. Plant 11, 460–472 (2018).
Steketee, C. J., Schapaugh, W. T., Carter, T. E. & Li, Z. Genome-wide association analyses reveal genomic regions controlling canopy wilting in soybean. G3 10, 1413–1425 (2020).
Kaler, A. S., Ray, J. D., Schapaugh, W. T., King, C. A. & Purcell, L. C. Genome-wide association mapping of canopy wilting in diverse soybean genotypes. Theor. Appl. Genet. 130, 2203–2217 (2017).
Wu, C. et al. Genome-wide association mapping of flooding tolerance in soybean. Mol. Breed. 40, 4 (2019).
Assefa, T. et al. Deconstructing the genetic architecture of iron deficiency chlorosis in soybean using genome-wide approaches. BMC Plant Biol 20, 42 (2020).
Mamidi, S. et al. Genome-wide association analysis identifies candidate genes associated with iron deficiency chlorosis in soybean. Plant Genome 4, 154–164 (2011).
Hesketh, J. D., Myhre, D. L. & Willey, C. R. Temperature control of time intervals between vegetative and reproductive events in soybeans1. Crop Sci 13, 250–254 (1973).
Wang, N. et al. Genomic prediction across years in a maize doubled haploid breeding program to accelerate early-stage testcross testing. Theor. Appl. Genet. 133, 2869–2879 (2020).
Bai, Y., Ali, S., Liu, S., Zhou, J. & Tang, Y. Characterization of plant laccase genes and their functions. Gene 852, 147060 (2023).
Tran, L.-S. P. et al. Molecular characterization of stress-inducible GmNAC genes in soybean. Mol. Genet. Genomics 281, 647–664 (2009).
Melo, B. P. et al. Revisiting the Soybean GmNAC Superfamily. Front. Plant Sci. 9, 1864 (2018).
Beuchat, J. et al. BRX promotes Arabidopsis shoot growth. New Phytol 188, 23–29 (2010).
Dhanapal, A. P. et al. Genome-wide association mapping of soybean chlorophyll traits based on canopy spectral reflectance and leaf extracts. BMC Plant Biol 16, 174 (2016).
Zhou, R. et al. Drought stress had a predominant effect over heat stress on three tomato cultivars subjected to combined stress. BMC Plant Biol 17, 24 (2017).
Al-Yassin, A., Grando, S., Kafawin, O., Tell, A. & Ceccarelli, S. Heritability estimates in contrasting environments as influenced by the adaptation level of barley germ plasm. Ann. Appl. Biol. 147, 235–244 (2005).
Kumar, R., Venuprasad, R. & Atlin, G. N. Genetic analysis of rainfed lowland rice drought tolerance under naturally-occurring stress in eastern India: Heritability and QTL effects. Field Crops Res 103, 42–52 (2007).
Longmei, N. et al. Genome wide association mapping for heat tolerance in sub-tropical maize. BMC Genomics 22, 154 (2021).
Yuan, Y. et al. Genome-Wide Association Mapping and Genomic Prediction Analyses Reveal the Genetic Architecture of Grain Yield and Flowering Time Under Drought and Heat Stress Conditions in Maize. Front. Plant Sci. 9 (2019).
Soltani, A., Weraduwage, S. M., Sharkey, T. D. & Lowry, D. B. Elevated temperatures cause loss of seed set in common bean (Phaseolus vulgaris L.) potentially through the disruption of source-sink relationships. BMC Genomics 20, 312 (2019).
Zhou, X. et al. Phytoene Synthase: The key rate-limiting enzyme of carotenoid biosynthesis in plants. Front. Plant Sci. 13, 884720 (2022).
Young, A. J. The photoprotective role of carotenoids in higher plants. Physiol. Plant. 83, 702–708 (1991).
Li, Z. et al. OsBRXL4 regulates shoot gravitropism and rice tiller angle through affecting LAZY1 nuclear localization. Mol. Plant 12, 1143–1156 (2019).
Zhang, Y. et al. Cysteine-rich receptor-like protein kinases: emerging regulators of plant stress responses. Trends Plant Sci 28, 776–794 (2023).
Su, H.-G. et al. Genome-wide identification, evolution, and expression of gdsl-type esterase/lipase gene family in soybean. Front. Plant Sci. 11, 726 (2020).
Seetharam, K. et al. Genomic regions associated with heat stress tolerance in tropical maize (Zea mays L.). Sci. Rep. 11, 13730 (2021).
Chen, J. et al. Genome-wide association study of developing leaves’ heat tolerance during vegetative growth stages in a sorghum association panel. Plant Genome 10 (2017).
Crossa, J. et al. Genomic selection in plant breeding: methods, models, and perspectives. Trends Plant Sci 22, 961–975 (2017).
Howard, R. & Jarquin, D. Genomic prediction using canopy coverage image and genotypic information in soybean via a hybrid model. Evol. Bioinform. Online 15, 1176934319840026 (2019).
de Azevedo Peixoto, L. et al. Leveraging genomic prediction to scan germplasm collection for crop improvement. PLoS ONE 12, e0179191 (2017).
Yu, X. et al. Genomic prediction contributing to a promising global strategy to turbocharge gene banks. Nat Plants 2, 16150 (2016).
Ud-Din, N., Carver, B. F. & Clutter, A. C. Genetic analysis and selection for wheat yield in drought-stressed and irrigated environments. Euphytica 62, 89–96 (1992).
Moellers, T. C. et al. Main and epistatic loci studies in soybean for Sclerotinia sclerotiorum resistance reveal multiple modes of resistance in multi-environments. Sci. Rep. 7, 3554 (2017).
Oliveira, M. F., Nelson, R. L., Geraldi, I. O., Cruz, C. D. & de Toledo, J. F. F. Establishing a soybean germplasm core collection. Field Crops Res 119, 277–289 (2010).
Song, Q. et al. Genetic characterization of the soybean nested association mapping population. Plant Genome 10 (2017).
Lund, R. E. Tables for an approximate test for outliers in linear models. Technometrics 17, 473–476 (1975).
Lozano-Isla, F. inti: Tools and statistical procedures in plant science. R Package Version 0 (2021).
Piepho, H.-P. & Möhring, J. Computing heritability and selection response from unbalanced plant breeding trials. Genetics 177, 1881–1888 (2007).
Alvarado, G. et al. META-R: A software to analyze data from multi-environment plant breeding trials. Crop J 8, 745–756 (2020).
Li, D., Dutta, S. & Roy, V. Model based screening embedded bayesian variable selection for ultra-high dimensional settings. J. Comput. Graph. Stat. 32, 61–73 (2023).
Song, Q. et al. Fingerprinting soybean germplasm and its utility in genomic research. G3 5, 1999–2006 (2015).
Rairdin, A. et al. Deep learning-based phenotyping for genome wide association studies of sudden death syndrome in soybean. Front. Plant Sci. 13, 966244 (2022).
Van der Laan, L. et al. Genomic and phenomic prediction for soybean seed yield, protein, and oil. Plant Genome 18, e70002 (2025).
Brown, A. V. et al. A new decade and new data at SoyBase, the USDA-ARS soybean genetics and genomics database. Nucleic Acids Res. 49, D1496–D1501 (2021).
Frichot, E. & François, O. L. E. A. An R package for landscape and ecological association studies. Methods Ecol. Evol. 6, 925–929 (2015).
Endelman, J. B. Ridge regression and other kernels for genomic selection with R package rrBLUP. Plant Genome 4, 250–255 (2011).
Acknowledgements
The authors thank the undergraduate and graduate students for their assistance during the data collection. We express our gratitude to Jennifer Hicks and Antonella Ferela for their assistance in procuring materials and organizing assistance for this project. We thank Aaron Brand and Peter Lawlor for ensuring the greenhouses were running in the appropriate settings. We appreciate support from Dr. Phillip Dixon and Ms. Laritza Lorenzo for their guidance and feedback on the statistical analysis, and Dr. Jacqueline Campbell for assistance in the search for candidate genes. The authors sincerely appreciate the funding support from Iowa Soybean Association, USDA CRIS project IOW04714, AI Institute for Resilient Agriculture (USDA-NIFA #2021-67021-35329), COALESCE: COntext Aware LEarning for Sustainable CybEr-Agricultural Systems (CPS Frontier #1954556), Smart Integrated Farm Network for Rural Agricultural Communities (SIRAC) (NSF S&CC #1952045), Raymond F. Baker Center for Plant Breeding, and Plant Sciences Institute.
Author information
Authors and Affiliations
Contributions
L.V. and A.K.S. conceptualized and designed the experiment. L.V. performed data collection with the assistance of lab members, performed the data analysis, and wrote the first draft manuscript with feedback from AKS. L.A.P. performed the genomic projection analysis and provided feedback on the draft manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Van der Laan, L., de Azevedo Peixoto, L. & Singh, A.K. Genetic dissection of heat stress tolerance in soybean through genome-wide association studies and use of genomic prediction to enhance breeding applications. npj Sci. Plants 1, 9 (2025). https://doi.org/10.1038/s44383-025-00010-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44383-025-00010-8