Abstract
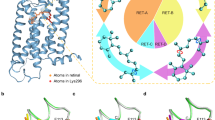
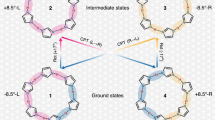
The dramatic progress in the understanding of the dynamics of biomolecules has been largely fuelled by computer simulations based on the law of classical mechanics1. However in some respects biomolecules are at the borders of the domain of applicability of classical mechanics. The role of quantum mechanical effects in biomolecular structure and function is therefore worth investigating. Here we present preliminary results from a quantum simulation of a protein and contrast them with results from full classical simulations. The most significant differences are found in motions of high frequency, such as bond stretching or the torsional oscillation of groups that bear hydrogen atoms. The amplitudes of such motions are significantly increased by the penetration of atoms into classically forbidden regions. These differences will directly influence the rates of such processes as proton and electron transfer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
McCammon, J. A. & Harvey, S. C. Dynamics of Proteins and Nucleic Acids (Cambridge University Press, 1987).
Frauenfelder, H. & Wolynes, P. G. Science 229, 337–345 (1985).
Tunneling in Biological Systems (eds Chance, B., et al.) (Academic, New York, 1979).
DeVault, D. Quantum-mechanical Tunnelling in Biological Systems 2nd edn (Cambridge University Press 1984).
Mayo, S. L., Ellis, W. R. Jr., Crutchley, R. J. & Gray, H. B. Science 233, 948–952 (1986).
Kihara, T. & McCray, J. A. Biochim. biophys. Acta 292, 297–309 (1973).
Albery, W. J. & Knowles, J. R. J. Am. chem. Soc. 99, 637–638 (1977).
Goldanskii, V. I., Krupyanskii, Yu. F. & Flerov, V. N. Doklady Biophysics 272, 209–212 (1984).
Ringe, D. & Petsko, G. A. Prog. Biophys. molec. Biol. 45, 197–235 (1985).
Parak, F., Frolov, E. N., Mössbauer, R. L. & Goldanskii, V. I. J. molec. Biol. 145, 825–833 (1981).
Friedman, H. L. A Course in Statistical Mechanics (Prentice-Hall, Englewood Cliffs, 1985).
Feynman, R. P. Statistical Mechanics (Benjamin, London, 1972).
Chandler, D. & Wolynes, P. G. J. chem. Phys. 74, 4078–4095 (1981).
Berne, B. J. & Thiriumalai, D. A. Rev. phys. Chem. 37, 401–424 (1986).
van Gunsteren, W. F., Berendsen, H. J. C., Hermans, J., Hol, W. G. J. & Postma, J. P. M. Proc. natn. Acad. Sci. U.S.A. 80, 4315–4319 (1983).
Takano, T. & Dickerson, R. E. J. molec. Biol. 153, 79–94 (1981).
Northrup, S. H., Pear, M. R., Morgan, J. D., McCammon, J. A. & Karplus, M. J. molec. Biol. 153, 1087–1109 (1981).
Moore, G. R. et al. Faraday Discuss. Chem. Soc. 74, 311–329 (1982).
Wolynes, P. G. J. chem. Phys. 87, 6559–6561.(1987).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Zheng, C., Wong, C., McCammon, J. et al. Quantum simulation of ferrocytochrome c. Nature 334, 726–728 (1988). https://doi.org/10.1038/334726a0
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/334726a0