ABSTRACT
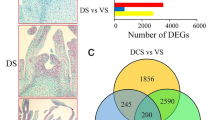
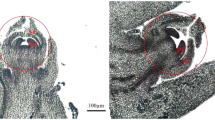
The homologous genes FLORICAULA (FLO) in Antirrhinum and LEAFY (LFY) in Arabidopsis are known to regulate the initiation of flowering in these two distantly related plant species. These genes are necessary also for the expression of downstream genes that control floral organ identity. We used Arabidopsis LFY cDNA as a probe to clone and sequence a papaya ortholog of LFY, PFL. It encodes a protein that shares 61% identity with the Arabidopsis LFY gene and 71% identity with the LFY homologs of the two woody tree species: California sycamore (Platanus racemosa) and black cottonwood (Populus trichocarpa). Despite the high sequence similarity within two conserved regions, the N-terminal proline-rich motif in papaya PFL differs from other members in the family. This difference may not affect the gene function of papaya PFL, since an equally divergent but a functional LFY ortholog NEEDLY of Pinus radiata has been reported. Genomic and BAC Southern analyses indicated that there is only one copy of PFL in the papaya genome. In situ hybridization experiments demonstrated that PFL is expressed at a relatively low level in leaf primordia, but it is expressed at a high level in the floral meristem. Quantitative PCR analyses revealed that PFL was expressed in flower buds of all three sex types - male, female, and hermaphrodite with marginal difference between hermaphrodite and unisexual flowers. These data suggest that PFL may play a similar role as LFY in flower development and has limited effect on sex differentiation in papaya.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Liljegren SJ, Gustafson-Brown C, Pinyopich A, Ditta GS, Yanofsky MF . Interactions among APETALA1, LEAFY, and TERMINAL FLOWER1 specify meristem fate. Plant Cell 1999; 11:1007–18.
Wagner D, Sablowski RWM, Meyerowitz EM . Transcriptional activation of APETALA1 by LEAFY. Science 1999; 285:582–4.
Weigel D, Meyerowitz EM . Activation of floral homeotic genes in Arabidopsis. Science 1993; 261:1723–26.
Parcy F, Nilsson O, Busch MA, Lee I, Weigel D . A genetic framework for floral patterning. Nature 1998; 395:561–6.
Blázquez MA, Soowal L, Lee I, Weigel D . LEAFY expression and flower initiation in Arabidopsis. Development 1997; 124:3835–44.
Haughn GW, Somerville CR . Genetic control of morphogenesis in Arabidopsis. Dev Genet 1988; 9:73–89.
Meyerowitz EM, Bowman JL, Brockman LL, et al. A genetic and molecular model for flower development in Arabidopsis thaliana. Dev Suppl 1991; 1:157–67.
Schultz EA, Haughn GW . LEAFY, a homeotic gene that regulates inflorescence development in Arabidopsis. Plant Cell 1991; 3:771–81.
Weigel D, Alvarez J, Smyth DR, Yanofsky MF, Meyerowitz EM . LEAFY controls floral meristem identity in Arabidopsis. Cell 1992; 69:843–59.
Mandel MA, Yanofsky MF . A gene triggering flower formation in Arabidopsis. Nature 1995; 377:522–4.
Weigel D, Nilsson O . A development switch sufficient for flower initiation in diverse plants. Nature 1995; 377:495–500.
Blázquez MA, Weigel D . Integration of floral inductive signals in Arabidopsis. Nature 2000; 404:889–92.
Busch MA, Bomblies K, Weigel D . Activation of a floral homeotic gene in Arabidopsis. Science 1999; 285:585–7.
Blázquez MA . Flower development pathways. J Cell Sci 2000; 113:3547–48.
Ng M, Yanofsky MF . Three ways to learn the ABCs. Curr Opin Plant Biol 2000; 3:47–52.
Ma H, de Pamphilis C . The ABCs of floral evolution. Cell 2000; 101:5–8.
Coen ES, Romero JM, Doyle S, et al. floricaula: A homeotic gene required for flower development in Antirrhinum majus. Cell 1990; 63:1311–22
Mouradov A, Glassick T, Hamdorf B, et al. NEEDLY, a Pinus radiata ortholog of FLORICAULA/LEAFY genes, expressed in both reproductive and vegetative meristems. PNAS 1998; 95:6537–42.
Molinero-Rosales N, Jamilena M, Zurita S, et al. The tomato orthologue of FLORICAULA and LEAFY, controls flowering time and floral meristem identity. Plant J 1999; 20:685–93.
Kelly A, Bonnlander MB, Meeks-Wagner DR . NFL, the tobacco homolog of Floricaula and Leafy, is transcriptionally expressed in both vegetative and floral meristems. Plant Cell 1995; 7:225–34.
Kyozuka J, Konishi S, Nemoto K, Izawa T, Shimamoto K . Down-regulation of RFL, the FLO/LFY homolog of rice, accompanied with panicel branch initiation. PNAS 1998; 95:1979–82
Hofer J ., Turner L ., Hellens R, et al. UNIFOLIATA regulates leaf and flower morphogenesis in pea. Curr Biol 1997; 7:581–7.
Gocal GFW, King RW, Blundell CA, et al. Evolution of floral meristem identity genes. Analysis of Lolium temulentum genes related to APETALA1 and LEAFY of Arabidopsis. Plant Physiol 2001; 125:1788–801.
Bomblies K, Wang R -L, Ambrose BA, et al. Duplicate FLORICAULA/LEAFY homologs zfl1 and zfl2 control inflorescence architecture and flower patterning in maize. Development 2003; 130:2385–95.
Ahearn KP, Johnson HA, Weigel D, Wagner D Ry . NFL1, a Nicotiana tabacum LEAFY-Like gene, controls meristem initiation and floral structure. Plant Cell Physiol 2001; 42:1130–9
Peña L, Martin-Trillo M, Juarez J, et al. Constitutive expression of Arabidopsis LEAFY or APETALA1 genes in citrus reduces their generation time. Nat Biotechnol, 2001; 19:263–7.
Chujo A, Zhang Z, Kishino H, Shimamoto K, Kyozuka J . Partial conservation of LFY function between rice and Arabidopsis. Plant Cell Physiol 2003; 44:1311–9.
Bremer K, Chase MW, Stevens PF . An ordinal classification for the families of flowering plants. Ann Mo Bot Gard 1998; 85:531–53.
Liu Z, Moore PH, Ma H, et al. A primitive Y chromosome in papaya marks incipient sex chromosome evolution. Nature 2004; 427:348–52.
Hall TC, Buchbinder BU, Pyne JW, Sun SM, Bliss FA . Messenger RNA for G1 protein of French bean seeds: cell-free translation and product characterization. Proc Natl Acad Sci U S A 1978; 75:3196–200.
Tai TH, Tanksley, SD . A rapid and inexpensive method for isolation of total DNA from dehydrated plant tissue. Plant Mol Biol Rep 1990; 8:297–303.
Sambrook J, Fritsch EF, Maniatis T . Molecular cloning, 2nd edn. Cold Spring Harbor: Cold Spring Harbor Laboratory Press, 1989.
Chittenden LM, Schertz KF, Lin YR, Wing RA, Paterson AH . A detailed RFLP map of Sorghum bicolor x S. propinquum, suitable for high-density mapping, suggests ancestral duplication of Sorghum chromosomes or chromosomal segments. Theor Appl Genet 1994; 87:925–33.
Jackson D . In-situ hybridization in plants. In: Bowles, DJ, Gurr SJ and McPherson, M. Eds. Molecular plant pathology: A practical approach. Oxford University Press: London 1991:163–74.
Nakasone HY, Storey WB . Studies on the inheritance of fruiting height of Carica papaya L. Proc Amer Soc Hort Sci 1955; 66:168–82.
Carmona MJ, Cubas P, MartÍnez-Zapater JM . VFL, the grapevine FLORICAULA/LEAFY ortholog, is expressed in meristematic regions independently of their fate. Plant Physiol 2002; 130:68–77.
Lamb RS, Hill TA, Tan QKG, Irish VF . Regulation of APETALA3 floral homeotic gene expression by meristem identity genes. Development 2002; 129:2079–86.
William DA, Su Y, Smith MR, et al. Genomic identification of direct target genes of LEAFY. Proc Natl Acad Sci U S A 2004; 101:1775–80.
Maizel A, Busch MA, Tanahashi T, et al. The floral regulator LEAFY evolves by substitutions in the DNA binding domain. Science 2005; 308:260–3.
Southerton SG, Strauss SH, Olive MR, et al. Eucalyptus has a functional equivalent of Arabidopsis floral meristem identity gene LEAFY. Plant Mol Biol 1998; 37:897–910.
Wada M, Cao Q-F, Kotoda N, Soejima J-I, Masuda T . Apple has two orthologues of FLORICAULA/LEAFY involved in flowering. Plant Mol Biol 2002; 49:567–77.
Acknowledgements
We thank Marc CREAPAU, Peizhu GUAN, Denise STEIGER and Jim CARR for technical assistance, Elliot MEYEROWITZ for providing the Arabidopsis LFY cDNA clone, and Mingli WANG for review the manuscript. This work was supported by a USDA-ARS Cooperative Agreement (CA 58-3020-8-134) with the Hawaii Agriculture Research Center.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
YU, Q., MOORE, P., ALBERT, H. et al. Cloning and characterization of a FLORICAULA/LEAFY ortholog, PFL, in polygamous papaya. Cell Res 15, 576–584 (2005). https://doi.org/10.1038/sj.cr.7290327
Received:
Revised:
Accepted:
Issue date:
DOI: https://doi.org/10.1038/sj.cr.7290327
Keywords
This article is cited by
-
Genome-wide Transcriptome Analysis Reveals the Gene Regulatory Network in Star Fruit Flower Blooming
Tropical Plant Biology (2023)
-
Evolution and expression of LEAFY genes in ferns and lycophytes
EvoDevo (2022)
-
An ortholog of LEAFY in Jatropha curcas regulates flowering time and floral organ development
Scientific Reports (2016)
-
Cloning and characterization of a FLO/LFY ortholog in Gossypium hirsutum L.
Plant Cell Reports (2013)
-
Identification and Characterization of PpLFL, a Homolog of FLORICAULA/LEAFY in Peach (Prunus persica)
Plant Molecular Biology Reporter (2012)