Abstract
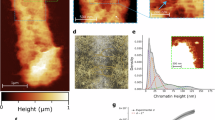
The organization of the higher order structure of chromatin in chicken erythrocytes has been examined with tapping-mode scanning force microscopy under conditions close to their native environment. Reproducible high-resolution AFM images of chromatin compaction at several levels can be demonstrated. An extended beads-on-a-string (width of ∼ 15-20nm, height of ∼ 2-3nm for each individual nucleosome) can be consistently observed. Furthermore, superbeads (width of ∼ 40nm, height of ∼ 7nm) are demonstrated. Visualization of the solenoid conformation at the level of 30nm chromatin fiber is attained either by using AFM or by using electron microscopy. In addition, tightly coiled chromatin fibers (∼ 50-60nm and ∼ 90-110nm) can be revealed. Our data suggest that the chromatin in the interphase nucleus of chicken erythrocyte represents a high-order conformation and AFM provides useful high-resolution structural information concerning the folding pattern of interphase chromatin fibers.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Sedat J, L Manuelidis . A direct approach to the structure of eukaryotic chromosomes. Coldspring harbor symp. Quant Biol 1997; 42:331–50.
Paulson JR, UK Laemmli, The structure of histone depleted metaphase chromosomes. Cell 1977; 12:817–28.
Marsden MPF, UK Laemmli, Metaphase chromosome structure: evidence for a radial loopmodel. Cell 1979; 17:849–58.
Adolph KW . Organization of chromosomes in mitotic HeLa cells. Exp Cell Res 1980; 125:95–103.
Rattner JB, CC Lin . Radial loops and helical coils coexist in metaphase chromosomes. Cell 1985; 42:291–6.
Boy de la Tour E, UK Laemmli . The metaphase scaffold is helically folded: sister chromatidshave predominantly opposite helical handedness. Cell 1988; 55:937–44.
Leuba SH, Guoliang Yang, Charles Robert, Bruno Samori, Kensal van Holde, Jordanka Zla-tanova, Carlos Bustamante . Three-dimensional structure of extended chromatin fibers as re-vealed by tapping-mode scanning force microscopy. Proc Natl Acad Sci USA 1994; 91:11621–5.
Zlatanova J, Leuba SH, Guoliang Yang, Carlos Bustamante, Kensal van Holde . Linker DNA accessibility in chromatin fibers of different conformations: A reevaluation. Proc Natl Acad Sci USA 1994; 91:5277–80.
Martin LD, Vesenka JP, Henderson E, Dobbs DL . Visualization of nucleosomal substructure innative chromatin by atomic force microscopy. Biochemistry 1995; 34:4610–6.
Guoliang Yang, Sanford H Leuba, Carlos Bustamante, Jordanka Zlatanova, Kensal Van Holde . Role of linker histones in extended chromatin fibre structure. Structural Biology 1994; 1:761–3.
Lyubchenko YL . et al Atomic force microscopy of DNA and bacteriophage in air, water andpropanol: the role of adhesion forces. Nucleic Acids Res 1993; 21:1117–23.
Jing-de Zhu, Xiao-ping Sun, Fan Wang . The DNA intercalator, ethidium bromide, alters thepattern of DNase I hypersensitive sites of the βA-globin gene in chicken erythrocytes. Biochem Biophys Acta 1991;1089:158–66.
Gall J . Chromosome fibers from an interphase nucleus. Science 1963; 139 120–1.
Belmont AS, Bruce K . Visualization of G1 chromosomes: A folded, Twisted, Supercoiledchromonema model of interphase chromatid structure. J Cell Biol 1994; 127:287–302.
Fritzsche W, Schaper A, Jovin TM . Probing chromatin with the scanning force microscope. Chromosoma 1994; 103:231–6.
Finch JT, Klug A . Solenoidal model for superstructure in chromatin. Proc Natl Acad Sci USA 1976; 73:1897–901.
McGhee JD, Rau DC, Charney E, Felsenfeld G . Orientation of the nucleosome within the higherorder structure of chromatin. Cell 1980; 22:87–96.
Worcel A, Strogatz S, Riley D . Structure of chromatin and the linking number of DNA. ProcNatl Acad Sci USA 1981;78:1461–5.
Woodcock CLF, Frado L-LY, Rattner JB . The higher-order structure of chromatin: Evidencefor a helical ribbon arrangement. J Cell Biol 1984; 99 42–52.
Woodcock CL, Grigoryev SA, Horowitz RA, Whitaker N . A chromatin folding model that in-corporates linker variability generates fibers resembling the native structures. Proc Natl Acad Sci USA 1993; 90:9021–5.
Horowitz RA, Agard DA, Sedat JW, Woodcock CL . The three-dimensional architecture of chro-matin in situ: Electron tomography reveals fibers composed of a continuously variable Zig-Zagnucleosomal ribbon. J Cell Biol 1994; 125:1–10.
Acknowledgements
We are very grateful to Prof. WANG Ya-Hui for helpful suggestions and for critical reading of the manuscript. We also thank GAO Qi-Rong for excellent assistance in preparing the electron microscopic pictures. This project was supported by grants from National Natural Science Foundation, and from Chinese Academy of Sciences.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Qian, R., Liu, Z., Zhou, M. et al. Visualization of chromatin folding patterns in chicken erythrocytes by atomic force microscopy (AFM). Cell Res 7, 143–150 (1997). https://doi.org/10.1038/cr.1997.15
Received:
Revised:
Accepted:
Issue date:
DOI: https://doi.org/10.1038/cr.1997.15