Abstract
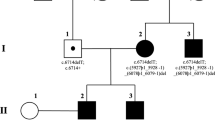
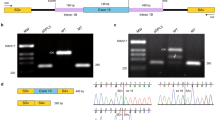
Fast and efficient high-throughput techniques are essential for the molecular diagnosis of highly heterogeneous hereditary diseases, such as retinitis pigmentosa (RP). We had previously approached RP genetic testing by devising a chip based on co-segregation analysis for the autosomal recessive forms. In this study, we aimed to design a diagnostic tool for all the known genes (40 up to now) responsible for the autosomal dominant and recessive RP and Leber congenital amaurosis (LCA). This new chip analyzes 240 single nucleotide polymorphisms (SNPs) (6 per gene) on a high-throughput genotyping platform (SNPlex, Applied Biosystems), and genetic diagnosis is based on the co-segregation analysis of SNP haplotypes in independent families. In a single genotyping step, the number of RP candidates to be screened for mutations is considerably reduced, and in the most informative families, all the candidates are ruled out at once. In a panel of RP Spanish pedigrees, the disease chip became a crucial tool for selecting those suitable for genome-wide RP gene search, and saved the burdensome direct mutational screening of every known RP gene. In a large adRP family, the chip allowed ruling out of all but the causative gene, and identification of an unreported null mutation (E181X) in PRPF31. Finally, on the basis of the conservation of the SNP haplotype linked to this pathogenic variant, we propose that the E181X mutation spread through a cohort of geographically isolated families by a founder effect.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Rivolta C, Sharon D, DeAngelis MM, Dryja TP : Retinitis pigmentosa and allied diseases: numerous diseases, genes, and inheritance patterns. Hum Mol Genet 2002; 11: 1219–1227.
Hims MM, Diager SP, Inglehearn CF : Retinitis pigmentosa: genes, proteins and prospects. Dev Ophthalmol 2003; 37: 109–125.
Pomares E, Marfany G, Brion MJ, Carracedo A, Gonzalez-Duarte R : Novel high-throughput SNP genotyping cosegregation analysis for genetic diagnosis of autosomal recessive retinitis pigmentosa and Leber congenital amaurosis. Hum Mutat 2007; 28: 511–516.
Dryja TP, McGee TL, Reichel E et al: A point mutation of the rhodopsin gene in one form of retinitis pigmentosa. Nature 1990; 343: 364–366.
Farrar GJ, McWilliam P, Bradley DG et al: Autosomal dominant retinitis pigmentosa: Linkage to rhodopsin and evidence for genetic heterogeneity. Genomics 1990; 8: 35–40.
Rosenfeld PJ, Cowley GS, McGee TL, Sandberg MA, Berson EL, Dryja TP : A null mutation in the rhodopsin gene causes rod photoreceptor dysfunction and autosomal recessive retinitis pigmentosa. Nat Genet 1992; 1: 209–213.
Bessant DA, Payne AM, Mitton KP et al: A mutation in NRL is associated with autosomal dominant retinitis pigmentosa. Nat Genet 1999; 21: 355–356.
Nishiguchi KM, Friedman JS, Sandberg MA, Swaroop A, Berson EL, Dryja TP : Recessive NRL mutations in patients with clumped pigmentary retinal degeneration and relative preservation of blue cone function. Proc Natl Acad Sci USA 2004; 101: 17819–17824.
Celesia GG, Bodis-Wollner I, Chatrian GE, Harding GF, Sokol S, Spekreijse H : Recommended standards for electroretinograms and visual evoked potentials. Report of an IFCN committee. Electroencephalogr Clin Neurophysiol 1993; 87: 421–436.
Vithana EN, Abu-Safieh L, Pelosini L et al: Expression of PRPF31 mRNA in patients with autosomal dominant retinitis pigmentosa: a molecular clue for incomplete penetrance? Invest Ophthalmol Vis Sci 2003; 44: 4204–4209.
Sato M, Nakazawa M, Usui T, Tanimoto N, Abe H, Ohguro H : Mutations in the gene coding for guanylate cyclase-activating protein 2 (GUCA1B gene) in patients with autosomal dominant retinal dystrophies. Graefes Arch Clin Exp Ophthalmol 2005; 243: 235–242.
Rivolta C, McGee TL, Rio Frio T, Jensen RV, Berson EL, Dryja TP : Variation in retinitis pigmentosa-11 (PRPF31 or RP11) gene expression between symptomatic and asymptomatic patients with dominant RP11 mutations. Hum Mutat 2006; 27: 644–653.
Rio Frio T, Wade NM, Ransijn A, Berson EL, Beckmann JS, Rivolta C : Premature termination codons in PRPF31 cause retinitis pigmentosa via haploinsufficiency due to nonsense-mediated mRNA decay. J Clin Invest 2008; 118: 1519–1531.
Abu-Safieh L, Vithana EN, Mantel I et al: A large deletion in the adRP gene PRPF31: evidence that haploinsufficiency is the cause of disease. Mol Vis 2006; 12: 384–388.
Rio Frio T, Civic N, Ransijn A, Beckmann JS, Rivolta C : Two trans-acting eQTLs modulate the penetrance of PRPF31 mutations. Hum Mol Genet 2008; 17: 3154–3165.
Acknowledgements
We are grateful to the members of all the families for their participation in this study. We are obliged to A Mayor for sample collection, helpful discussions and constant support to our research. EP was a recipient of an FPI fellowship from the Spanish Ministry of Education and Science (MEC), and is now under contract by CIBERER. MR was a recipient of a scholarship from Bidons Egara and is now a recipient of an FPU fellowship from MEC. PM and JP are under contract by CIBERER. This study was funded by Fundaluce (2004), Grant BFU2006-04562 (Ministerio de Educación y Ciencia) and CIBERER INTRA/07–08/718.1 to RG-D. This study was approved by the Bioethics Committee of the University of Barcelona (Barcelona, Spain).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pomares, E., Riera, M., Permanyer, J. et al. Comprehensive SNP-chip for retinitis pigmentosa-Leber congenital amaurosis diagnosis: new mutations and detection of mutational founder effects. Eur J Hum Genet 18, 118–124 (2010). https://doi.org/10.1038/ejhg.2009.114
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/ejhg.2009.114
Keywords
This article is cited by
-
Molecular characterization of AIPL1 gene region in the Iranian population: application of novel informative haplotypes and detection of mutational founder effect
Genes & Genomics (2017)
-
SAQC: SNP Array Quality Control
BMC Bioinformatics (2011)