Abstract
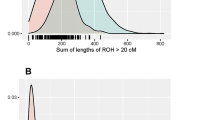
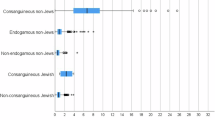
Consanguineous marriages have been widely practiced in several global communities with varying rates depending on religion, culture, and geography. In consanguineous marriages, parents pass to their children autozygous segments known as homozygous by descent segments. In this study, single-nucleotide polymorphisms were analyzed in 165 unrelated Lebanese people from Greek Orthodox, Maronite, Shiite and Sunni communities. Runs of homozygosity, total inbreeding levels, remote consanguinity, and population admixture and structure were estimated. The inbreeding coefficient value was estimated to be 1.61% in offspring of unrelated parents over three generations and 8.33% in offspring of first cousins. From these values, remote consanguinity values, resulting from genetic drift or recurrent consanguineous unions, were estimated in offspring of unrelated and first-cousin parents to be 0.61 and 1.2%, respectively. This remote consanguinity value suggests that for any unrelated marriages in Lebanon, the mates could be related as third cousins or as second cousins once removed. Under the assumption that 25% of marriages occur between first cousins, the mean inbreeding value of 2.3% may explain the increased incidence of recessive disease in offspring. Our analysis reveals a common ancestral population in the four Lebanese communities we studied.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
Change history
17 September 2015
This paper has been corrected since online publication and a corrigendum also appears in this issue
References
Bittles AH, Black ML : Evolution in health and medicine Sackler colloquium: consanguinity, human evolution, and complex diseases. Proc Natl Acad Sci USA 2010; 107: 1779–1786.
Alkuraya FS : Autozygome decoded. Genet Med 2010; 12: 765–771.
Hamamy H, Antonarakis SE, Cavalli-Sforza LL et al: Consanguineous marriages, pearls and perils: Geneva International Consanguinity Workshop Report. Genet Med Off J Am Coll Med Genet 2011; 13: 841–847.
Cavalli-Sforza LL, Bodmer WF The Genetics of Human Populations 1971.
Leutenegger AL, Prum B, Génin E et al: Estimation of the inbreeding coefficient through use of genomic data. Am J Hum Genet 2003; 73: 516–523.
Leutenegger AL, Sahbatou M, Gazal S et al: Consanguinity around the world: what do the genomic data of the HGDP-CEPH diversity panel tell us? Eur J Hum Genet 2011; 19: 583–587.
Browning SR, Browning BL : High-resolution detection of identity by descent in unrelated individuals. Am J Hum Genet 2010; 86: 526–539.
Haber M, Gauguier D, Youhanna S et al: Genome-wide diversity in the levant reveals recent structuring by culture. PLoS Genet 2013; 9: e1003316.
Miller SA, Dykes DD, Polesky HF : A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 1988; 16: 1215.
International HapMap 3 Consortium. et al.. Integrating common and rare genetic variation in diverse human populations. Nature 2010; 467: 52–58.
Cann HM, de Toma C, Cazes L et al: A human genome diversity cell line panel. Science 2002; 296: 261–262.
Li JZ, Absher DM, Tang H et al: Worldwide human relationships inferred from genome-wide patterns of variation. Science 2008; 319: 1100–1104.
Rosenberg NA : Standardized subsets of the HGDP-CEPH Human Genome Diversity Cell Line Panel, accounting for atypical and duplicated samples and pairs of close relatives. Ann Hum Genet 2006; 70: 841–847.
International HapMap Consortium International HapMap Consortium, Frazer KA International HapMap Consortium, Ballinger DG International HapMap Consortium, Cox DR et al: A second generation human haplotype map of over 3.1 million SNPs. Nature 2007; 449: 851–861.
Bødker JS, Gyrup C, Johansen P et al: Performance comparison of Affymetrix SNP6.0 and cytogenetic 2.7M whole-genome microarrays in complex cancer samples. Cytogenet Genome Res 2013; 139: 80–87.
Kirin M, McQuillan R, Franklin CS et al: Genomic runs of homozygosity record population history and consanguinity. PLoS One 2010; 5: e13996.
Kearney HM, Kearney JB, Conlin LK : Diagnostic implications of excessive homozygosity detected by SNP-based microarrays: consanguinity, uniparental disomy, and recessive single-gene mutations. Clin Lab Med 2011; 31: 595–613.
McQuillan R, Leutenegger AL, Abdel-Rahman R et al: Runs of homozygosity in European populations. Am J Hum Genet 2008; 83: 359–372.
Wright S. : Evolution and the Genetics of Populations; Vol. 2. The Theory of Gene Frequencies. Chicago: University of Chicago Press, 1968.
Patterson N, Price AL, Reich D : Population structure and eigenanalysis. PLoS Genet 2006; 2: 2074–2093.
Li LH, Ho SF, Chen CH et al: Long contiguous stretches of homozygosity in the human genome. Hum Mutat 2006; 27: 1115–1121.
Gibson J, Morton NE, Collins A : Extended tracts of homozygosity in outbred human populations. Hum Mol Genet 2006; 15: 789–795.
Khlat M : Consanguineous marriage and reproduction in Beirut, Lebanon. Am J Hum Genet 1988; 43: 188–196.
Khlat M : Consanguineous marriages in Beirut: time trends, spatial distribution. Soc Biol 1988; 35: 324–330.
Barbour B, Salameh P : Consanguinity in Lebanon: prevalence, distribution and determinants. J Biosoc Sci 2009; 41: 505–517.
Ten Kate LP, Teeuw M, Henneman L et al: Autosomal recessive disease in children of consanguineous parents: inferences from the proportion of compound heterozygotes. J Community Genet 2010; 1: 37–40.
Gialluisi A, Pippucci T, Anikster Y et al: Estimating the allele frequency of autosomal recessive disorders through mutational records and consanguinity: the Homozygosity Index (HI). Ann Hum Genet 2012; 76: 159–167.
Rosenberg NA, Pemberton TJ, Li JZ et al: Runs of homozygosity and parental relatedness. Genet Med 2013; 15: 753–754.
Rehder CW, David KL, Hirsch B et al: American College of Medical Genetics and Genomics: standards and guidelines for documenting suspected consanguinity as an incidental finding of genomic testing. Genet Med 2013; 15: 150–152.
Acknowledgements
We are grateful to the research council of Saint-Joseph University-Lebanon. This project is supported and funded by the Agence de la Biomedecine (France), Saint-Joseph University (Lebanon) and the French Ministry of Higher Education and Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Jalkh, N., Sahbatou, M., Chouery, E. et al. Genome-wide inbreeding estimation within Lebanese communities using SNP arrays. Eur J Hum Genet 23, 1364–1369 (2015). https://doi.org/10.1038/ejhg.2014.246
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/ejhg.2014.246
This article is cited by
-
People from Ibiza: an unexpected isolate in the Western Mediterranean
European Journal of Human Genetics (2019)
-
First molecular study in Lebanese patients with Cockayne syndrome and report of a novel mutation in ERCC8 gene
BMC Medical Genetics (2018)
-
Runs of homozygosity: windows into population history and trait architecture
Nature Reviews Genetics (2018)
-
Contribution of copy number variants (CNVs) to congenital, unexplained intellectual and developmental disabilities in Lebanese patients
Molecular Cytogenetics (2015)