Abstract
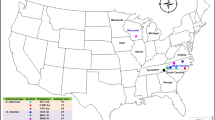
S-allele diversity in Solanum carolinense was surveyed in two natural populations, located in Tennessee and North Carolina, with a molecular assay to determine the genotype of individual plants. A total of 13 different S-alleles were identified and sequenced. There is high overlap between the two populations sampled, with 10 alleles shared in common, one allele found only in Tennessee, and two found only in North Carolina. The number of alleles in this species appears to be extremely low compared with other species with gametophytic self-incompatibility. Sequence comparisons show that most alleles are extremely different one from another in their primary sequence and a phylogenetic analysis indicates extensive trans-specific evolution of S-lineages. In addition, some alleles appear to be derived much more recently. The implications of these observations are discussed in the light of recent theoretical results on S-allele population diversity and persistence.
Similar content being viewed by others
Article PDF
References
Ai, Y, Singh, A, Coleman, C E, Ioerger, T R, Kheyrpour, A, and Kao, T-H. 1990. Self-incompatibility in Petunia inflata: isolation and characterization of cdnas encoding three 5-allele-associated proteins. Sex Plant Reprod, 3, 130–138.
Anderson, M A, Cornish, E C, Mau, S-L, Williams, E G, Hoggart, R, Atkinson, A, Bonig, I, Grego, B, Simpson, R, Roche, P J, Haley, J D, Penschow, J D, Niall, H D, Tregear, G W, Coghlan, J P, Crawford, R J, and Clarke, A E. 1986. Cloning of cdna for a stylar glycoprotein associated with expression of self-incompatibility in Nicotiana alata. Nature, 321, 38–44.
Brace, J, King G J and Ockendon, D J. 1993. Development of a method for the identification of S-alleles in Brassica oleracea based on digestion of PCR-amplified restriction endonucleases. Sex Plant Reprod, 6, 133–138.
Brace, J, King G J and Ockendon, D J. 1994. A molecular approach to the identification of S-alleles in Brassica oleracea. Sex Plant Reprod, 7, 203–208.
Campbell, J M, and Lawrence, M J. 1981. The population genetics of the self-incompatibility polymorphism in Papaver rhoeas. II. The number and frequency of S-alleles in a natural population (R106). Heredity, 46, 81–90.
Clark, A G. 1993. Evolutionary inferences from molecular characterization of self-incompatibility alleles. In: Takahata, N. and Clark, A. G. (eds) Mechanisms of Molecular Evolution, pp. 79–108. Sinauer Associates, Sunderland, MA.
Clark, A G, and Kao, T-H. 1991. Excess nonsynonymous substitution at shared polymorphic sites among self-incompatibility alleles of Solanaceae. Proc Natl Acad Sci USA, 88, 9823–9827.
Clarke, A E, and Newbigin, E. 1993. Molecular aspects of self-incompatibility in flowering plants. Ann Rev Genet, 27, 257–279.
Emerson, S. 1938. The genetics of self-incompatibility in Oenothera organensis. Genetics, 23, 190–202.
Emerson, S. 1939. A preliminary survey of the Oenothera organensis population. Genetics, 24, 524–537.
Foote, H C C, Ride, J P, Franklin-Tong, V E, Walker, E A, Lawrence, M J, and Franklin, F C. 1994. Cloning and expression of a novel class of self-incompatibility (S) gene from Papaver rhoeas L. Proc Natl Acad Sci USA, 91, 2265–2269.
Franklin-Tong, V E, and Franklin, F C H. 1993. Gametophytic self-incompatibility: contrasting mechanisms for Nicotiana and Papaver. Trends Cell Biol, 3, 340–345.
Hinata, K, Watanabe, M, Toriyama, K, and Isogai, A. 1993. A review of recent studies on homomorphic self-incompatibility. Int Rev Cytol, 143, 257–296.
Ioerger, T R, Clark, A G, and Kao, T-H. 1990. Polymorphism at the self-incompatibility locus in Solanaceae predates speciation. Proc Natl Acad Sci USA, 87, 9732–9735.
Ioerger, T R, Gohlke, J R, Xu, B, and Kao, T-H. 1991. Primary structural features of the self-incompatibility protein in Solanaceae. Sex Plant Reprod, 4, 81–87.
Jost, W, Bak, H, Glund, K, Terpstra, P, and Beintema, J J. 1991. Amino acid sequence of an extracellular, phosphate-starvation-induced ribonuclease from cultured tomato (Lycopersicon esculentum) cells. Eur J Biochem, 198, 1–6.
Kaufmann, H, Salamini, F, and Thompson, R D. 1991. Sequence variability and gene structure at the self-incompatibility locus of Solanum tuberosum. Mol Gen Genet, 226, 457–466.
Kheyrpour, A, Bintrim, S B, Ioerger, T R, Remy, R, Hammond, S A, and Kao, T-H. 1990. Sequence diversity of pistil 5-proteins associated with gametophytic self-incompatibility in Nicotiana alata. Sex Plant Reprod, 3, 88–97.
Kirch, H H, Uhrig, H, Lottspeich, F, Salamini, F, and Thompson, R D. 1989. Characterization of proteins associated with self-incompatibility in Solanum tuberosum. Theor Appl Genet, 78, 581–588.
Kumar, S, Tamura, K, and Nei, M. 1993. MEGA: Molecular Evolutionary Genetics Analysis, version 1.02. The Pennsylvania State University, University Park, PA.
Lane, M D, and Lawrence, M J. 1993. The population genetics of the self-incompatibility polymorphism in Papaver rhoeas. VII. The number of S-alleles in the species. Heredity, 71, 596–602.
Lawrence, M J. 1975. The genetics of self-incompatibility in Papaver rhoeas. Proc R Soc Lond B, 188, 275–285.
Lawrence, M J, and Franklin-Tong, V E. 1994. The population genetics of the self-incompatibility polymorphism in Papaver rhoeas. IX. Evidence of an extra effect of selection acting on the 5-locus. Heredity, 72, 353–364.
Lawrence, M J, Lane, M D, O'Donnell, S, and Franklin-Tong, V E. 1993. The population genetics of the self-incompatibility polymorphism in Papaver rhoeas. V. Cross-classification of the S-alleles of samples from three natural populations. Heredity, 71, 581–590.
Lawrence, M J, and O'Donnell, S. 1981. The population genetics of the self-incompatibility polymorphism in Papaver rhoeas. III. The number and frequency of S-alleles in two further natural populations (R102 and R104). Heredity, 47, 53–61.
Lawrence, M J, O'Donnell, S, Lane, M D, and Marshall, D F. 1994. The population genetics of the self-incompatibility polymorphism in Papaver rhoeas. VIII. Sampling effects as a possible cause of unequal allele frequencies. Heredity, 72, 345–352.
Lee, H-S, Huang, S, and Kao, T-H. 1994. S proteins control rejection of incompatible pollen in Petunia inflata. Nature, 367, 560–563.
Lee, H-S, Singh, A, and Kao, T-H. 1992. RNase X2, a pistil-specific ribonuclease from Petunia inflata, shares sequence similarity with solanaceous S proteins. Plant Mol Biol, 20, 1131–1141.
Levin, D A. 1993. S-gene polymorphism in Phlox drum-mondii. Heredity, 71, 193–198.
Li, X, Nield, J, Hayman, D, and Langridge, P. 1994. Cloning a putative self-incompatibility gene from the pollen of the grass Phalaris coerulescens. Plant Cell, 6, 1923–1932.
Mantel, N. 1974. Approaches to a health research occupancy problem. Biometrics, 30, 355–362.
Nagylaki, T. 1975. The deterministic behavior of self-incompatibility alleles. Genetics, 79, 545–550.
Nasrallah, J B, Kao, T-H, Chen, C-H, Goldberg, M L, and Nasrallah, M E. 1987. Amino-acid sequence of glycoproteins encoded by three alleles of the 5 locus of Brassica oleracea. Nature, 326, 617–619.
O'Donnell, S, Lane, M D, and Lawrence, M J. 1993. The population genetics of the self-incompatibility polymorphism in Papaver rhoeas. VI. Estimation of the overlap between the allelic complements of a pair of populations. Heredity, 71, 591–595.
O'Donnell, S, and Lawrence, M J. 1984. The population genetics of the self-incompatibility polymorphism in Papaver rhoeas. IV. The estimation of the number of alleles in a population. Heredity, 53, 495–507.
Ockendon, D J. 1974. Distribution of self-incompatibility alleles and breeding structure of open-pollinated cultivars of Brussels sprouts. Heredity, 33, 159–171.
Paxman, G J. 1963. The maximum likelihood estimation of the number of self-sterility alleles in a population. Genetics, 48, 1029–1032.
Royo, J, Kowyama, Y, and Clarke, A E. 1994. Cloning and nucleotide sequence of two 5-RNases from Lyco-persicon peruvianum (L.) Mill. Plant Physiol, 105, 751–752.
Saba-El-Leil, M K, Rivard, S, Morse, D, and Cappado-Cia, M. 1994. The 511 and 513 self-incompatibility alleles in Solanum chacoense Bitt. are remarkably similar. Plant Mol Biol, 24, 571–583.
Saitou, N, and Nei, M. 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol, 4, 406–425.
Taylor, C B, and Green, P J. 1991. Genes with homology to fungal and 5-gene RNases are expressed in Arabi-dopsis thaliana. PI Physiol, 96, 980–984.
Uyenoyama, M K. 1991. On the evolution of genetic incompatibility systems. VI. A three-locus modifier model for the origin of gametophytic self-incompatibility. Genetics, 128, 453–469.
Uyenoyama, M K. 1995. A generalized least squares estimate for the origin of sporophytic self-incompatibility. Genetics, 139, 975–992.
Vekemans, X, and Slatkin, M. 1994. Gene and allelic genealogies at a gametophytic self-incompatibility locus. Genetics, 137, 1157–1165.
Wright, S. 1939. The distribution of self-sterility alleles in populations. Genetics, 24, 538–552.
Wright, S. 1960. On the number of self-incompatibility alleles maintained in equilibrium by a given mutation rate in a population of a given size: a re-examination. Biometrics, 16, 61–85.
Wright, S. 1965. The distribution of self-incompatibility alleles in populations. Evolution, 18, 609–619.
Xu, B, Mu, J, Nevins, D L, Grun, P, and Kao, T-H. 1990. Cloning and sequencing of cdnas encoding two self-incompatibility associated proteins in Solanum chacoense. Mol Gen Genet, 224, 341–346.
Yokoyama, S, and Hetherington, L E. 1982. The expected number of self-incompatibility alleles in finite plant populations. Heredity, 48, 299–303.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Richman, A., Kao, TH., Schaeffer, S. et al. S-allele sequence diversity in natural populations of Solanum carolinense (Horsenettle). Heredity 75, 405–415 (1995). https://doi.org/10.1038/hdy.1995.153
Received:
Issue date:
DOI: https://doi.org/10.1038/hdy.1995.153
Keywords
This article is cited by
-
Nutrient stress can have opposite effects on the ability of plants to tolerate foliar herbivory and floral herbivory
Oecologia (2023)
-
Simple Sequence Repeat-Based Genetic Diversity and Analysis of Molecular Variance among on-Farm Native Potato Landraces from the Influence Zone of Camisea Gas Project, Northern Ayacucho, Peru
American Journal of Potato Research (2020)
-
Herbivory and inbreeding affect growth, reproduction, and resistance in the rhizomatous offshoots of Solanum carolinense (Solanaceae)
Evolutionary Ecology (2019)
-
Functional gametophytic self-incompatibility in a peripheral population of Solanum peruvianum (Solanaceae)
Heredity (2011)
-
Molecular and genetic characterization of novel S-RNases from a natural population of Nicotiana alata
Plant Cell Reports (2010)