Abstract
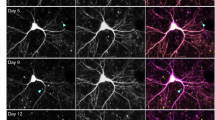
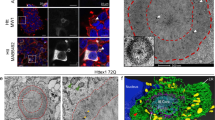
Huntington's disease is caused by an abnormal polyglutamine expansion within the protein huntingtin and is characterized by microscopic inclusion bodies of aggregated huntingtin and by the death of selected types of neuron. Whether inclusion bodies are pathogenic, incidental or a beneficial coping response is controversial. To resolve this issue we have developed an automated microscope that returns to precisely the same neuron after arbitrary intervals, even after cells have been removed from the microscope stage. Here we show, by survival analysis, that neurons die in a time-independent fashion but one that is dependent on mutant huntingtin dose and polyglutamine expansion; many neurons die without forming an inclusion body. Rather, the amount of diffuse intracellular huntingtin predicts whether and when inclusion body formation or death will occur. Surprisingly, inclusion body formation predicts improved survival and leads to decreased levels of mutant huntingtin elsewhere in a neuron. Thus, inclusion body formation can function as a coping response to toxic mutant huntingtin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Davies, S. W. et al. Formation of neuronal intranuclear inclusions underlies the neurological dysfunction in mice transgenic for the HD mutation. Cell 90, 537–548 (1997)
Becher, M. W. et al. Intranuclear neuronal inclusions in Huntington's disease and dentatorubral and pallidoluysian atrophy—correlation between the density of inclusions and IT-15 CAG triplet repeat length. Neurobiol. Dis. 4, 387–397 (1998)
DiFiglia, M. et al. Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science 277, 1990–1993 (1997)
Ordway, J. M. et al. Ectopically expressed CAG repeats cause intranuclear inclusions and a progressive late onset neurological phenotype in the mouse. Cell 91, 753–763 (1997)
Saudou, F., Finkbeiner, S., Devys, D. & Greenberg, M. E. Huntingtin acts in the nucleus to induce apoptosis, but death does not correlate with the formation of intranuclear inclusions. Cell 95, 55–66 (1998)
Klement, I. A. et al. Ataxin-1 nuclear localization and aggregation: Role in polyglutamine-induced disease in SCA1 transgenic mice. Cell 95, 41–53 (1998)
Cummings, C. J. et al. Mutation of the E6–AP ubiquitin ligase reduces nuclear inclusion frequency while accelerating polyglutamine-induced pathology in SCA1 mice. Neuron 24, 879–892 (1999)
Taylor, J. P. et al. Aggresomes protect cells by enhancing the degradation of toxic polyglutamine-containing protein. Hum. Mol. Genet. 12, 749–757 (2003)
Shimohata, T. et al. Expanded polyglutamine stretches form an ‘aggresome’. Neurosci. Lett. 323, 215–218 (2002)
Sisodia, S. S. Nuclear inclusions in glutamine repeat disorders: Are they pernicious, coincidental or beneficial? Cell 95, 1–4 (1998)
Bence, N. F., Sampat, R. M. & Kopito, R. R. Impairment of the ubiquitin–proteasome system by protein aggregation. Science 292, 1552–1555 (2001)
Ross, C. A. Intranuclear neuronal inclusions: A common pathogenic mechanism for glutamine-repeat neurodegenerative diseases. Neuron 19, 1147–1150 (1997)
Poirier, M. A. et al. Huntingtin spheroids and protofibrils as precursors in polyglutamine fibrilization. J. Biol. Chem. 277, 41032–41037 (2002)
Chen, S., Berthelier, V., Yang, W. & Wetzel, R. Polyglutamine aggregation behavior in vitro supports a recruitment mechanism of cytotoxicity. J. Mol. Biol. 311, 173–182 (2001)
Wyttenbach, A. et al. Effects of heat shock, heat shock protein 40 (HDJ-2), and proteasome inhibition on protein aggregation in cellular models of Huntington's disease. Proc. Natl Acad. Sci. USA 97, 2898–2903 (2000)
Muchowski, P. J., Ning, K., D'Souza-Schorey, C. & Fields, S. Requirement of an intact microtubule cytoskeleton for aggregation and inclusion body formation by a mutant huntingtin fragment. Proc. Natl Acad. Sci. USA 99, 727–732 (2002)
Perutz, M. F. & Windle, A. H. Cause of neural death in neurodegenerative disease attributable to expansion of glutamine repeats. Nature 412, 143–144 (2001)
Arrasate, M., Brooks, L., Chang, P., Mitra, S. & Finkbeiner, S. Longitudinal analysis to identify pathogenic factors in a striatal model of Huntington's disease. Soc. Neurosci. Abstr. 29, 209.8 (2003)
Collett, D. Modeling Survival Data in Medical Research (Chapman & Hall, London, 1994)
Therneau, T. M. & Grambsch, P. M. Modeling Survival Data: Extending the Cox Model (Springer, New York, 2000)
Kazantsev, A., Preisinger, E., Dranovsky, A., Goldgaber, D. & Housman, D. Insoluble detergent-resistant aggregates form between pathological and nonpathological lengths of polyglutamine in mammalian cells. Proc. Natl Acad. Sci. USA 96, 11404–11409 (1999)
Goldberg, Y. P. et al. Cleavage of huntingtin by apopain, a proapoptotic cysteine protease, is modulated by the polyglutamine tract. Nature Genet. 13, 442–449 (1996)
Scherzinger, E. et al. Huntingtin-encoded polyglutamine expansions form amyloid-like protein aggregates in vitro and in vivo. Cell 90, 549–558 (1997)
Wellington, C. L. et al. Caspase cleavage of gene products associated with triplet expansion disorders generates truncated fragments containing the polyglutamine. J. Biol. Chem. 273, 9158–9167 (1998)
Kim, Y. J. et al. Caspase 3-cleaved N-terminal fragments of wild-type and mutant huntingtin are present in normal and Huntington's disease brains, associate with membranes, and undergo calpain-dependent proteolysis. Proc. Natl Acad. Sci. USA 98, 12784–12789 (2001)
Mende-Mueller, L. M., Toneff, T., Hwang, S. R., Chesselet, M. F. & Hook, V. Y. H. Tissue-specific proteolysis of huntingtin (htt) in human brain: Evidence of enhanced levels of N- and C-terminal htt fragments in Huntington's disease striatum. J. Neurosci. 21, 1830–1837 (2001)
Mangiarini, L. et al. Exon 1 of the HD gene with an expanded CAG repeat is sufficient to cause a progressive neurological phenotype in transgenic mice. Cell 87, 493–506 (1996)
Campbell, R. E. et al. A monomeric red fluorescent protein. Proc. Natl Acad. Sci. USA 99, 7877–7882 (2002)
Strebel, A., Harr, T., Bachmann, F., Wernli, M. & Erb, P. Green fluorescent protein as a novel tool to measure apoptosis and necrosis. Cytometry 43, 126–133 (2001)
MacDonald, M. E. in Trinucleotide Diseases and Instability (ed. Oostra, B. A.) 47–75 (Springer, Berlin, 1998)
Clarke, G. et al. A one-hit model of cell death in inherited neuronal degenerations. Nature 406, 195–199 (2000)
Reiner, A. et al. Differential loss of striatal projection neurons in Huntington disease. Proc. Natl Acad. Sci. USA 85, 5733–5737 (1988)
Richfield, E. K., Maguire-Zeiss, K. A., Vonkeman, H. E. & Voorn, P. Preferential loss of preproenkephalin versus preprotachykinin neurons from the striatum of Huntington's disease patients. Ann. Neurol. 38, 852–861 (1995)
Rajan, R. S., Illing, M. E., Bence, N. F. & Kopito, R. R. Specificity in intracellular protein aggregation and inclusion body formation. Proc. Natl Acad. Sci. USA 98, 13060–13065 (2001)
Moulder, K. L., Onodera, O., Burke, J. R., Strittmatter, W. J. & Johnson, E. M. Jr Generation of neuronal intranuclear inclusions by polyglutamine-GFP: Analysis of inclusion clearance and toxicity as a function of polyglutamine length. J. Neurosci. 19, 705–715 (1999)
Gutekunst, C. A. et al. Nuclear and neuropil aggregates in Huntington's disease: Relationship to neuropathology. J. Neurosci. 19, 2522–2534 (1999)
Preisinger, E., Jordan, B. M., Kazantsev, A. & Housman, D. Evidence for a recruitment and sequestration mechanism in Huntington's disease. Phil. Trans. R. Soc. Lond. B 354, 1029–1034 (1999)
Kuemmerle, S. et al. Huntingtin aggregates may not predict neuronal death in Huntington's disease. Ann. Neurol. 46, 842–849 (1999)
Kim, M. et al. Mutant huntingtin expression in clonal striatal cells: Dissociation of inclusion formation and neuronal survival by caspase inhibition. J. Neurosci. 19, 964–973 (1999)
Hack, N. J. et al. Green fluorescent protein as a quantitative tool. J. Neurosci. Methods 95, 177–184 (2000)
Yoo, S.-Y. et al. SCA7 knockin mice model human SCA7 and reveal gradual accumulation of mutant ataxin-7 in neurons and abnormalities in short-term plasticity. Neuron 37, 383–401 (2003)
Persichetti, F. et al. Differential expression of normal and mutant Huntington's disease gene alleles. Neurobiol. Dis. 3, 183–190 (1996)
Kegel, K. B. et al. Huntingtin is present in the nucleus, interacts with the transcriptional corepressor C-terminal binding protein, and represses transcription. J. Biol. Chem. 277, 7466–7476 (2002)
Peters, M. F. et al. Nuclear targeting of mutant huntingtin increases toxicity. Mol. Cell. Neurosci. 14, 121–128 (1999)
Ravikumar, B. et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nature Genet. 36, 585–595 (2004)
Sánchez, I., Mahlke, C. & Yuan, J. Pivotal role of oligomerization in expanded polyglutamine neurodegenerative disorders. Nature 421, 373–379 (2003)
Apostol, B. L. et al. A cell-based assay for aggregation inhibitors as therapeutics of polyglutamine-repeat disease and validation in Drosophila. Proc. Natl Acad. Sci. USA 100, 5950–5955 (2003)
Tanaka, M. et al. Trehalose alleviates polyglutamine-mediated pathology in a mouse model of Huntington disease. Nature Med. 10, 148–154 (2004)
Finkbeiner, S. et al. CREB: A major mediator of neuronal neurotrophin responses. Neuron 19, 1031–1047 (1997)
Aiken, C. T., Tobin, A. J. & Schweitzer, E. S. A cell-based screen for drugs to treat Huntington's disease. Neurobiol. Dis. 16, 546–555 (2004)
Acknowledgements
We thank A. Kazantzev, D. Housman and the Hereditary Disease Foundation for pcDNA3.1-Htt (Q25, Q47, Q72, Q103)-GFP plasmids; R. Truant for the PCR template (GFP–109-17Q-βgal) used to create pGW1-Httex1-Q17-GFP; R. Tsien for mRFP cDNA; D. Bredesen, S. Prusiner, S. Lindquist, R. Edwards, A. Tobin, E. Signer, C. Johnson, P. Muchowski and members of the Finkbeiner laboratory for useful discussions; S. Ordway and G. Howard for editorial assistance; K. Nelson for administrative assistance; and E. Oliver and D. Murphy for their interest and support. Primary support for this work was provided by the National Institute of Neurological Disease and Stroke (S.F). Additional support was provided by the National Institute of Aging and the J. David Gladstone Institutes (S.F.). M.A. is a MECD–Fulbright Fellow and is supported by the Hillblom Foundation. S.M. is supported by the NIH–NIGMS UCSF Medical Scientist Training Program and a fellowship from the UCSF Hillblom Center for the Biology of Aging. E.S. is supported by the National Institute of Neurological Disease and Stroke, the Hereditary Disease Foundation, and the High Q Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure 1 (download JPG )
Fluorescence of mRFP provides a measure of neuronal morphology and viability that is independent of httex1-GFP. (JPG 129 kb)
Supplementary Figure 2 (download JPG )
Abrupt loss of fluorescence from transfected mRFP is a sensitive and specific assay of neuronal death. (JPG 56 kb)
Supplementary Figure 3 (download JPG )
Httex1-Q47–GFP increases the risk of death significantly compared with httex1-Q17–GFP. (JPG 29 kb)
Supplementary Figure 4 (download JPG )
Single-neuron levels of GFP estimated by imaging correlate well with levels measured by immunocytochemistry. (JPG 21 kb)
Supplementary Figure 5 (download JPG )
Neurons with similar levels of httex1-Q47-GFP that form IBs on the second day survive better than neurons that do not. (JPG 31 kb)
Supplementary Figure 6 (download JPG )
IB formation is associated with reduced death risk and increased survival among neurons transfected with httex1-Q47-GFP that are alive beginning on the sixth day. (JPG 24 kb)
Supplementary Figure 7 (download JPG )
Neurons with IBs are viable as assessed by annexinV staining. (JPG 143 kb)
Supplementary Figure 8 (download JPG )
IB formation is associated with reduced death risk and increased survival among stably transfected PC12 cells that were induced to express httex1-Q103-GFP and were alive beginning on the third day. (JPG 33 kb)
Rights and permissions
About this article
Cite this article
Arrasate, M., Mitra, S., Schweitzer, E. et al. Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 431, 805–810 (2004). https://doi.org/10.1038/nature02998
Received:
Accepted:
Issue date:
DOI: https://doi.org/10.1038/nature02998
This article is cited by
-
Elucidation of Site-Specific Ubiquitination on Chaperones in Response to Mutant Huntingtin
Cellular and Molecular Neurobiology (2024)
-
Comparison of Huntington’s disease phenotype progression in male and female heterozygous FDNQ175 mice
Molecular Brain (2023)
-
Enhanced liquidity of p62 droplets mediated by Smurf1 links Nrf2 activation and autophagy
Cell & Bioscience (2023)
-
Beneficial behavioral effects of chronic cerebral dopamine neurotrophic factor (CDNF) infusion in the N171-82Q transgenic model of Huntington’s disease
Scientific Reports (2023)
-
Nuclear and cytoplasmic spatial protein quality control is coordinated by nuclear–vacuolar junctions and perinuclear ESCRT
Nature Cell Biology (2023)