Abstract
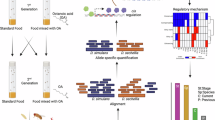
We developed a method for systematically comparing gene expression patterns across organisms using genome-wide comparative analysis of DNA microarray experiments. We identified analogous gene expression programs comprising shared patterns of regulation across orthologous genes. Biological features of these patterns could be identified as highly conserved subpatterns that correspond to Gene Ontology categories. Here, we demonstrate these methods by analyzing a specific biological process, aging, and show that similar analysis can be applied to a range of biological processes. We found that two highly diverged animals, the nematode Caenorhabditis elegans and the fruit fly Drosophila melanogaster, implement a shared adult-onset expression program of genes involved in mitochondrial metabolism, DNA repair, catabolism, peptidolysis and cellular transport. Most of these changes were implemented early in adulthood. Using this approach to search databases of gene expression data, we found conserved transcriptional signatures in larval development, embryogenesis, gametogenesis and mRNA degradation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
DeRisi, J.L., Iyer, V.R. & Brown, P.O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686 (1997).
Lipshutz, R.J., Fodor, S.P., Gingeras, T.R. & Lockhart, D.J. High density synthetic oligonucleotide arrays. Nat. Genet. 21, 20–24 (1999).
Chung, C.H., Bernard, P.S. & Perou, C.M. Molecular portraits and the family tree of cancer. Nat. Genet. 32 Suppl, 533–540 (2002).
Ramaswamy, S., Ross, K.N., Lander, E.S. & Golub, T.R. A molecular signature of metastasis in primary solid tumors. Nat. Genet. 33, 49–54 (2003).
Hughes, T.R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000).
Mootha, V.K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Wang, D.Y., Kumar, S. & Hedges, S.B. Divergence time estimates for the early history of animal phyla and the origin of plants, animals and fungi. Proc. R. Soc. Lond. B Biol. Sci. 266, 163–171 (1999).
Pletcher, S.D. et al. Genome-wide transcript profiles in aging and calorically restricted Drosophila melanogaster. Curr. Biol. 12, 712–723 (2002).
Hill, A.A., Hunter, C.P., Tsung, B.T., Tucker-Kellogg, G. & Brown, E.L. Genomic analysis of gene expression in C. elegans. Science 290, 809–812 (2000).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Lee, C.K., Klopp, R.G., Weindruch, R. & Prolla, T.A. Gene expression profile of aging and its retardation by caloric restriction. Science 285, 1390–1393 (1999).
Kayo, T., Allison, D.B., Weindruch, R. & Prolla, T.A. Influences of aging and caloric restriction on the transcriptional profile of skeletal muscle from rhesus monkeys. Proc. Natl. Acad. Sci. USA 98, 5093–5098 (2001).
Lund, J. et al. Transcriptional profile of aging in C. elegans. Curr. Biol. 12, 1566–1573 (2002).
Zou, S., Meadows, S., Sharp, L., Jan, L.Y. & Jan, Y.N. Genome-wide study of aging and oxidative stress response in Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 97, 13726–13731 (2000).
Sherlock, G. et al. The Stanford Microarray Database. Nucleic Acids Res. 29, 152–155 (2001).
Finkel, T. & Holbrook, N.J. Oxidants, oxidative stress and the biology of ageing. Nature 408, 239–247 (2000).
Lithgow, G.J., White, T.M., Melov, S. & Johnson, T.E. Thermotolerance and extended life-span conferred by single-gene mutations and induced by thermal stress. Proc. Natl. Acad. Sci. USA 92, 7540–7544 (1995).
Murphy, C.T. et al. Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans. Nature 424, 277–283 (2003).
Kimura, K.D., Tissenbaum, H.A., Liu, Y. & Ruvkun, G. daf-2, an insulin receptor-like gene that regulates longevity and diapause in Caenorhabditis elegans. Science 277, 942–946 (1997).
Kenyon, C., Chang, J., Gensch, E., Rudner, A. & Tabtiang, R.A. C. elegans mutant that lives twice as long as wild type. Nature 366, 461–464 (1993).
Arbeitman, M.N. et al. Gene expression during the life cycle of Drosophila melanogaster. Science 297, 2270–2275 (2002).
Jiang, M. et al. Genome-wide analysis of developmental and sex-regulated gene expression profiles in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 98, 218–223 (2001).
Gaudet, J. & Mango, S.E. Regulation of organogenesis by the Caenorhabditis elegans FoxA protein PHA-4. Science 295, 821–825 (2002).
Chu, S. et al. The transcriptional program of sporulation in budding yeast. Science 282, 699–705 (1998).
Reinke, V. et al. A global profile of germline gene expression in C. elegans. Mol. Cell 6, 605–616 (2000).
Raghavan, A. et al. Genome-wide analysis of mRNA decay in resting and activated primary human T lymphocytes. Nucleic Acids Res. 30, 5529–5538 (2002).
Wang, Y. et al. Precision and functional specificity in mRNA decay. Proc. Natl. Acad. Sci. USA 99, 5860–5865 (2002).
Boldrick, J.C. et al. Stereotyped and specific gene expression programs in human innate immune responses to bacteria. Proc. Natl. Acad. Sci. USA 99, 972–977 (2002).
Detweiler, C.S., Cunanan, D.B. & Falkow, S. Host microarray analysis reveals a role for the Salmonella response regulator phoP in human macrophage cell death. Proc. Natl. Acad. Sci. USA 98, 5850–5855 (2001).
Guillemin, K., Salama, N.R., Tompkins, L.S. & Falkow, S. Cag pathogenicity island-specific responses of gastric epithelial cells to Helicobacter pylori infection. Proc. Natl. Acad. Sci. USA 99, 15136–15141 (2002).
Cuadras, M.A., Feigelstock, D.A., An, S. & Greenberg, H.B. Gene expression pattern in Caco-2 cells following rotavirus infection. J. Virol. 76, 4467–4482 (2002).
Whitney, A.R. et al. Individuality and variation in gene expression patterns in human blood. Proc. Natl. Acad. Sci. USA 100, 1896–1901 (2003).
Dillin, A. et al. Rates of behavior and aging specified by mitochondrial function during development. Science 298, 2398–2401 (2002).
Dillin, A., Crawford, D.K. & Kenyon, C. Timing requirements for insulin/IGF-1 signaling in C. elegans. Science 298, 830–834 (2002).
Somani, S.M. et al. Influence of age on caloric expenditure during exercise. Int. J. Clin. Pharmacol. Ther. Toxicol. 30, 1–6 (1992).
Bluher, M., Kahn, B. & Kahn, C. Extended longevity in mice lacking the insulin receptor in adipose tissue. Science 299, 572–574 (2003).
Wang, W., Cherry, J.M., Botstein, D. & Li, H. A systematic approach to reconstructing transcription networks in Saccharomycescerevisiae. Proc. Natl. Acad. Sci. USA 99, 16893–16898 (2002).
Segal, E. et al. Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data. Nat. Genet. 34, 166–176 (2003).
Whitfield, M.L. et al. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol. Biol. Cell. 13, 1977–2000 (2002).
Teichmann, S.A. & Babu, M.M. Conservation of gene co-regulation in prokaryotes and eukaryotes. Trends Biotechnol. 20, 407–410 (2002).
Alter, O., Brown, P.O. & Botstein, D. Generalized singular value decomposition for comparative analysis of genome-scale expression data sets of two different organisms. Proc. Natl. Acad. Sci. USA 100, 3351–3356 (2003).
van Noort, V., Snel, B. & Huynen, M.A. Predicting gene function by conserved co-expression. Trends Genet. 19, 238–242 (2003).
Stuart, J.M., Segal, E., Koller, D. & Kim, S.K. A gene-coexpression network for global discovery of conserved genetic modules. Science 302, 249–255 (2003).
Gygi, S.P. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Edgar, R., Domrachev, M. & Lash, A.E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Brazma, A. et al. ArrayExpress—a public repository for microarray gene expression data at the EBI. Nucleic Acids Res. 31, 68–71 (2003).
Stoeckert, C.J., Jr., Causton, H.C. & Ball, C.A. Microarray databases: standards and ontologies. Nat. Genet. 32 Suppl, 469–473 (2002).
Rubin, G.M. et al. Comparative genomics of the eukaryotes. Science 287, 2204–2215 (2000).
Hsin, H. & Kenyon, C. Signals from the reproductive system regulate the lifespan of C. elegans. Nature 399, 362–366 (1999).
Lewis, J.A. & Fleming, J.T. Basic culture methods. in Methods in Cell Biology, Volume 48: Caenorhabditis elegans: Modern Biological Analysis of an Organism (eds. Epstein, H.F. & Shakes, D.C.) 4–30 (Academic Press, San Diego, California, 1995).
Acknowledgements
We thank A. Malmberg, C. Patil, J. DeRisi and I. Herskowitz for discussions and comments on the manuscript; A. Dillin and J. Lehrer-Graiwer for assistance in building the C. elegans microarrays; S. Meadows for assistance with D. melanogaster expression profiling; and J. DeRisi and H. Bennett for advice, instruction and use of their equipment. This work was supported by a grant from the US National Institute on Deafness and Other Communication Disorders to C.I.B., a grant from the Ellison Foundation to C.K., and a Sandler Grant, Packard Fellowship and Life Sciences Informatics grant to H.L. S.A.M. was a Howard Hughes Medical Institute graduate research fellow; C.T.M. is a Life Sciences Research postdoctoral fellow; C.I.B. and Y.N.J. are investigators of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
McCarroll, S., Murphy, C., Zou, S. et al. Comparing genomic expression patterns across species identifies shared transcriptional profile in aging. Nat Genet 36, 197–204 (2004). https://doi.org/10.1038/ng1291
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/ng1291
This article is cited by
-
Ageing as a software design flaw
Genome Biology (2023)
-
Regulation of the antennal transcriptome of the dengue vector, Aedes aegypti, during the first gonotrophic cycle
BMC Genomics (2021)
-
A method to analyze time expression profiles demonstrated in a database of chili pepper fruit development
Scientific Reports (2021)
-
Proteomics profiling and pathway analysis of hippocampal aging in rhesus monkeys
BMC Neuroscience (2020)
-
Unique age-related transcriptional signature in the nervous system of the long-lived red sea urchin Mesocentrotus franciscanus
Scientific Reports (2020)