Abstract
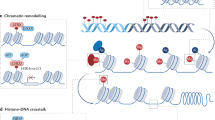
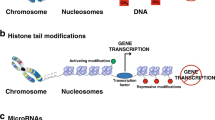
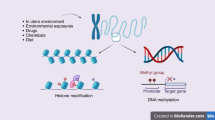
Epigenetics is the study of the heritable changes in gene expression that occur without a change in the DNA sequence itself. These heritable epigenetic changes include chromatin folding and attachment to the nuclear matrix, packaging of DNA around nucleosomes, histone modifications, and DNA methylation. The epigenome is particularly susceptible to dysregulation during gestation, neonatal development, puberty, and old age. Nevertheless, it is most vulnerable to environmental factors during embryogenesis because the DNA synthetic rate is high, and the elaborate DNA methylation patterning and chromatin structure required for normal tissue development is established during early development. Metastable epialleles are alleles that are variably expressed in genetically identical individuals due to epigenetic modifications established during early development and are thought to be particularly vulnerable to environmental influences. The viable yellow agouti (Avy) allele, whose expression is correlated to DNA methylation, is a murine metastable epiallele, which has been used as an epigenetic biosensor for environmental factors affecting the fetal epigenome. In this review, we introduce epigenetic gene regulation, describe important epigenetic phenomenon in mammals, summarize literature linking the early environment to developmental plasticity of the fetal epigenome, and promote the necessity to identify epigenetically labile genes in the mouse and human genomes.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
Abbreviations
- A :
-
agouti
- A vy :
-
viable yellow agouti
- Axin fu :
-
axin fused
- CpG:
-
cytosine-guanine dinucleotide
- IAP:
-
intracisternal A particle
- LOI:
-
loss of imprinting
- RISC:
-
RNA-induced silencing complex
- RITS:
-
RNA-induced initiation of transcriptional silencing
- SAM:
-
S-adenosylmethionine
References
Waterland RA, Jirtle RL 2003 Transposable elements: targets for early nutritional effects on epigenetic gene regulation. Mol Cell Biol 23: 5293–5300
Waterland RA, Jirtle RL 2004 Early nutrition, epigenetic changes at transposons and imprinted genes, and enhanced susceptibility to adult chronic diseases. Nutrition 20: 63–68
Morgan HD, Sutherland HG, Martin DI, Whitelaw E 1999 Epigenetic inheritance at the agouti locus in the mouse. Nat Genet 23: 314–318
Rakyan VK, Chong S, Champ ME, Cuthbert PC, Morgan HD, Luu KV, Whitelaw E 2003 Transgenerational inheritance of epigenetic states at the murine AxinFu allele occurs after maternal and paternal transmission. Proc Natl Acad Sci U S A 100: 2538–2543
Waddington CH 1940 Organisers and Genes. Cambridge University Press, Cambridge, UK
Holliday R, Pugh J 1975 DNA modification mechanisms and gene activity during development. Science 187: 226–232
Willard HF, Brown CJ, Carrel L, Hendrich B, Miller AP 1993 Epigenetic and chromosomal control of gene expression: molecular and genetic analysis of X chromosome inactivation. Cold Spring Harb Symp Quant Biol 58: 315–322
Monk M 1988 Genomic imprinting. Genes Dev 2: 921–925
Murrell A, Rakyan VK, Beck S 2005 From genome to epigenome. Hum Mol Genet 14: R3–R10
Lane N, Dean W, Erhardt S, Hajkova P, Surani A, Walter J, Reik W 2003 Resistance of IAPs to methylation reprogramming may provide a mechanism for epigenetic inheritance in the mouse. Genesis 35: 88–93
Goll MG, Bestor TH 2005 Eukaryotic methyltransferases. Annu Rev Biochem 74: 481–514
Vire E, Brenner C, Deplus R, Blanchon L, Fraga M, Didelot C, Morey L, Van Eynde A, Bernard D, Vanderwinden J, Bollen M, Esteller M, Di Croce L, de Launoit Y, Fuks F 2006 The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439: 871–874
Murphy SK, Jirtle RL 2003 Imprinting evolution and the price of silence. Bioessays 25: 577–588
Gardiner-Garden M, Frommer M 1987 Cpg islands in vertebrate genomes. J Mol Biol 196: 261–282
Saha A, Wittmeyer J, Cairns BR 2006 Chromatin remodelling: the industrial revolution of DNA around histones. Nat Rev Mol Cell Biol 7: 437–447
Caiafa P, Zampieri M 2005 DNA methylation and chromatin structure: the puzzling CpG islands. J Cell Biochem 94: 257–265
Cheung P, Lau P 2005 Epigenetic regulation by histone methylation and histone variants. Mol Endocrinol 19: 563–573
Verdel A, Jia S, Gerber S, Sugiyama T, Gygi S, Grewal SI, Moazed D 2004 RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303: 672–676
Matzke MA, Birchler JA 2005 RNAi-mediated pathways in the nucleus. Nat Rev Genet 6: 24–35
Bestor TH 2003 Cytosine methylation mediates sexual conflict. Trends Genet 19: 185–190
Yoder JA, Walsh CP, Bestor TH 1997 Cytosine methylation and the ecology of intragenomic parasite. Trends Genet 13: 335–340
Rakyan VK, Blewitt ME, Druker R, Preis JI, Whitelaw E 2002 Metastable epialleles in mammals. Trends Genet 18: 348–351
Dolinoy DC, Wiedman J, Waterland R, Jirtle RL 2006 Maternal genistein alters coat color and protects Avy mouse offspring from obesity by modifying the fetal epigenome. Environ Health Perspect 114: 567–572
Waterland RA, Dolinoy DC, Lin JR, Smith CA, Shi X, Tahiliani KG 2006 Maternal methyl supplements increase offspring DNA methylation at AxinFused. Genesis 44: 401–406
Duhl DM, Vrieling H, Miller KA, Wolff GL, Barsh GS 1994 Neomorphic agouti mutations in obese yellow mice. Nat Genet 8: 59–65
Miltenberger RJ, Mynatt RL, Wilkinson JE, Woychik RP 1997 The role of the agouti gene in the Yellow Obese syndrome. J Nutr 127: 1902S–1907S.
Cooney CA, Dave AA, Wolff GL 2002 Maternal methyl supplements in mice affect epigenetic variation and DNA methylation of offspring. J Nutr 132: 2393S–2400S.
Zeng L, Fagotto F, Zhang T, Hsu W, Vasicek TJ, Perry WL, Lee JJ, Tilghman SM, Gumbiner BM, Costantini F 1997 The mouse fused locus encodes axin, an inhibitor of the wnt signaling pathway that regulates embryonic axis formation. Cell 90: 181–192
Flood WD, Ruvinsky A 2001 Alternative splicing and expressivity of the Axin(Fu) allele in mice. Heredity 87: 146–152
Reik W, Walter J 2001 Genomic imprinting: parental influence on the genome. Nat Rev Genet 2: 21–32
Wilkins JF, Haig D 2003 What good is genomic imprinting: the function of parent-specific gene expression. Nat Rev Genet 4: 359–368
Haig D, Graham C 1991 Genomic imprinting and the strange case of the insulin-like growth factor II receptor. Cell 64: 1045–1046
Killian JK, Byrd JC, Jirtle JV, Munday BL, Stoskopf MK, MacDonald RG, Jirtle RL 2000 M6P/IGF2R imprinting evolution in mammals. Mol Cell 5: 707–716
Killian JK, Nolan CM, Wylie AA, Li T, Vu TH, Hoffman AR, Jirtle RL 2001 Divergent evolution in M6P/IGF2R imprinting from the Jurassic to the Quaternary. Hum Mol Genet 10: 1721–1728
Evans HK, Weidman JR, Cowley DO, Jirtle RL 2005 Comparative Phylogenetic Analysis of Blcap/Nnat Reveals Eutherian-Specific Imprinted Gene. Mol Biol Evol 22: 1740–1748
Suzuki S, Renfree M, Pask A, Shaw G, Kobayashi S, Kohda T, Kaneko-Ishino T, Ishino F 2005 Genomic imprinting of IGF2, p57(KIP2) and PEG1/MEST in a marsupial, the tammar wallaby. Mech Dev 122: 213–222
Weidman JR, Maloney KA, Jirtle RL 2006 Comparative phylogenetic analysis reveals multiple non-imprinted isoforms of opossum Dlk1. Mamm Genome 17: 157–167
Niemitz EL, Feinberg AP 2004 Epigenetics and assisted reproductive technology: a call for investigation. Am J Hum Genet 74: 599–609
Feinberg AP, Tycko B 2004 The history of cancer epigenetics. Nat Rev Cancer 4: 143–153
Feinberg AP 2004 The epigenetics of cancer etiology. Semin Cancer Biol 14: 427–432
Cruz-Correa M, Cui H, Giardiello FM, Powe NR, Hylind L, Robinson A, Hutcheon DF, Kafonek DR, Brandenburg S, Wu Y, He X, Feinberg AP 2004 Loss of imprinting of insulin growth factor II gene: a potential heritable biomarker for colon neoplasia predisposition. Gastroenterology 126: 964–970
Jirtle RL 2004 IGF2 loss of imprinting: a potential heritable risk factor for colorectal cancer. Gastroenterology 126: 1190–1193
Feinberg AP, Tycko B 2004 The history of cancer epigenetics. Nat Rev Cancer 4: 143–153
Falls JG, Pulford DJ, Wylie AA, Jirtle RL 1999 Genomic imprinting: implications for human disease. Am J Pathol 154: 635–647
Waterland RA, Lin J-R, Smith CA, Jirtle RL 2006 Post-weaning diet affects genomic imprinting at the insulin-like growth factor 2 (Igf2) locus. Hum Mol Genet 15: 705–716
Reik W, Dean W, Walter J 2001 Epigenetic reprogramming in mammalian development. Science 293: 1089–1093
Morgan HD, Santos F, Green K, Dean W, Reik W 2005 Epigenetic reprogramming in mammals. Hum Mol Genet 14: R47–R58
Mayer W, Niveleau A, Walter J, Fundele R, Haaf T 2000 Demethylation of the zygotic paternal genome. Nature 403: 501–502
Santos F, Hendrich B, Reik W, Dean W 2002 Dynamic reprogramming of DNA methylation in the early mouse embryo. Dev Biol 241: 172–182
Wolff GL 1978 Influence of maternal phenotype on metabolic differentiation of agouti locus mutants in the mouse. Genetics 88: 529–539
Belyaev DK, Ruvinsky AO, Borodin PM 1981 Inheritance of alternative states of the fused gene in mice. J Hered 72: 107–112
Rakyan VK, Preis J, Morgan HD, Whitelaw E 2001 The marks, mechanisms and memory of epigenetic states in mammals. Biochem J 356: 1–10
Blewitt ME, Vickaryous NK, Paldi A, Koseki H, Whitelaw E 2006 Dynamic reprogramming of DNA methylation at an epigenetically sensitive allele in mice. PLoS Genet 2: 399–405
Anway MD, Cupp AS, Uzumcu M, Skinner MK 2005 Epigenetic transgenerational actions of endocrine disruptors and male fertility. Science 308: 1466–1469
Barker DJ 2004 The developmental origins of adult disease. J Am Coll Nutr 23: 588S–595S
Allred CD, Allred KF, Ju YH, Virant SM, Helferich WG 2001 Soy diets containing varying amounts of genistein stimulate growth of estrogen-dependent (MCF-7) tumors in a dose-dependent manner. Cancer Res 61: 5045–5050
Jiang YH, Bressler J, Beaudet AL 2004 Epigenetics and human disease. Annu Rev Genomics Hum Genet 5: 479–510
Luedi PP, Hartemink AJ, Jirtle RL 2005 Genome-wide prediction of imprinted murine genes. Genome Res 15: 875–884
McMillen IC, Robinson JS 2005 Developmental origins of the metabolic syndrome: prediction, plasticity, and programming. Physiol Rev 85: 571–633
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was supported by DOE grant DE-FG02-05ER64101 and National Institutes of Health grants ES13053, ES08823, ES015165, and T32-ES07031.
Rights and permissions
About this article
Cite this article
Dolinoy, D., Das, R., Weidman, J. et al. Metastable Epialleles, Imprinting, and the Fetal Origins of Adult Diseases. Pediatr Res 61, 30–37 (2007). https://doi.org/10.1203/pdr.0b013e31804575f7
Received:
Accepted:
Issue date:
DOI: https://doi.org/10.1203/pdr.0b013e31804575f7
This article is cited by
-
Batch-effect detection, correction and characterisation in Illumina HumanMethylation450 and MethylationEPIC BeadChip array data
Clinical Epigenetics (2022)
-
Epigenetic contribution to obesity
Mammalian Genome (2020)
-
Exposure to NO2, CO, and PM2.5 is linked to regional DNA methylation differences in asthma
Clinical Epigenetics (2018)
-
The gymnastics of epigenomics in rice
Plant Cell Reports (2018)
-
Intrauterine Inflammation, Epigenetics, and Microbiome Influences on Preterm Infant Health
Current Pathobiology Reports (2018)