Abstract
Background:
The childhood salivary microbiome, which plays an important role in healthy development, may be influenced by breast milk consumption. The composition of the milk microbiome and the role it plays in the establishment of the infant microbiome are not well understood.
Methods:
Here, we sequenced the bacterial 16S rRNA gene to characterize microbial communities in breast milk and 5-year-old child saliva from 10 low-income, Mexican-American mother–child pairs with a high prevalence of obesity.
Results:
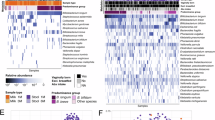
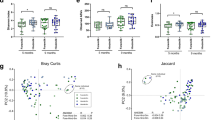
Members of the genus Streptococcus dominated both milk and salivary microbial communities in most subjects. Staphylococcus was observed predominately in milk samples while Prevotella was more prevalent in child saliva. No statistically significant relationships were observed between maternal and child microbiomes or between child microbiome and BMI. However, prepregnancy BMI was correlated with both lower Streptococcus abundance (r = −0.67) and higher microbial diversity (r = 0.77) in breast milk (P < 0.05 for both). Diversity estimates were notably similar to data from other low-income cohorts or children.
Conclusion:
These findings contribute to the currently limited state of knowledge regarding the breast milk and salivary microbiomes in mother–child pairs and may inform future studies seeking to elucidate the relationship between early-life microbial exposures and pediatric health.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Grice EA, Segre JA. The human microbiome: our second genome. Annu Rev Genomics Hum Genet 2012;13:151–70.
Koeth RA, Wang Z, Levison BS, et al. Intestinal microbiota metabolism of l -carnitine, a nutrient in red meat, promotes atherosclerosis. Nat Med 2013;19:576–85.
de Sousa Moraes LF, Grzeskowiak LM, de Sales Teixeira TF, Gouveia Peluzio Mdo C. Intestinal microbiota and probiotics in celiac disease. Clin Microbiol Rev 2014;27:482–9.
Ley RE. Obesity and the human microbiome. Curr Opin Gastroenterol 2010;26:5–11.
Wade WG. The oral microbiome in health and disease. Pharmacol Res 2013;69:137–43.
Docktor MJ, Paster BJ, Abramowicz S, et al. Alterations in diversity of the oral microbiome in pediatric inflammatory bowel disease. Inflamm Bowel Dis 2012;18:935–42.
Groer MW, Luciano AA, Dishaw LJ, Ashmeade TL, Miller E, Gilbert JA. Development of the preterm infant gut microbiome: a research priority. Microbiome 2014;2:38.
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012;486:207–14.
Zaura E, Nicu EA, Krom BP, Keijser BJ. Acquiring and maintaining a normal oral microbiome: current perspective. Front Cell Infect Microbiol 2014;4:85.
Dominguez-Bello MG, Costello EK, Contreras M, et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci USA 2010;107:11971–5.
Cephas KD, Kim J, Mathai RA, et al. Comparative analysis of salivary bacterial microbiome diversity in edentulous infants and their mothers or primary care givers using pyrosequencing. PLoS One 2011;6:e23503.
Douglass JM, Li Y, Tinanoff N. Association of mutans streptococci between caregivers and their children. Pediatr Dent 2008;30:375–87.
Li Y, Ismail AI, Ge Y, Tellez M, Sohn W. Similarity of bacterial populations in saliva from African-American mother–child dyads. J Clin Microbiol 2007;45:3082–5.
Sampaio-Maia B, Monteiro-Silva F. Acquisition and maturation of oral microbiome throughout childhood: an update. Dent Res J (Isfahan) 2014;11:291–301.
Fernández L, Langa S, Martín V, et al. The human milk microbiota: origin and potential roles in health and disease. Pharmacol Res 2013;69:1–10.
Holgerson PL, Vestman NR, Claesson R, et al. Oral microbial profile discriminates breast-fed from formula-fed infants. J Pediatr Gastroenterol Nutr 2013;56:127–36.
Ajslev TA, Andersen CS, Gamborg M, Sørensen TI, Jess T. Childhood overweight after establishment of the gut microbiota: the role of delivery mode, pre-pregnancy weight and early administration of antibiotics. Int J Obes (Lond) 2011;35:522–9.
Kalliomäki M, Collado MC, Salminen S, Isolauri E. Early differences in fecal microbiota composition in children may predict overweight. Am J Clin Nutr 2008;87:534–8.
Eskenazi B, Harley K, Bradman A, et al. Association of in utero organophosphate pesticide exposure and fetal growth and length of gestation in an agricultural population. Environ Health Perspect 2004;112:1116–24.
Aagaard K, Petrosino J, Keitel W, et al. The Human Microbiome Project strategy for comprehensive sampling of the human microbiome and why it matters. FASEB J 2013;27:1012–22.
Cabrera-Rubio R, Collado MC, Laitinen K, Salminen S, Isolauri E, Mira A. The human milk microbiome changes over lactation and is shaped by maternal weight and mode of delivery. Am J Clin Nutr 2012;96:544–51.
Hunt KM, Foster JA, Forney LJ, et al. Characterization of the diversity and temporal stability of bacterial communities in human milk. PLoS One 2011;6:e21313.
Jost T, Lacroix C, Braegger C, Chassard C. Assessment of bacterial diversity in breast milk using culture-dependent and culture-independent approaches. Br J Nutr 2013;110:1253–62.
Collado MC, Isolauri E, Laitinen K, Salminen S. Effect of mother’s weight on infant’s microbiota acquisition, composition, and activity during early infancy: a prospective follow-up study initiated in early pregnancy. Am J Clin Nutr 2010;92:1023–30.
Galley JD, Bailey M, Kamp Dush C, Schoppe-Sullivan S, Christian LM. Maternal obesity is associated with alterations in the gut microbiome in toddlers. PLoS One 2014;9:e113026.
Lin J. Divergence measures based on the Shannon entropy. IEEE Trans Inform Theory 1991;37:145–51.
Ling Z, Liu X, Wang Y, Li L, Xiang C. Pyrosequencing analysis of the salivary microbiota of healthy Chinese children and adults. Microb Ecol 2013;65:487–95.
Zaura E, Keijser BJ, Huse SM, Crielaard W. Defining the healthy “core microbiome” of oral microbial communities. BMC Microbiol 2009;9:259.
Laitinen K, Poussa T, Isolauri E ; Nutrition, Allergy, Mucosal Immunology and Intestinal Microbiota Group. Probiotics and dietary counselling contribute to glucose regulation during and after pregnancy: a randomised controlled trial. Br J Nutr 2009;101:1679–87.
Dogra S, Sakwinska O, Soh SE, et al.; GUSTO Study Group. Dynamics of infant gut microbiota are influenced by delivery mode and gestational duration and are associated with subsequent adiposity. mBio 2015;6:e02419–14.
Caporaso JG, Lauber CL, Walters WA, et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 2012;6:1621–4.
Caporaso JG, Lauber CL, Walters WA, et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 2011;108:Suppl 1:4516–22.
Caporaso JG, Kuczynski J, Stombaugh J, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 2010;7:335–6.
Bokulich NA, Subramanian S, Faith JJ, et al. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 2013;10:57–9.
Rideout JR, He Y, Navas-Molina JA, et al. Subsampled open-reference clustering creates consistent, comprehensive OTU definitions and scales to billions of sequences. PeerJ 2014;2:e545.
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 2010;26:266–7.
DeSantis TZ, Hugenholtz P, Larsen N, et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 2006;72:5069–72.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010;26:2460–1.
McMurdie PJ, Holmes S. Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput Biol 2014;10:e1003531.
Hill MO. Diversity and evenness: a unifying notation and its consequences. Ecology 1973;54:427–32.
Acknowledgements
We are grateful to the laboratory and field staff and participants of the CHAMACOS study for their contributions. We thank Amelia Wallace for assistance with preparation of samples for 16S rRNA gene sequencing and Rosana Weldon for the development of the breast milk collection protocol. The contents of this work are solely the responsibility of the authors and do not necessarily represent the official views of NIEHS or EPA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Davé, V., Street, K., Francis, S. et al. Bacterial microbiome of breast milk and child saliva from low-income Mexican-American women and children. Pediatr Res 79, 846–854 (2016). https://doi.org/10.1038/pr.2016.9
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/pr.2016.9
This article is cited by
-
A cross-sectional study evidences regulations of leukocytes in the colostrum of mothers with obesity
BMC Medicine (2022)
-
Maternal breast milk microbiota and immune markers in relation to subsequent development of celiac disease in offspring
Scientific Reports (2022)
-
The hidden universe of human milk microbiome: origin, composition, determinants, role, and future perspectives
European Journal of Pediatrics (2022)
-
Differential analysis of the bacterial community in colostrum samples from women with gestational diabetes mellitus and obesity
Scientific Reports (2021)
-
DNA extraction approaches substantially influence the assessment of the human breast milk microbiome
Scientific Reports (2020)