Abstract
Background and aims
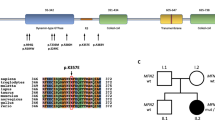
Mutations in neurofilament genes have been linked to several neuromuscular disorders. The neurofilament heavy (NEFH) gene was identified as the causative gene of Charcot–Marie–Tooth disease type 2CC (CMT2CC) in 2016, with a toxic gain of function mechanism caused by the translation and aggregation of cryptic amyloidogenic element (CAE) in the 3′ untranslated region (UTR). But the NEFH-related clinical and genetic spectrums are still unclear in Japan.
Methods
We analyzed all variants in the NEFH gene from our in-house whole-exome sequencing data, established from Japanese nationwide patients with neuromuscular disorders, including Charcot–Marie–Tooth (CMT) disease and spinal muscular atrophy (SMA).
Results
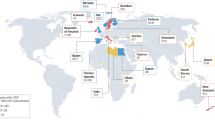
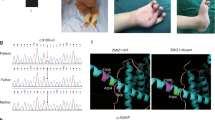
We identified a c.3017dup (p.Pro1007Alafs*56) variant in NEFH from three families clinically diagnosed with CMT, and one family with SMA. In addition to the patients presented with typical peripheral neuropathies, pyramidal signs were observed from one CMT patient. Whereas the SMA patients showed severe characteristic weakness of triceps brachii and quadriceps femoris. All of these four families reside in Kagoshima Prefecture of Japan, and a following haplotype analysis strongly suggests a founder effect.
Interpretation
This is the original report referring to a founder mutation in NEFH. The clinical diversity in our study, comprising CMT, with or without pyramidal signs, and SMA, suggest an extensive involvement of peripheral nerve, anterior horn cells, or both. Our findings broaden the phenotypic spectrum of NEFH-related disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Yuan A, Rao MV, Veeranna, Nixon RA. Neurofilaments and neurofilament proteins in health and disease. Cold Spring Harb Perspect Biol. 2017;9:a018309.
Mersiyanova IV, Perepelov AV, Polyakov AV, Sitnikov VF, Dadali EL, Oparin RB, et al. A new variant of Charcot-Marie-Tooth disease type 2 is probably the result of a mutation in the neurofilament-light gene. Am J Hum Genet. 2000;67:37–46.
Jordanova A. Mutations in the neurofilament light chain gene (NEFL) cause early onset severe Charcot-Marie-Tooth disease. Brain. 2003;126:590–7.
Abe A, Numakura C, Saito K, Koide H, Oka N, Honma A, et al. Neurofilament light chain polypeptide gene mutations in Charcot-Marie-Tooth disease: nonsense mutation probably causes a recessive phenotype. J Hum Genet. 2009;54:94–97.
Corrado L, Carlomagno Y, Falasco L, Mellone S, Godi M, Cova E, et al. A novel peripherin gene (PRPH) mutation identified in one sporadic amyotrophic lateral sclerosis patient. Neurobiol Aging. 2011;32:e1–6.
Figlewicz DA, Krizus A, Martinoli MG, Meininger V, Dib M, Rouleau GA, et al. Variants of the heavy neurofilament subunit are associated with the development of amyotrophic lateral sclerosis. Hum Mol Genet. 1994;3:1757–61.
Tomkins J, Usher P, Slade JY, Ince PG, Curtis A, Bushby K, et al. Novel insertion in the KSP region of the neurofilament heavy gene in amyotrophic lateral sclerosis (ALS). Neuroreport. 1998;17:3967–70.
Rebelo AP, Abrams AJ, Cottenie E, Horga A, Gonzalez M, Bis DM, et al. Cryptic amyloidogenic elements in the 3’ UTRs of neurofilament genes trigger axonal neuropathy. Am J Hum Genet. 2016;98:597–614.
Nam DE, Jung SC, Yoo DH, Choi SS, Seo SY, Kim GH, et al. Axonal Charcot-Marie-Tooth neuropathy concurrent with distal and proximal weakness by translational elongation of the 3’ UTR in NEFH. J Peripher Nerv Syst. 2017;22:200–7.
Jacquier A, Delorme C, Belotti E, Juntas-Morales R, Solé G, Dubourg O, et al. Cryptic amyloidogenic elements in mutant NEFH causing Charcot-Marie-Tooth 2 trigger aggresome formation and neuronal death. Acta Neuropathol Commun. 2017;5:55.
Bian X, Lin P, Li J, Long F, Duan R, Yuan Q, et al. Whole-genome linkage analysis with whole-exome sequencing identifies a novel frameshift variant in NEFH in a Chinese family with Charcot-Marie-Tooth 2: a novel variant in NEFH for Charcot-Marie-Tooth 2. Neurodegener Dis. 2018;18:74–83.
Ikenberg E, Reilich P, Abicht A, Heller C, Schoser B, Walter MC. Charcot-Marie-Tooth disease type 2CC due to a frameshift mutation of the neurofilament heavy polypeptide gene in an Austrian family. Neuromuscul Disord. 2019;29:392–7.
Aruta F, Severi D, Iovino A, Spina E, Barghigiani M, Ruggiero L, et al. Proximal weakness involvement in the first Italian case of Charcot-Marie-Tooth 2CC harboring a novel frameshift variant in NEFH. J Peripher Nerv Syst. 2021;26:231–4.
Charalampopoulou A, Veltsista D, Taraviras S, Chroni E. First case report of Charcot-Marie-Tooth disease type 2CC with a frameshift mutation of NEFH gene in Greece. Neurol Sci. 2021;42:4377–9.
Pipis M, Cortese A, Polke JM, Poh R, Vandrovcova J, Laura M, et al. Charcot-Marie-Tooth disease type 2CC due to NEFH variants causes a progressive, non-length-dependent, motor-predominant phenotype. J Neurol Neurosurg Psychiatry. 2022;93:48–56.
Yan J, Qiao L, Peng H, Liu A, Wu J, Huang J. A novel missense pathogenic variant in NEFH causing rare Charcot-Marie-Tooth neuropathy type 2CC. Neurol Sci. 2021;42:757–63.
Ando M, Hashiguchi A, Okamoto Y, Yoshimura A, Hiramatsu Y, Yuan J, et al. Clinical and genetic diversities of Charcot-Marie-Tooth disease with MFN2 mutations in a large case study. J Peripher Nerv Syst. 2017;22:191–9.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Higuchi Y, Hashiguchi A, Yuan J, Yoshimura A, Mitsui J, Ishiura H, et al. Mutations in MME cause an autosomal-recessive Charcot-Marie-Tooth disease type 2. Ann Neurol. 2016;79:659–72.
Acknowledgements
The authors appreciate Tomoko Ohnishi at Kagoshima University, for their great technical assistance. The authors are supported by Enago (www.enago.jp) for reviewing the English in this report. We appreciate the Division of Gene Research, Research Support Centre, Kagoshima University, for the use of their facilities. This work was supported by Grants-in-Aid from the Research Committee of Ataxia, Health Labour Sciences Research Grant, the Ministry of Health, Labour and Welfare, Japan (201610002B). This research is also supported by the Research program for conquering intractable disease from Japan agency for Medical Research and development (AMED) (201442014A, 201442071A, 17929553) and JSPS KAKENHI Grant Numbers JP18H02742, JP20K16604, JP21K15702, JP21H02842.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ando, M., Higuchi, Y., Okamoto, Y. et al. An NEFH founder mutation causes broad phenotypic spectrum in multiple Japanese families. J Hum Genet 67, 399–403 (2022). https://doi.org/10.1038/s10038-022-01019-y
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s10038-022-01019-y