Abstract
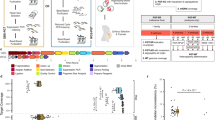
In this study, we aimed to apply preimplantation genetic testing for monogenic disorders (PGT-M) based on mutated allele revealed by sequencing with aneuploidy and linkage analyses (MARSALA) to block the transmission of inborn errors of metabolism (IEMs). After the disease-causing variants were identified through genetic testing, four carrier couples having children affected with IEMs, including methylmalonic aciduria, glutaric acidemia type 1, beta-ketothiolase deficiency, and ornithine transcarbamylase deficiency, sought PGT-M. A series of PGT procedures involving intracytoplasmic sperm injection, blastocyst culture, biopsy of trophectoderm cells, and next-generation sequencing (NGS)-based MARSALA, was performed to provide comprehensive chromosome screening and variant gene analysis. Finally, embryos were selected for transfer, and prenatal diagnosis was conducted to confirm the PGT-M results. All four carrier couples obtained transferrable embryos after PGT. The results of the prenatal diagnosis were consistent with the PGT results, and all couples gave birth to healthy babies free of IEMs. The results of this study confirm that NGS-based MARSALA is an effective approach for families with IEMs to prevent the subsequent transmission of pathological genetic variants to the next generation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
El-Hattab AW. Inborn errors of metabolism. Clin Perinatol. 2015;42:413–39.
Illsinger S, Das AM. Impact of selected inborn errors of metabolism on prenatal and neonatal development. IUBMB Life. 2010;62:403–13.
Campeau PM, Scriver CR, Mitchell JJ. A 25-year longitudinal analysis of treatment efficacy in inborn errors of metabolism. Mol Genet Metab. 2008;95:11–16.
Wang W, Yang J, Xue J, Mu W, Zhang X, Wu W. et al. A comprehensive multiplex PCR based exome-sequencing assay for rapid bloodspot confirmation of inborn errors of metabolism. BMC Med Genet. 2019;20:3
Jiang M, Liu L, Mei H, Li X, Cheng J, Cai Y. Detection of inborn errors of metabolism using GC-MS: over 3 years of experience in southern China. J Pediatr Endocrinol Metab. 2015;28:375–80.
Yang C, Zhou C, Xu P, Jin X, Liu W, Wang W, et al. Newborn screening and diagnosis of inborn errors of metabolism: A 5-year study in an eastern Chinese population. Clin Chim Acta. 2020;502:133–8.
Carvalho F, Moutou C, Dimitriadou E, Dreesen J, Giménez Z, Goossens V, et al. ESHRE PGT Consortium good practice recommendations for the detection of monogenic disorders. Hum Reprod Open. 2020;2020:hoaa018
Handyside AH, Kontogianni EH, Hardy K, Winston RM. Pregnancies from biopsied human preimplantation embryos sexed by Y-specific DNA amplification. Nature. 1990;344:768–70.
Yan L, Huang L, Xu L, Huang J, Ma F, Zhu X. et al. Live births after simultaneous avoidance of monogenic diseases and chromosome abnormality by next-generation sequencing with linkage analyses. Proc Natl Acad Sci USA. 2015;112:15964–9.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Yan L, Cao Y, Chen ZJ, Du J, Wang S, Huang H. et al. Chinese experts’ consensus guideline on preimplantation genetic testing of monogenic disorders. Hum Reprod. 2023;38:ii3–ii13.
Riggs ER, Andersen EF, Cherry AM, Kantarci S, Kearney H, Patel A. et al. Technical standards for the interpretation and reporting of constitutional copy-number variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics (ACMG) and the Clinical Genome Resource (ClinGen). Genet Med. 2020;22:245–57.
An Y, Shen Y, Ma Y, Wang H. Research needs for birth defect prevention and control in China in the genomic screening era. BMJ. 2024;386:e078637.
Zhang C, Zhang P, Yan Y, Zhou B, Wang Y, Tian X. et al. The spectrum of phenylalanine hydroxylase variants and genotype-phenotype correlation in phenylketonuria patients in Gansu, China. Hum Genom. 2023;17:36
Xiang L, Tao J, Deng K, Li X, Li Q, Yuan X. et al. Phenylketonuria incidence in China between 2013 and 2017 based on data from the Chinese newborn screening information system: a descriptive study. BMJ Open. 2019;9:e031474
Minucci A, Moradkhani K, Hwang MJ, Zuppi C, Giardina B, Capoluongo E. Glucose-6-phosphate dehydrogenase (G6PD) mutations database: review of the “old” and update of the new mutations. Blood Cells Mol Dis. 2012;48:154–65.
Rubio C, Rodrigo L, Mir P, Mateu E, Peinado V, Milán M, et al. Use of array comparative genomic hybridization (array-CGH) for embryo assessment: clinical results. Fertil Steril. 2013;99:1044–8.
Handyside AH, Robinson MD, Simpson RJ, Omar MB, Shaw MA, Grudzinskas JG, et al. Isothermal whole genome amplification from single and small numbers of cells: a new era for preimplantation genetic diagnosis of inherited disease. Mol Hum Reprod. 2004;10:767–72.
Natesan SA, Bladon AJ, Coskun S, Qubbaj J, Prates R, Munne S, et al. Genome-wide karyomapping accurately identifies the inheritance of single-gene defects in human preimplantation embryos in vitro. Genet Med. 2014;16:838–45.
Xiong L, Huang L, Tian F, Lu S, Xie XS. Bayesian model for accurate MARSALA (mutated allele revealed by sequencing with aneuploidy and linkage analyses). J Assist Reprod Genet. 2019;36:1263–71.
Minasi MG, Colasante A, Riccio T, Ruberti A, Casciani V, Scarselli F, et al. Correlation between aneuploidy, standard morphology evaluation and morphokinetic development in 1730 biopsied blastocysts: a consecutive case series study. Hum Reprod. 2016;31:2245–54.
van Montfoort A, Carvalho F, Coonen E, Kokkali G, Moutou C, Rubio C, et al. ESHRE PGT Consortium data collection XIX-XX: PGT analyses from 2016 to 2017(†). Hum Reprod Open. 2021;2021:hoab024.
Shaulov T, Zhang L, Chung JT, Son WY, Buckett W, Ao A. Outcomes of preimplantation genetic testing for single gene defects in a privately funded period and publicly funded period: a North-American single center experience. J Reprod Infertil. 2020;21:107–15.
Ben-Nagi J, Jones B, Naja R, Amer A, Sunkara S, SenGupta S. et al. Live birth rate is associated with oocyte yield and number of biopsied and suitable blastocysts to transfer in preimplantation genetic testing (PGT) cycles for monogenic disorders and chromosomal structural rearrangements. Eur J Obstet Gynecol Reprod Biol X. 2019;4:100055
Cram DS, Zhou D. Next generation sequencing: Coping with rare genetic diseases in China. Intractable Rare Dis Res. 2016;5:140–4.
Acknowledgements
We are grateful to the families for their participation in the study. Furthermore, we especially thank the staff at Fujun Genetic Biotechnology Co., Ltd. and Yikon Genomics Co., Ltd.
Funding
This work was supported by the Natural Science Foundation of Fujian Province, China [No. 2023J011341] to Zhihong Wang.
Author information
Authors and Affiliations
Contributions
Xiaoli Li, Qiuxiang Huang, and Fuchun Zhong conceived and designed the study. Xiaoli Li, Qiuxiang Huang, Fuchun Zhong, Zhibiao Chen, Juan Lin, and Zhongli Fan conducted and performed the experiments. Yun Liu, Fenghua Lan, and Zhihong Wang contributed to the genetic counseling. Xiaoli Li, Qiuxiang Huang, Fuchun Zhong, and Zhihong Wang are responsible for data collection and analysis. Xiaoli Li and Qiuxiang Huang wrote the manuscript. Zhihong Wang received funding support and supervised the study. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The protocols for this study were evaluated and approved by the Ethics Committee of Fuzhou General Hospital (Ethics approval no. 2013027).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, X., Huang, Q., Zhong, F. et al. Preimplantation genetic testing for inborn errors of metabolism: observations from a reproductive genetic laboratory in China. J Hum Genet 70, 113–119 (2025). https://doi.org/10.1038/s10038-024-01307-9
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s10038-024-01307-9