Abstract
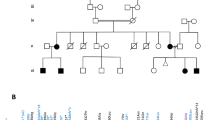
Pathogenic SNF2 related chromatin remodeling ATPase 1 (SMARCA1) variants have been reported in patients with X-linked intellectual disability (XLID) characterized by macrocephaly and variable neurological symptoms. Here, we report two unrelated male patients with XLID due to novel SMARCA1 variants detected by exome sequencing. Patient 1 showed macrocephaly, behavioral difficulty, and learning disability with a hemizygous SMARCA1 variant (NM_003069.5:c.1795 C > T p.[Gln599*]) leading to nonsense-mediated decay. Patient 2 had ataxia and speech delay with a hemizygous missense variant (NM_003069.5:c.1343 G > T p.[Arg448Leu]). Structural modeling suggested that the missense variant, p.(Arg448Leu) might destabilize interactions between SMARCA1 and nucleosomal DNA, thereby contributing to the abberant effect of mutant SMARCA1 protein. Both variants were inherited from their unaffected healthy mothers. This study suggests that hemizygous variants impairing SMARCA1 function can cause XLID with other variable features, such as macrocephaly and ataxia, in men.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Schwartz CE, Louie RJ, Toutain A, Skinner C, Friez MJ, Stevenson RE. X-Linked intellectual disability update 2022. Am J Med Genet A. 2023;191:144–59.
Lee YR, Khan K, Armfield-Uhas K, Srikanth S, Thompson NA, Pardo M, et al. Mutations in FAM50A suggest that Armfield XLID syndrome is a spliceosomopathy. Nat Commun. 2020;11:3698.
Iurlaro M, Masoni F, Flyamer IM, Wirbelauer C, Iskar M, Burger L, et al. Systematic assessment of ISWI subunits shows that NURF creates local accessibility for CTCF. Nat Genet. 2024;56:1203–12.
Picketts D, Mirzaa G, Yan K, Relator R, Timpano S, Yalcin B, et al. Pathogenic variants in SMARCA1 cause an X-linked neurodevelopmental disorder modulated by NURF complex composition. Res Sq. 2023. https://doi.org/10.21203/rs.3.rs-3317938/v1.
Takahashi Y, Date H, Oi H, Adachi T, Imanishi N, Kimura E, et al. Six years’ accomplishment of the Initiative on Rare and Undiagnosed Diseases: nationwide project in Japan to discover causes, mechanisms, and cures. J Hum Genet. 2022;67:505–13.
Kameyama S, Mizuguchi T, Fukuda H, Moey LH, Keng WT, Okamoto N, et al. Biallelic null variants in ZNF142 cause global developmental delay with familial epilepsy and dysmorphic features. J Hum Genet. 2022;67:169–73.
Chio US, Palovcak E, Smith AAA, Autzen H, Muñoz EN, Yu Z, et al. Functionalized graphene-oxide grids enable high-resolution cryo-EM structures of the SNF2h-nucleosome complex without crosslinking. Nat Commun. 2024;15:2225.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Leonard JD, Narlikar GJ. A nucleotide-driven switch regulates flanking DNA length sensing by a dimeric chromatin remodeler. Mol Cell. 2015;57:850–59.
Schubach M, Maass T, Nazaretyan L, Roner S, Kircher M. CADD v1.7: using protein language models, regulatory CNNs and other nucleotide-level scores to improve genome-wide variant predictions. Nucleic Acids Res. 2024;52:D1143–D54.
Yip DJ, Corcoran CP, Alvarez-Saavedra M, DeMaria A, Rennick S, Mears AJ, et al. Snf2l regulates Foxg1-dependent progenitor cell expansion in the developing brain. Dev Cell. 2012;22:871–8.
Zhang X, Theotokis PI, Li N, SHaRe Investigators, Wright CF, Samocha KE, et al. Genetic constraint at single amino acid resolution in protein domains improves missense variant prioritisation and gene discovery. Genome Med. 2024;16:88.
Acknowledgements
We thank the participants for their cooperation. We also thank Ms. K. Takabe, Mr. T. Miyama, Ms. N. Watanabe, Ms. M. Sato, and Ms. S. Sugimoto at the Department of Human Genetics, Yokohama City University Graduate School of Medicine, Japan for their technical assistance. We thank Rachel James, PhD, from Edanz (https://jp.edanz.com/ac) for editing a draft of this manuscript.
Funding
This work was supported by the Japan Agency for Medical Research and Development (AMED) under grant numbers JP24ek0109674, JP24ek0109760, JP24ek0109617, JP24ek0109648, and JP24ek0109677 (N. Matsumoto); Japan Society for the Promotion of Science (JSPS) KAKENHI under grant numbers JP23K27568 and JP23K18278 (T. Mizuguchi), JP24K18862 (M. Sakamoto), JP23K07229 (Y. Uchiyama), JP23K15353 (N. Tsuchida), JP21K07869 (E. Koshimizu), JP23K27520 (S. Miyatake), and JP24K02230 (N. Matsumoto); and the Takeda Science Foundation (T. Mizuguchi, N. Matsumoto).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nishimura, N., Mizuguchi, T., Hamada, K. et al. Hemizygous SMARCA1 variants cause X-linked intellectual disability. J Hum Genet 70, 359–363 (2025). https://doi.org/10.1038/s10038-025-01346-w
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s10038-025-01346-w