Abstract
Background
The incidence of obesity has significantly increased worldwide. However, it is still unclear about the genetic susceptibility of obesity.
Methods
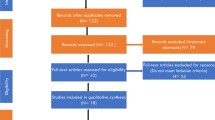
Here we performed the largest European meta-analysis of genome-wide association study, including 98,421 obesity cases and 2,108,019 healthy controls.
Results
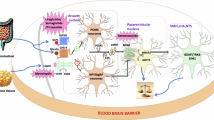
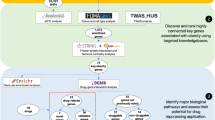
We identified 322 novel genome-wide significant obesity-associated loci and 23 of 32 known loci. SNP-based heritability analyses revealed that common variants explain 17.19 ± 0.59% of genetic risk for obesity, whereas MiXeR predicted an estimated 1.6 million effective sample sizes explaining 90% of obesity-associated phenotypic variance. Across 345 obesity-associated loci, 2000 likely causal genes are indicated, and 410 causal genes are prioritized. Tissue specificity enrichment analyses demonstrated that obesity-related causal genes mainly expressed in brain putamen basal ganglia, hippocampus, amygdala, substantia nigra, and caudate basal ganglia. The genetic correlation and gene-set analyses showed that apart from obesity-related diseases, some brain diseases and mood (e.g., broad depression, neuroticism, mood swings), inflammatory and allergic diseases diseases (e.g., asthma, spondyloarthritis, Hashimoto thyroiditis), cardiovascular diseases (e.g., hypertension, myocardial infarction, coronary artery disease), and lung disease (e.g., interstitial lung disease, chronic obstructive pulmonary disease, lung cancer) have the positive correlations with obesity. Gene-drug interaction analysis suggested that obesity-associated genes overlapped with targets of current medications for obesity. Finally, we used this meta-analysis to explore some potential targets (e.g., GLP1R, SIGMAR1, MC4R) and drug repurposing (e.g., iloprost, flunarizine, edrophonium chloride) for obesity.
Conclusions
We identified 345 genome-wide significant loci, including 322 novel loci for obesity. Based on 345 loci, we provided new biological insights to the etiology of obesity. Of clinical interest, we provided some potential targets and drug repurposing for obesity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Vecchie A, Dallegri F, Carbone F, Bonaventura A, Liberale L, Portincasa P, et al. Obesity phenotypes and their paradoxical association with cardiovascular diseases. Eur J Intern Med. 2018;48:6–17.
Schetz M, De Jong A, Deane AM, Druml W, Hemelaar P, Pelosi P, et al. Obesity in the critically ill: a narrative review. Intensive Care Med. 2019;45:757–69.
Koskinas KC, Van Craenenbroeck EM, Antoniades C, Bluher M, Gorter TM, Hanssen H, et al. Obesity and cardiovascular disease: an ESC clinical consensus statement. Eur J Prev Cardiol. 2025;32:184–220.
Engin A. The definition and prevalence of obesity and metabolic syndrome: correlative clinical evaluation based on phenotypes. Adv Exp Med Biol. 2024;1460:1–25.
Elmaleh-Sachs A, Schwartz JL, Bramante CT, Nicklas JM, Gudzune KA, Jay M. Obesity management in adults: a review. JAMA. 2023;330:2000–15.
Zhang X, Ma N, Lin Q, Chen K, Zheng F, Wu J, et al. Body roundness index and all-cause mortality among US adults. JAMA Netw Open. 2024;7:e2415051.
Lopez-Jimenez F, Almahmeed W, Bays H, Cuevas A, Di Angelantonio E, le Roux CW, et al. Obesity and cardiovascular disease: mechanistic insights and management strategies. A joint position paper by the World Heart Federation and World Obesity Federation. Eur J Prev Cardiol. 2022;29:2218–37.
Reid TJ, Korner J. Medical and surgical treatment of obesity. Med Clin North Am. 2022;106:837–52.
Heianza Y, Qi L. Gene-diet interaction and precision nutrition in obesity. Int J Mol Sci. 2017;18:787.
Zhao J, Bradfield JP, Li M, Wang K, Zhang H, Kim CE, et al. The role of obesity-associated loci identified in genome-wide association studies in the determination of pediatric BMI. Obesity. 2009;17:2254–7.
den Hoed M, Ekelund U, Brage S, Grontved A, Zhao JH, Sharp SJ, et al. Genetic susceptibility to obesity and related traits in childhood and adolescence: influence of loci identified by genome-wide association studies. Diabetes. 2010;59:2980–8.
Bradfield JP, Taal HR, Timpson NJ, Scherag A, Lecoeur C, Warrington NM, et al. A genome-wide association meta-analysis identifies new childhood obesity loci. Nat Genet. 2012;44:526–31.
Berndt SI, Gustafsson S, Magi R, Ganna A, Wheeler E, Feitosa MF, et al. Genome-wide meta-analysis identifies 11 new loci for anthropometric traits and provides insights into genetic architecture. Nat Genet. 2013;45:501–12.
Mansour Aly D, Dwivedi OP, Prasad RB, Karajamaki A, Hjort R, Thangam M, et al. Genome-wide association analyses highlight etiological differences underlying newly defined subtypes of diabetes. Nat Genet. 2021;53:1534–42.
Winkler TW, Day FR, Croteau-Chonka DC, Wood AR, Locke AE, Magi R, et al. Quality control and conduct of genome-wide association meta-analyses. Nat Protoc. 2014;9:1192–212.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
Turner SD. qqman: an R package for visualizing GWAS results using Q–Q and Manhattan plots. J Open Source Softw. 2018;3:731.
Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Rentzsch P, Witten D, Cooper GM, Shendure J, Kircher M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 2019;47:D886–94.
Dong S, Zhao N, Spragins E, Kagda MS, Li M, Assis P, et al. Annotating and prioritizing human non-coding variants with RegulomeDB v.2. Nat Genet. 2023;55:724–6.
Roadmap Epigenomics C, Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, et al. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518:317–30.
Ernst J, Kellis M. ChromHMM: automating chromatin-state discovery and characterization. Nat Methods. 2012;9:215–6.
Bulik-Sullivan BK, Loh PR, Finucane HK, Ripke S, Yang J, Schizophrenia Working Group of the Psychiatric Genomics. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:291–5.
Holland D, Frei O, Desikan R, Fan CC, Shadrin AA, Smeland OB, et al. Beyond SNP heritability: polygenicity and discoverability of phenotypes estimated with a univariate Gaussian mixture model. PLoS Genet. 2020;16:e1008612.
Consortium GT. Human Genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science. 2015;348:648–60.
Quan C, Ping J, Lu H, Zhou G, Lu Y. 3DSNP 2.0: update and expansion of the noncoding genomic variant annotation database. Nucleic Acids Res. 2022;50:D950–5.
de Leeuw CA, Mooij JM, Heskes T, Posthuma D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput Biol. 2015;11:e1004219.
GTEx Consortium; Laboratory, Data Analysis & Coordinating Center (LDACC)—Analysis Working Group; Statistical Methods groups—Analysis Working Group; Enhancing GTEx (eGTEx) groups; NIH Common Fund; NIH/NCI; NIH/NHGRI, et al. Genetic effects on gene expression across human tissues. Nature. 2017;550:204–13.
Gusev A, Ko A, Shi H, Bhatia G, Chung W, Penninx BW, et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet. 2016;48:245–52.
Zhu Z, Zhang F, Hu H, Bakshi A, Robinson MR, Powell JE, et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet. 2016;48:481–7.
Weeks EM, Ulirsch JC, Cheng NY, Trippe BL, Fine RS, Miao J, et al. Leveraging polygenic enrichments of gene features to predict genes underlying complex traits and diseases. Nat Genet. 2023;55:1267–76.
Finan C, Gaulton A, Kruger FA, Lumbers RT, Shah T, Engmann J, et al. The druggable genome and support for target identification and validation in drug development. Sci Transl Med. 2017;9:eaag1166.
Freshour SL, Kiwala S, Cotto KC, Coffman AC, McMichael JF, Song JJ, et al. Integration of the Drug-Gene Interaction Database (DGIdb 4.0) with open crowdsource efforts. Nucleic Acids Res. 2021;49:D1144–51.
Wishart DS, Feunang YD, Guo AC, Lo EJ, Marcu A, Grant JR, et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 2018;46:D1074–82.
Zhang M, Su W, Deng J, Zhai B, Zhu G, Gao R, et al. Multi-ancestry genome-wide meta-analysis with 472,819 individuals identifies 32 novel risk loci for psoriasis. J Transl Med. 2025;23:133.
International League Against Epilepsy Consortium on Complex Epilepsies. GWAS meta-analysis of over 29,000 people with epilepsy identifies 26 risk loci and subtype-specific genetic architecture. Nat Genet. 2023;55:1471–82.
Mirza N, Stevelink R, Taweel B, Koeleman BPC, Marson AG. International League Against Epilepsy Consortium on Complex Epilepsies. Using common genetic variants to find drugs for common epilepsies. Brain Commun. 2021;3:fcab287.
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet. 2014;46:310–5.
Hao RH, Yang TL, Rong Y, Yao S, Dong SS, Chen H, et al. Gene expression profiles indicate tissue-specific obesity regulation changes and strong obesity relevant tissues. Int J Obes. 2018;42:363–9.
Yoganathan P, Karunakaran S, Ho MM, Clee SM. Nutritional regulation of genome-wide association obesity genes in a tissue-dependent manner. Nutr Metab. 2012;9:65.
Pan S, Liu X, Liu T, Zhao Z, Dai Y, Wang YY, et al. Causal inference of genetic variants and genes in amyotrophic lateral sclerosis. Front Genet. 2022;13:917142.
Markham A. Setmelanotide: first approval. Drugs. 2021;81:397–403.
Malone JI, Hansen BC. Does obesity cause type 2 diabetes mellitus (T2DM)? Or is it the opposite?. Pediatr Diabetes. 2019;20:5–9.
Apovian CM. Obesity: definition, comorbidities, causes, and burden. Am J Manag Care. 2016;22:s176–85.
Jastreboff AM, le Roux CW, Stefanski A, Aronne LJ, Halpern B, Wharton S, et al. Tirzepatide for obesity treatment and diabetes prevention. N Engl J Med. 2025;392:958–71.
Garvey WT, Frias JP, Jastreboff AM, le Roux CW, Sattar N, Aizenberg D, et al. Tirzepatide once weekly for the treatment of obesity in people with type 2 diabetes (SURMOUNT-2): a double-blind, randomised, multicentre, placebo-controlled, phase 3 trial. Lancet. 2023;402:613–26.
Ladenheim EE. Liraglutide and obesity: a review of the data so far. Drug Des Devel Ther. 2015;9:1867–75.
Chao AM, Tronieri JS, Amaro A, Wadden TA. Semaglutide for the treatment of obesity. Trends Cardiovasc Med. 2023;33:159–66.
No authors listed. Naltrexone/bupropion for obesity. Drug Ther Bull. 2017;55:126–9.
Bernal-Sore I, Navarro-Marquez M, Osorio-Fuentealba C, Diaz-Castro F, Del Campo A, Donoso-Barraza C, et al. Mifepristone enhances insulin-stimulated Akt phosphorylation and glucose uptake in skeletal muscle cells. Mol Cell Endocrinol. 2018;461:277–83.
Acknowledgements
We thank European Molecular Biology Laboratory-European Bioinformatics Institute (NHGRI-EBI), MRC Integrative Epidemiology Unit (IEU), and FinnGen Biobank for providing the summary statistics of obesity-associated GWAS. We thank the GTEx project for providing summary data for eQTL analysis. We thank the Yang Lab for providing the V8 release of the GTEx eQTL/sQTL summary data in SMR binary (BESD) format. We thank Junghyun Jung in the Mancuso lab for providing pre-computed predictive models for TWAS analysis.
Funding
This study was supported by grants from the National Natural Science Foundation of China (32270933), the R&D Program of the Beijing Municipal Education Commission (KZ202210025035), and the Chinese Institutes for Medical Research, Beijing (CX24PY07). The funding source had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; and decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
RW and GL conceived, initiated, and supervised the project. WS helped with the study design. RG, WS, JD, BZ, GZ, JQ, ZB, and HX collected and analyzed the data. RG, WS, JD, and BZ collected GWAS data and performed the meta-analysis, MAGMA, TWAS, SMR, and drug repurposing analysis. GZ, JQ, ZB, and HX collected GO/KEGG data and performed enrichment, gene set, and genetic overlap analysis. RW wrote a draft of the manuscript. All authors critically reviewed and revised the manuscript and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The Ethics Committee of Beijing Institute of Brain Disorders in Capital Medical University approved this secondary analysis of publicly available datasets that were properly anonymized. Informed consent was obtained at the time of the ethically approved original data collection and is not required for this secondary analysis. All methods were performed in accordance with the relevant guidelines and regulations.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gao, R., Su, W., Deng, J. et al. Genome-wide meta-analysis with 2,206,440 individuals identifies 322 novel risk loci for obesity. Int J Obes (2025). https://doi.org/10.1038/s41366-025-01979-z
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41366-025-01979-z