Abstract
Pediatric acute lymphoblastic leukemia (pALL) is the most common childhood malignancy, yet its etiology remains incompletely understood. However, over the course of three waves of germline genetic research, several non-environmental causes have been identified. Beginning with trisomy 21, seven overt cancer predisposition syndromes (CPSs)—characterized by broad clinical phenotypes that include an elevated risk of pALL—were first described. More recently, newly described CPSs conferring high risk of pALL are increasingly covert, with six exhibiting only minimal or no non-cancer features. These 13 CPSs now represent the principal known hereditary causes of pALL, and human pangenomic data indicates a strong negative selection against mutations in the genes associated with these conditions. Collectively they affect approximately 1 in 450 newborns, of which just a minority will develop the disease. As evidenced by tailored leukemia care protocols for children with trisomy 21, there is growing recognition that CPSs warrant specialized diagnostic, therapeutic, and long-term management strategies. In this review, we investigate the evidence that the 12 other CPSs associated with high risk of pALL may also see benefits from specialized care — even if these needs are often incompletely mapped or addressed in the clinic. Given the rarity of each syndrome, collaborative international research and shared data initiatives will be crucial for advancing knowledge and improving outcomes for these patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hjalgrim LL, Rostgaard K, Schmiegelow K, Söderhäll S, Kolmannskog S, Vettenranta K, et al. Age- and sex-specific incidence of childhood leukemia by immunophenotype in the Nordic countries. J Natl Cancer Inst. 2003;95:1539–1544.
Malard F, Mohty M. Acute lymphoblastic leukaemia. Lancet Lond Engl. 2020;395:1146–62.
Tulstrup M, Stoltze UK, Schmiegelow K, Yang JJ. Epidemiology and Etiology of Childhood ALL. In: Vora A (ed). Childhood Acute Lymphoblastic Leukemia. Springer International Publishing: Cham, 2017, pp 1–27.
Kamper-Jørgensen M, Woodward A, Wohlfahrt J, Benn CS, Simonsen J, Hjalgrim H, et al. Childcare in the first 2 years of life reduces the risk of childhood acute lymphoblastic leukemia. Leukemia. 2008;22:189–93.
Søegaard SH, Rostgaard K, Kamper-Jørgensen M, Schmiegelow K, Hjalgrim H. Childcare attendance and risk of childhood acute lymphoblastic leukaemia: A register study based on the Danish childcare database. Int J Cancer. 2023;152:1817–26.
Jeha S, Pei D, Choi J, Cheng C, Sandlund JT, Coustan-Smith E, et al. Improved CNS Control of Childhood Acute Lymphoblastic Leukemia Without Cranial Irradiation: St Jude Total Therapy Study 16. J Clin Oncol J Am Soc Clin Oncol. 2019;37:3377–91.
Pui C-H, Yang JJ, Bhakta N, Rodriguez-Galindo C. Global efforts toward the cure of childhood acute lymphoblastic leukaemia. Lancet Child Adolesc Health. 2018;2:440–54.
Oskarsson T, Söderhäll S, Arvidson J, Forestier E, Frandsen TL, Hellebostad M et al. Treatment-related mortality in relapsed childhood acute lymphoblastic leukemia. Pediatr Blood Cancer. 2018; 65. https://doi.org/10.1002/pbc.26909.
Ehrlich BS, McNeil MJ, Pham LTD, Chen Y, Rivera J, Acuna C, et al. Treatment-related mortality in children with cancer in low-income and middle-income countries: a systematic review and meta-analysis. Lancet Oncol. 2023;24:967–77.
Schmiegelow K, Müller K, Mogensen SS, Mogensen PR, Wolthers BO, Stoltze UK, et al. Non-infectious chemotherapy-associated acute toxicities during childhood acute lymphoblastic leukemia therapy. F1000Research. 2017;6:444. https://doi.org/10.12688/f1000research.10768.1
Björk-Eriksson T, Boström M, Bryngelsson I-L, Lähteenmäki PM, Jarfelt M, Kalm M, et al. Mortality Among Pediatric Patients With Acute Lymphoblastic Leukemia in Sweden From 1988 to 2017. JAMA Netw Open. 2022;5:e2243857.
Klco JM, Mullighan CG. Advances in germline predisposition to acute leukemias and myeloid neoplasms. Nat Rev Cancer. 2021;21:122–37.
Shendure J, Balasubramanian S, Church GM, Gilbert W, Rogers J, Schloss JA, et al. DNA sequencing at 40: past, present and future. Nature. 2017;550:345–53.
Down JL. Observations on an ethnic classification of idiots. 1866. Ment Retard. 1995;33:54–56.
Krivit W, Good RA. Simultaneous occurrence of mongolism and leukemia; report of a nationwide survey. AMA J Dis Child. 1957;94:289–93.
Lampert F. [Acute lymphoblastic leukemia in sibblings with progressive cerebellar ataxia (Louis-Bar syndrome)]. Dtsch Med Wochenschr. 1969;94:217–20.
McEvoy MW, Mann JR. Neurofibromatosis with leukaemia. Br Med J. 1971;3:641.
Fraumeni JF, Manning MD, Mitus WJ. Acute childhood leukemia: epidemiologic study by cell type of 1263 cases at the Children’s Cancer Research Foundation in Boston, 1947-65. J Natl Cancer Inst. 1971;46:461–70.
Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD, et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature. 2007;446:758–64.
Moriyama T, Relling MV, Yang JJ. Inherited genetic variation in childhood acute lymphoblastic leukemia. Blood. 2015;125:3988–96.
Jeon S, de Smith AJ, Li S, Chen M, Chan TF, Muskens IS, et al. Genome-wide trans-ethnic meta-analysis identifies novel susceptibility loci for childhood acute lymphoblastic leukemia. Leukemia. 2022;36:865–8.
Kullo IJ, Lewis CM, Inouye M, Martin AR, Ripatti S, Chatterjee N. Polygenic scores in biomedical research. Nat Rev Genet. 2022;23:524–32.
Hasle H, Clemmensen IH, Mikkelsen M. Risks of leukaemia and solid tumours in individuals with Down’s syndrome. Lancet. 2000;355:165–9.
Buitenkamp TD, Izraeli S, Zimmermann M, Forestier E, Heerema NA, Heuvel-eibrink MMVD, et al. Acute lymphoblastic leukemia in children with Down syndrome: a retrospective analysis from the Ponte di Legno study group. Blood. 2015;123:70–78.
Izraeli S. The acute lymphoblastic leukemia of Down Syndrome – Genetics and Pathogenesis. Eur J Med Genet. 2015;59:10–13.
Brown AL, de Smith AJ, Gant VU, Yang W, Scheurer ME, Walsh KM, et al. Inherited genetic susceptibility to acute lymphoblastic leukemia in Down syndrome. Blood. 2019;134:1227–37.
Forestier E, Izraeli S, Beverloo B, Haas O, Pession A, Michalová K, et al. Cytogenetic features of acute lymphoblastic and myeloid leukemias in pediatric patients with Down syndrome: an iBFM-SG study. Blood. 2008;111:1575–83.
Hertzberg L, Vendramini E, Ganmore I, Cazzaniga G, Schmitz M, Chalker J, et al. Down syndrome acute lymphoblastic leukemia, a highly heterogeneous disease in which aberrant expression of CRLF2 is associated with mutated JAK2: a report from the International BFM Study Group. Blood. 2010;115:1006–17.
Tal N, Shochat C, Geron I, Bercovich D, Izraeli S. Interleukin 7 and thymic stromal lymphopoietin: from immunity to leukemia. Cell Mol Life Sci CMLS. 2014;71:365–78.
Shochat C, Tal N, Gryshkova V, Birger Y, Bandapalli OR, Cazzaniga G, et al. Novel activating mutations lacking cysteine in type I cytokine receptors in acute lymphoblastic leukemia. Blood. 2014;124:106–10.
Shochat C, Tal N, Bandapalli OR, Palmi C, Ganmore I, te Kronnie G, et al. Gain-of-function mutations in interleukin-7 receptor-α (IL7R) in childhood acute lymphoblastic leukemias. J Exp Med. 2011;208:901–8.
Bercovich D, Ganmore I, Scott LM, Wainreb G, Birger Y, Elimelech A, et al. Mutations of JAK2 in acute lymphoblastic leukaemias associated with Down’s syndrome. Lancet Lond Engl. 2008;372:1484–92.
Schwartzman O, Savino AM, Gombert M, Palmi C, Cario G, Schrappe M, et al. Suppressors and activators of JAK-STAT signaling at diagnosis and relapse of acute lymphoblastic leukemia in Down syndrome. Proc Natl Acad Sci USA. 2017;114:E4030–E4039.
Roberts KG, Li Y, Payne-Turner D, Harvey RC, Yang Y-L, Pei D, et al. Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia. N Engl J Med. 2014;371:1005–15.
Ensor HM, Schwab C, Russell LJ, Richards SM, Morrison H, Masic D, et al. Demographic, clinical, and outcome features of children with acute lymphoblastic leukemia and CRLF2 deregulation: results from the MRC ALL97 clinical trial. Blood. 2011;117:2129–36.
Rand V, Parker H, Russell LJ, Schwab C, Ensor H, Irving J, et al. Genomic characterization implicates iAMP21 as a likely primary genetic event in childhood B-cell precursor acute lymphoblastic leukemia. Blood. 2011;117:6848–55.
Li Z, Chang T-C, Junco JJ, Devidas M, Li Y, Yang W, et al. Genomic landscape of Down syndrome-associated acute lymphoblastic leukemia. Blood. 2023;142:172–84.
Geron I, Savino AM, Fishman H, Tal N, Brown J, Turati VA, et al. An instructive role for Interleukin-7 receptor α in the development of human B-cell precursor leukemia. Nat Commun. 2022;13:659.
Jardine L, Webb S, Goh I, Quiroga Londoño M, Reynolds G, Mather M, et al. Blood and immune development in human fetal bone marrow and Down syndrome. Nature. 2021;598:327–31.
Rabin KR, Devidas M, Chen Z, Ji L, Kairalla J, Hitzler JK, et al. Outcomes in Children, Adolescents, and Young Adults With Down Syndrome and ALL: A Report From the Children’s Oncology Group. J Clin Oncol J Am Soc Clin Oncol. 2024;42:218–27.
Bohnstedt C, Taskinen M, Zeller B, Björgvinsdóttir H, Hafsteinsdottir S, Schmiegelow K. Poor treatment compliance in children with down syndrome and acute lymphoblastic leukemia. J Pediatr Hematol Oncol. 2009;31:79–80.
Rav ES, Wahba A, Patnaik A, Toruner G, Hittle A, Toepfer L, et al. A balancing act: Blinatumomab use in a rare occurrence of Philadelphia chromosome-positive B-cell acute lymphoblastic leukemia in an adolescent patient with Down syndrome. Pediatr Blood Cancer. 2023;70:e30191.
Rothblum-Oviatt C, Wright J, Lefton-Greif MA, McGrath-Morrow SA, Crawford TO, Lederman HM. Ataxia telangiectasia: a review. Orphanet J Rare Dis. 2016;11:159.
Savitsky K, Bar-Shira A, Gilad S, Rotman G, Ziv Y, Vanagaite L, et al. A single ataxia telangiectasia gene with a product similar to PI-3 kinase. Science. 1995;268:1749–53.
Lee J-H, Paull TT. Cellular functions of the protein kinase ATM and their relevance to human disease. Nat Rev Mol Cell Biol. 2021;22:796–814.
Gilad S, Chessa L, Khosravi R, Russell P, Galanty Y, Piane M, et al. Genotype-phenotype relationships in ataxia-telangiectasia and variants. Am J Hum Genet. 1998;62:551–61.
Verhagen MMM, Abdo WF, Willemsen M, Hogervorst FBL, Smeets DFCM, Hiel JAP, et al. Clinical spectrum of ataxia-telangiectasia in adulthood. Neurology. 2009;73:430–7.
Micol R, Slama LB, Suarez F, Mignot LL, Beauté J, Mahlaoui N, et al. Morbidity and mortality from ataxia-telangiectasia are associated with ATM genotype. J Allergy Clin Immunol. 2011;128:382.
Verhagen MMM, Last JI, Hogervorst FBL, Smeets DFCM, Roeleveld N, Verheijen F, et al. Presence of ATM protein and residual kinase activity correlates with the phenotype in ataxia-telangiectasia: a genotype-phenotype study. Hum Mutat. 2012;33:561–71.
Fiévet A, Bellanger D, Rieunier G, Dubois d’Enghien C, Sophie J, Calvas P, et al. Functional classification of ATM variants in ataxia-telangiectasia patients. Hum Mutat. 2019;40:1713–30.
Suarez F, Mahlaoui N, Canioni D, Andriamanga C, Dubois d’Enghien C, Brousse N, et al. Incidence, presentation, and prognosis of malignancies in ataxia-telangiectasia: a report from the French national registry of primary immune deficiencies. J Clin Oncol J Am Soc Clin Oncol. 2015;33:202–8.
Pastorczak A, Attarbaschi A, Bomken S, Borkhardt A, van der Werff Ten Bosch J, Elitzur S, et al. Consensus Recommendations for the Clinical Management of Hematological Malignancies in Patients with DNA Double Stranded Break Disorders. Cancers. 2022;14:2000.
Elitzur S, Shiloh R, Loeffen JLC, Pastorczak A, Takagi M, Bomken S et al. ATM germline pathogenic variants affect outcomes in children with ataxia-telangiectasia and hematological malignancies. Blood. 2024; blood.2024024283.
Varon R, Demuth I, Chrzanowska KH. Nijmegen Breakage Syndrome. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Bean LJ et al. (eds). GeneReviews®. University of Washington, Seattle: Seattle (WA), 1993 http://www.ncbi.nlm.nih.gov/books/NBK1176/ (accessed 8 Sep 2024).
Varon R, Vissinga C, Platzer M, Cerosaletti KM, Chrzanowska KH, Saar K, et al. Nibrin, a novel DNA double-strand break repair protein, is mutated in Nijmegen breakage syndrome. Cell. 1998;93:467–76.
You Z, Chahwan C, Bailis J, Hunter T, Russell P. ATM activation and its recruitment to damaged DNA require binding to the C terminus of Nbs1. Mol Cell Biol. 2005;25:5363–79.
Stracker TH, Morales M, Couto SS, Hussein H, Petrini JHJ. The carboxy terminus of NBS1 is required for induction of apoptosis by the MRE11 complex. Nature. 2007;447:218–21.
Sharapova SO, Pashchenko OE, Bondarenko AV, Vakhlyarskaya SS, Prokofjeva T, Fedorova AS, et al. Geographical distribution, incidence, malignancies, and outcome of 136 eastern slavic patients with Nijmegen breakage syndrome and NBN Founder Variant c.657_661del5. Front Immunol. 2020;11:602482.
Wolska-Kusnierz B, Pastorczak A, Fendler W, Wakulinska A, Dembowska-Baginska B, Heropolitanska-Pliszka E, et al. Hematopoietic stem cell transplantation positively affects the natural history of cancer in Nijmegen breakage syndrome. Clin Cancer Res J Am Assoc Cancer Res. 2021;27:575–84.
Taalman RD, Hustinx TW, Weemaes CM, Seemanová E, Schmidt A, Passarge E, et al. Further delineation of the Nijmegen breakage syndrome. Am J Med Genet. 1989;32:425–31.
Jaspers NG, Taalman RD, Baan C. Patients with an inherited syndrome characterized by immunodeficiency, microcephaly, and chromosomal instability: genetic relationship to ataxia telangiectasia. Am J Hum Genet. 1988;42:66–73.
Chrzanowska KH, Kleijer WJ, Krajewska-Walasek M, Białecka M, Gutkowska A, Goryluk-Kozakiewicz B, et al. Eleven Polish patients with microcephaly, immunodeficiency, and chromosomal instability: the Nijmegen breakage syndrome. Am J Med Genet. 1995;57:462–71.
Slack J, Albert MH, Balashov D, Belohradsky BH, Bertaina A, Bleesing J, et al. Outcome of hematopoietic cell transplantation for DNA double-strand break repair disorders. J Allergy Clin Immunol. 2018;141:322–.e10.
Ababou M. Bloom syndrome and the underlying causes of genetic instability. Mol Genet Metab. 2021;133:35–48.
Chaganti RS, Schonberg S, German J. A manyfold increase in sister chromatid exchanges in Bloom’s syndrome lymphocytes. Proc Natl Acad Sci USA. 1974;71:4508–12.
Hoover A, Turcotte LM, Phelan R, Barbus C, Rayannavar A, Miller BS et al. Longitudinal clinical manifestations of Fanconi anemia: a systematized review. Blood Rev. 2024;68:101225.
Giardino S, Eikema D-J, Piepenbroek B, Algeri M, Ayas M, Faraci M et al. HLA-haploidentical stem cell transplantation in children with inherited bone marrow failure syndromes: a retrospective analysis on behalf of EBMT severe aplastic Anemia and pediatric diseases working parties. Am J Hematol. 2024;99:1066-76.
Lum SH, Eikema D-J, Piepenbroek B, Wynn R, Samarasinghe S, Dalissier A et al. Outcome of hematopoietic stem cell transplantation in 813 pediatric patients with fanconi anemia. Blood. 2024;144:1329–42.
Grant AR, Cushman BJ, Cavé H, Dillon MW, Gelb BD, Gripp KW, et al. Assessing the gene-disease association of 19 genes with the RASopathies using the ClinGen gene curation framework. Hum Mutat. 2018;39:1485–93.
Landry JP, Schertz KL, Chiang Y-J, Bhalla AD, Yi M, Keung EZ, et al. Comparison of cancer prevalence in patients with neurofibromatosis type 1 at an academic cancer center vs in the general population from 1985 to 2020. JAMA Netw Open. 2021;4:e210945.
Kratz CP, Jongmans MC, Cavé H, Wimmer K, Behjati S, Guerrini-Rousseau L, et al. Predisposition to cancer in children and adolescents. Lancet Child Adolesc Health. 2021;5:142–54.
Cavé H, Caye A, Strullu M, Aladjidi N, Vignal C, Ferster A, et al. Acute lymphoblastic leukemia in the context of RASopathies. Eur J Med Genet. 2016;59:173–8.
Shearer P, Parham D, Kovnar E, Kun L, Rao B, Lobe T, et al. Neurofibromatosis type I and malignancy: review of 32 pediatric cases treated at a single institution. Med Pediatr Oncol. 1994;22:78–83.
Galbiati M, Lettieri A, Micalizzi C, Songia S, Morerio C, Biondi A, et al. Natural history of acute lymphoblastic leukemia in neurofibromatosis type 1 monozygotic twins. Leukemia. 2013;27:1778–81.
Bader JL, Miller RW. Neurofibromatosis and childhood leukemia. J Pediatr. 1978;92:925–9.
Horigome H, Sumazaki R, Iwasaki N, Imoto N, Kinugasa H, Saito M, et al. Fatal eosinophilic heart disease in a child with neurofibromatosis-1 complicated by acute lymphoblastic leukemia. Heart Vessels. 2005;20:120–2.
Stiller CA, Chessells JM, Fitchett M. Neurofibromatosis and childhood leukaemia/lymphoma: a population-based UKCCSG study. Br J Cancer. 1994;70:969–72.
Klopfenstein KJ, Sommer A, Ruymann FB. Neurofibromatosis-Noonan syndrome and acute lymphoblastic leukemia: a report of two cases. J Pediatr Hematol Oncol. 1999;21:158–60.
Perilongo G, Felix CA, Meadows AT, Nowell P, Biegel J, Lange BJ. Sequential development of Wilms tumor, T-cell acute lymphoblastic leukemia, medulloblastoma and myeloid leukemia in a child with type 1 neurofibromatosis: a clinical and cytogenetic case report. Leukemia. 1993;7:912–5.
Molteni CG, Te Kronnie G, Bicciato S, Villa T, Tartaglia M, Basso G, et al. PTPN11 mutations in childhood acute lymphoblastic leukemia occur as a secondary event associated with high hyperdiploidy. Leukemia. 2010;24:232–5.
Zipper L, Wagener R, Fischer U, Hoffmann A, Yasin L, Brandes D, et al. Hyperdiploid acute lymphoblastic leukemia in children with LZTR1 germline variants. HemaSphere. 2024;8:e26.
Perrino MR, Das A, Scollon SR, Mitchell SG, Greer M-LC, Yohe ME et al. Update on Pediatric Cancer Surveillance Recommendations for Patients with Neurofibromatosis Type 1, Noonan Syndrome, CBL Syndrome, Costello Syndrome, and Related RASopathies. Clin Cancer Res Off J Am Assoc Cancer Res. 2024. https://doi.org/10.1158/1078-0432.CCR-24-1611.
Wimmer K, Kratz CP, Vasen HFA, Caron O, Colas C, Entz-Werle N, et al. Diagnostic criteria for constitutional mismatch repair deficiency syndrome: suggestions of the European consortium ‘care for CMMRD’ (C4CMMRD). J Med Genet. 2014;51:355–65.
Suerink M, Ripperger T, Messiaen L, Menko FH, Bourdeaut F, Colas C, et al. Constitutional mismatch repair deficiency as a differential diagnosis of neurofibromatosis type 1: consensus guidelines for testing a child without malignancy. J Med Genet. 2019;56:53–62.
Das A, MacFarland SP, Meade J, Hansford JR, Schneider KW, Kuiper RP, et al. Clinical updates and surveillance recommendations for DNA replication repair deficiency syndromes in children and young adults. Clin Cancer Res J Am Assoc Cancer Res. 2024;30:3378–87.
Wimmer K, Rosenbaum T, Messiaen L. Connections between constitutional mismatch repair deficiency syndrome and neurofibromatosis type 1. Clin Genet. 2017;91:507–19.
Ercan AB, Aronson M, Fernandez NR, Chang Y, Levine A, Liu ZA, et al. Clinical and biological landscape of constitutional mismatch-repair deficiency syndrome: an International Replication Repair Deficiency Consortium cohort study. Lancet Oncol. 2024;25:668–82.
Kroeze E, Weijers DD, Hagleitner MM, de Groot-Kruseman HA, Jongmans MCJ, Kuiper RP, et al. High prevalence of constitutional mismatch repair deficiency in a pediatric T-cell lymphoblastic lymphoma cohort. HemaSphere. 2022;6:e668.
González-Acosta M, Marín F, Puliafito B, Bonifaci N, Fernández A, Navarro M, et al. High-sensitivity microsatellite instability assessment for the detection of mismatch repair defects in normal tissue of biallelic germline mismatch repair mutation carriers. J Med Genet. 2020;57:269–73.
Gallon R, Phelps R, Hayes C, Brugieres L, Guerrini-Rousseau L, Colas C, et al. Constitutional microsatellite instability, genotype, and phenotype correlations in constitutional mismatch repair deficiency. Gastroenterology. 2023;164:579–.e8.
Chung J, Negm L, Bianchi V, Stengs L, Das A, Liu ZA, et al. Genomic microsatellite signatures identify germline mismatch repair deficiency and risk of cancer onset. J Clin Oncol J Am Soc Clin Oncol. 2023;41:766–77.
Guerrini-Rousseau L, Gallon R, Pineda M, Brugières L, Baert-Desurmont S, Corsini C et al. Report of the sixth meeting of the European Consortium ‘Care for CMMRD’ (C4CMMRD), Paris, France, November 16th 2022. Fam Cancer. 2024. https://doi.org/10.1007/s10689-024-00403-1.
Muir AW, Houston J, Marshall RJ, Bowman WC, Marshall IG. A comparison of the neuromuscular blocking and autonomic effects of two new short-acting muscle relaxants with those of succinylcholine in the anesthetized cat and pig. Anesthesiology. 1989;70:533–40.
Westdorp H, Kolders S, Hoogerbrugge N, de Vries IJM, Jongmans MCJ, Schreibelt G. Immunotherapy holds the key to cancer treatment and prevention in constitutional mismatch repair deficiency (CMMRD) syndrome. Cancer Lett. 2017;403:159–64.
Das A, Sudhaman S, Morgenstern D, Coblentz A, Chung J, Stone SC, et al. Genomic predictors of response to PD-1 inhibition in children with germline DNA replication repair deficiency. Nat Med. 2022;28:125–35.
Das A, Fernandez NR, Levine A, Bianchi V, Stengs LK, Chung J, et al. ComBined Immunotherapy Improves Outcome For Replication-repair-deficient (RRD) high-grade glioma failing anti-PD-1 monotherapy: a report from the international RRD Consortium. Cancer Discov. 2024;14:258–73.
Yang F, Brady SW, Tang C, Sun H, Du L, Barz MJ, et al. Chemotherapy and mismatch repair deficiency cooperate to fuel TP53 mutagenesis and ALL relapse. Nat Cancer. 2021;2:819–34.
van Engelen N, Roest M, van Dijk F, Sonneveld E, Bladergroen R, van Reijmersdal SV, et al. A novel germline PAX5 single exon deletion in a pediatric patient with precursor B-cell leukemia. Leukemia. 2023;37:1908–11.
Touat M, Li YY, Boynton AN, Spurr LF, Iorgulescu JB, Bohrson CL, et al. Mechanisms and therapeutic implications of hypermutation in gliomas. Nature. 2020;580:517–23.
Bougeard G, Renaux-Petel M, Flaman J-M, Charbonnier C, Fermey P, Belotti M, et al. Revisiting Li-Fraumeni Syndrome from TP53 mutation carriers. J Clin Oncol J Am Soc Clin Oncol. 2015;33:2345–52.
Amadou A, Achatz MIW, Hainaut P. Revisiting tumor patterns and penetrance in germline TP53 mutation carriers: temporal phases of Li-Fraumeni syndrome. Curr Opin Oncol. 2018;30:23–29.
Holmfeldt L, Wei L, Diaz-Flores E, Walsh M, Zhang J, Ding L, et al. The genomic landscape of hypodiploid acute lymphoblastic leukemia. Nat Genet. 2013;45:242–52.
Gu Z, Churchman ML, Roberts KG, Moore I, Zhou X, Nakitandwe J, et al. PAX5-driven subtypes of B-progenitor acute lymphoblastic leukemia. Nat Genet. 2019;51:296–307.
Jongmans MC, Loeffen JL, Waanders E, Hoogerbrugge PM, Ligtenberg MJ, Kuiper RP, Hoogerbrugge N, et al. Recognition of genetic predisposition in pediatric cancer patients: An easy-to-use selection tool. Eur J Med Genet. 2016;59:116–25.
Ripperger T, Bielack SS, Borkhardt A, Brecht IB, Burkhardt B, Calaminus G, et al. Childhood cancer predisposition syndromes-A concise review and recommendations by the Cancer Predisposition Working Group of the Society for Pediatric Oncology and Hematology. Am J Med Genet A. 2017;173:1017–37.
Goudie C, Witkowski L, Cullinan N, Reichman L, Schiller I, Tachdjian M, et al. Performance of the mcgill interactive pediatric oncogenetic guidelines for identifying cancer predisposition syndromes. JAMA Oncol. 2021;7:1806–14.
Hebert R, Cullinan N, Armstrong L, Blood KA, Brossard J, Brunga L, et al. Performance of the eHealth decision support tool, MIPOGG, for recognising children with Li-Fraumeni, DICER1, Constitutional mismatch repair deficiency and Gorlin syndromes. J Med Genet. 2023;60:1218–23.
Valdez JM, Nichols KE, Kesserwan C. Li-Fraumeni syndrome: a paradigm for the understanding of hereditary cancer predisposition. Br J Haematol. 2017;176:539–52.
Qian M, Cao X, Devidas M, Yang W, Cheng C, Dai Y, et al. TP53 Germline Variations influence the predisposition and prognosis of B-cell acute lymphoblastic leukemia in children. J Clin Oncol J Am Soc Clin Oncol. 2018;36:591–9.
Pinto EM, Ribeiro RC, Figueiredo BC, Zambetti GP. TP53-associated pediatric malignancies. Genes Cancer. 2011;2:485–90.
Gonzalez KD, Noltner KA, Buzin CH, Gu D, Wen-Fong CY, Nguyen VQ, et al. Beyond Li Fraumeni Syndrome: clinical characteristics of families with p53 germline mutations. J Clin Oncol J Am Soc Clin Oncol. 2009;27:1250–6.
Renaux-Petel M, Charbonnier F, Théry J-C, Fermey P, Lienard G, Bou J, et al. Contribution of de novo and mosaic TP53 mutations to Li-Fraumeni syndrome. J Med Genet. 2018;55:173–80.
Novel germline TP53 variant (p.(Phe109Ile)) confers high risk of cancer - PubMed. https://pubmed.ncbi.nlm.nih.gov/39317423/ (accessed 22 Oct 2024).
Noetzli L, Lo RW, Lee-Sherick AB, Callaghan M, Noris P, Savoia A, et al. Germline mutations in ETV6 are associated with thrombocytopenia, red cell macrocytosis and predisposition to lymphoblastic leukemia. Nat Genet. 2015;47:535–8.
Topka S, Vijai J, Walsh MF, Jacobs L, Maria A, Villano D, et al. Germline ETV6 Mutations Confer Susceptibility to Acute Lymphoblastic Leukemia and Thrombocytopenia. PLoS Genet. 2015;11:e1005262.
Zhang MY, Churpek JE, Keel SB, Walsh T, Lee MK, Loeb KR, et al. Germline ETV6 mutations in familial thrombocytopenia and hematologic malignancy. Nat Genet. 2015;47:180–5.
Di Paola J, Porter CC. ETV6-related thrombocytopenia and leukemia predisposition. Blood. 2019;134:663–7.
Moriyama T, Metzger ML, Wu G, Nishii R, Qian M, Devidas M, et al. Germline genetic variation in ETV6 and risk of childhood acute lymphoblastic leukaemia: A systematic genetic study. Lancet Oncol. 2015;16:1659–66.
Bloom M, Oak N, Baskin-Doerfler R, Feng R, Iacobucci I, Baviskar P, et al. ETV6 represses inflammatory response genes and regulates HSPC function during stress hematopoiesis in mice. Blood Adv. 2023;7:5608–23.
Nishii R, Baskin-Doerfler R, Yang W, Oak N, Zhao X, Yang W, et al. Molecular basis of ETV6-mediated predisposition to childhood acute lymphoblastic leukemia. Blood. 2021;137:364–73.
Escudero A, Takagi M, Auer F, Friedrich UA, Miyamoto S, Ogawa A, et al. Clinical and immunophenotypic characteristics of familial leukemia predisposition caused by PAX5 germline variants. Leukemia. 2022;36:2338–42.
Bettini LR, Fazio G, Saitta C, Piazza R, Palamini S, Buracchi C et al. Diverse mechanisms of leukemogenesis associated with PAX5 germline mutation. Leukemia. 2024;38:2479–82.
Georgopoulos K, Bigby M, Wang JH, Molnar A, Wu P, Winandy S, et al. The Ikaros gene is required for the development of all lymphoid lineages. Cell. 1994;79:143–56.
Georgopoulos K, Moore DD. Derfler B. Ikaros, an early lymphoid-specific transcription factor and a putative mediator for T cell commitment. Science. 1992;258:808–12.
Winandy S, Wu P, Georgopoulos K. A dominant mutation in the Ikaros gene leads to rapid development of leukemia and lymphoma. Cell. 1995;83:289–99.
Den Boer ML, van Slegtenhorst M, De Menezes RX, Cheok MH, Buijs-Gladdines JGCAM, Peters STCJM, et al. A subtype of childhood acute lymphoblastic leukaemia with poor treatment outcome: a genome-wide classification study. Lancet Oncol. 2009;10:125–34.
Boer JM, van der Veer A, Rizopoulos D, Fiocco M, Sonneveld E, de Groot-Kruseman HA, et al. Prognostic value of rare IKZF1 deletion in childhood B-cell precursor acute lymphoblastic leukemia: an international collaborative study. Leukemia. 2016;30:32–38.
Zhang J, McCastlain K, Yoshihara H, Xu B, Chang Y, Churchman ML, et al. Deregulation of DUX4 and ERG in acute lymphoblastic leukemia. Nat Genet. 2016;48:1481–9.
Kuehn HS, Boast B, Rosenzweig SD. Inborn errors of human IKAROS: LOF and GOF variants associated with primary immunodeficiency. Clin Exp Immunol. 2023;212:129–36.
Kuehn HS, Boisson B, Cunningham-Rundles C, Reichenbach J, Stray-Pedersen A, Gelfand EW, et al. Loss of B Cells in Patients with Heterozygous Mutations in IKAROS. N Engl J Med. 2016;374:1032–43.
Boutboul D, Kuehn HS, Van de Wyngaert Z, Niemela JE, Callebaut I, Stoddard J, et al. Dominant-negative IKZF1 mutations cause a T, B, and myeloid cell combined immunodeficiency. J Clin Invest. 2018;128:3071–87.
Kuehn HS, Niemela JE, Stoddard J, Mannurita SC, Shahin T, Goel S, et al. Germline IKAROS dimerization haploinsufficiency causes hematologic cytopenias and malignancies. Blood. 2021;137:349–63.
Hoshino A, Boutboul D, Zhang Y, Kuehn HS, Hadjadj J, Özdemir N, et al. Gain-of-function IKZF1 variants in humans cause immune dysregulation associated with abnormal T/B cell late differentiation. Sci Immunol. 2022;7:eabi7160.
Churchman ML, Qian M, Kronnie TeG, Zhang R, Yang W, Zhang H, et al. Germline genetic IKZF1 variation and predisposition to childhood acute lymphoblastic leukemia. Cancer Cell. 2018;33:937–.e8.
Song WJ, Sullivan MG, Legare RD, Hutchings S, Tan X, Kufrin D, et al. Haploinsufficiency of CBFA2 causes familial thrombocytopenia with propensity to develop acute myelogenous leukaemia. Nat Genet. 1999;23:166–75.
Latger-Cannard V, Philippe C, Bouquet A, Baccini V, Alessi M-C, Ankri A, et al. Haematological spectrum and genotype-phenotype correlations in nine unrelated families with RUNX1 mutations from the French network on inherited platelet disorders. Orphanet J Rare Dis. 2016;11:49.
Antony-Debré I, Duployez N, Bucci M, Geffroy S, Micol J-B, Renneville A, et al. Somatic mutations associated with leukemic progression of familial platelet disorder with predisposition to acute myeloid leukemia. Leukemia. 2016;30:999–1002.
Li Y, Yang W, Devidas M, Winter SS, Kesserwan C, Yang W, et al. Germline RUNX1 variation and predisposition to childhood acute lymphoblastic leukemia. J Clin Invest. 2021;131:e147898.
Schmiegelow K, Levinsen MF, Attarbaschi A, Baruchel A, Devidas M, Escherich G, et al. Second malignant neoplasms after treatment of childhood acute lymphoblastic leukemia. J Clin Oncol J Am Soc Clin Oncol. 2013;31:2469–76.
Turcotte LM, Neglia JP, Reulen RC, Ronckers CM, van Leeuwen FE, Morton LM, et al. Risk, risk factors, and surveillance of subsequent malignant neoplasms in survivors of childhood cancer: a review. J Clin Oncol J Am Soc Clin Oncol. 2018;36:2145–52.
Levinsen M, Rotevatn EØ, Rosthøj S, Nersting J, Abrahamsson J, Appell ML, et al. Pharmacogenetically based dosing of thiopurines in childhood acute lymphoblastic leukemia: influence on cure rates and risk of second cancer. Pediatr Blood Cancer. 2014;61:797–802.
Inskip PD, Sigurdson AJ, Veiga L, Bhatti P, Ronckers C, Rajaraman P, et al. Radiation-related new primary solid cancers in the childhood cancer survivor study: comparative radiation dose response and modification of treatment effects. Int J Radiat Oncol Biol Phys. 2016;94:800–7.
Yoshida M, Nakabayashi K, Yang W, Sato-Otsubo A, Tsujimoto S, Ogata-Kawata H, et al. NUDT15 variants confer high incidence of second malignancies in children with acute lymphoblastic leukemia. Blood Adv. 2021;5:5420–8.
Junk SV, Förster A, Schmidt G, Zimmermann M, Fedders B, Haermeyer B, et al. Germline variants in patients developing second malignant neoplasms after therapy for pediatric acute lymphoblastic leukemia—a case-control study. Leukemia. 2024;38:887–92.
Zwaan CM, Kaspers GJL, Pieters R, Hählen K, Janka-Schaub GE, van Zantwijk CH, et al. Different drug sensitivity profiles of acute myeloid and lymphoblastic leukemia and normal peripheral blood mononuclear cells in children with and without Down syndrome. Blood. 2002;99:245–51.
Maloney KW, Carroll WL, Carroll AJ, Devidas M, Borowitz MJ, Martin PL, et al. Down syndrome childhood acute lymphoblastic leukemia has a unique spectrum of sentinel cytogenetic lesions that influences treatment outcome: a report from the Children’s Oncology Group. Blood. 2010;116:1045–50.
Szmyd B, Mlynarski W, Pastorczak A. Genetic predisposition to lymphomas: Overview of rare syndromes and inherited familial variants. Mutat Res Rev Mutat Res. 2021;788:108386.
Junk SV, Klein N, Schreek S, Zimmermann M, Möricke A, Bleckmann K, et al. TP53, ETV6 and RUNX1 germline variants in a case series of patients developing secondary neoplasms after treatment for childhood acute lymphoblastic leukemia. Haematologica. 2019;104:e402–e405.
Ripperger T, Schlegelberger B. Acute lymphoblastic leukemia and lymphoma in the context of constitutional mismatch repair deficiency syndrome. Eur J Med Genet. 2016;59:133–42.
Guha T, Malkin D. Inherited TP53 Mutations and the Li-Fraumeni Syndrome. Cold Spring Harb Perspect Med. 2017;7:a026187.
Hendrickson PG, Luo Y, Kohlmann W, Schiffman J, Maese L, Bishop AJ, et al. Radiation therapy and secondary malignancy in Li-Fraumeni syndrome: A hereditary cancer registry study. Cancer Med. 2020;9:7954–63.
Ng AK, Kenney LB, Gilbert ES, Travis LB. Secondary malignancies across the age spectrum. Semin Radiat Oncol. 2010;20:67–78.
Lee JS, DuBois SG, Coccia PF, Bleyer A, Olin RL, Goldsby RE. Increased risk of second malignant neoplasms in adolescents and young adults with cancer. Cancer. 2016;122:116–23.
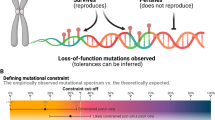
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alföldi J, Wang Q, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:434–43.
Stoltze UK, Foss-Skiftesvik J, Hansen T, van O, Rasmussen S, Karczewski KJ, et al. The evolutionary impact of childhood cancer on the human gene pool. Nat Commun. 2024;15:1881.
Amund Henriksen K, Van Overeem Hansen T, Wadt K, Schmiegelow K, Foss-Skiftesvik J, Stoltze UK. Evolutionary evidence precludes ELP1 as a high-penetrance pediatric cancer predisposition syndrome gene. Neuro-Oncol Adv. 2024;6:vdae165.
Zerella JR, Homan CC, Arts P, Lin X, Spinelli SJ, Venugopal P et al. Germline ERG haploinsufficiency defines a new syndrome with cytopenia and hematological malignancy predisposition. Blood. 2024; blood.2024024607.
Stanley KE, Giordano J, Thorsten V, Buchovecky C, Thomas A, Ganapathi M, et al. Causal Genetic Variants in Stillbirth. N Engl J Med. 2020;383:1107–16.
Stoltze UK, Hansen T, Foss-Skiftesvik J, Byrjalsen A, Henriksen K, Laspiur A et al. Causal germline variants in children with cancer. 2024. https://doi.org/10.2139/ssrn.4889581.
Funding
This work is part of the Danish nation-wide research program Childhood Oncology Network Targeting Research, Organisation & Life expectancy (CONTROL) and supported by the Danish Cancer Society (R-257-A14720 & R376-A22861) and the Danish Childhood Cancer Foundation (2019-5934, 2020-5769 & 2023-001063). GC is funded by the Italian Association for cancer Research (AIRC), IG 2023 grant #29175.
Author information
Authors and Affiliations
Contributions
US editorialized all sections, and was primary writer for all sections not listed below. SI authored section on t21. SE and SJ co-wrote the “ATM & NBS” section. SJ also led the “FA & BSyn” section. HC and MS co-authored “NF1 & RAS”, with AB assisting. RK prepared the “CMMRD & LFS” section, and GC wrote “ETV6 & PAX5.” EF contributed the “IKFZ1 & RUNX1” section, with JT assisting. LL authored the “radio/chemosusceptibility” section, with TM and SJ assisting. LH, AP, KW assisted with structuring, editorializing, and literature review/checking. US generated graphical elements and associated code, with MM assisting. All authors reviewed and approved the final manuscript and contributed to revisions where applicable.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Stoltze, U., Junk, S.V., Byrjalsen, A. et al. Overt and covert genetic causes of pediatric acute lymphoblastic leukemia. Leukemia 39, 1031–1045 (2025). https://doi.org/10.1038/s41375-025-02535-4
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41375-025-02535-4