Abstract
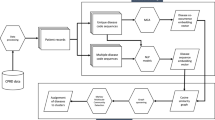
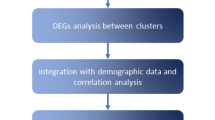
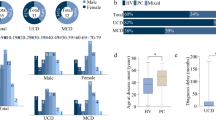
Erdheim-Chester disease (ECD) is a clonal-inflammatory neoplasm driven by mutations in MAPK pathway proto-oncogenes, such as BRAF. Clinical manifestations are protean, affecting virtually every system. This cohort study analyzed 661 patients with ECD to classify them based on clinical features and mutational profiles using unsupervised clustering. Nineteen clinical and mutational variables were subjected to hierarchical clustering combined with k-means. A three-cluster model emerged as the most stable. Most patients were classified according to key features, namely BRAFV600E mutation, and large-vessel, heart, and perirenal involvement. The “Widespread Disease” (WID) cluster (320 patients, 49%) was associated with the presence of the key features and the “Limited Disease” (LIM) cluster (282 patients, 42%) was associated with their absence. The “MAP2K1-RDD” cluster (MAP) was assigned 59 patients (9%), based on MAP2K1 mutation and/or overlapping Rosai-Dorfman-Destombes disease (RDD). Survival analysis revealed worse outcomes for WID compared to LIM (hazard ratio 1.54, 95% CI 1.09–2.17), while no significant survival difference was found for MAP. The identification of these clusters, based on mutational profiles, organ involvement and overlapping conditions, offers a data-driven validation of established clinical observations. These findings substantiate the role of the somatic mutation type in shaping the ECD phenotype.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Due to privacy and ethical restrictions, patient-level data cannot be shared publicly. Aggregated or de-identified data may be available upon reasonable request to the corresponding author, subject to institutional and ethical approvals. Code for generating a fake simulated dataset and carrying out the present analysis on it is available at: https://github.com/miki-tesi/ECD-clusters/.
References
Pegoraro F, Papo M, Maniscalco V, Charlotte F, Haroche J, Vaglio A. Erdheim-Chester disease: a rapidly evolving disease model. Leukemia. 2020;34:2840–57.
Haroche J, Cohen-Aubart F, Amoura Z. Erdheim-Chester disease. Blood. 2020;135:1311–8.
Emile JF, Abla O, Fraitag S, Horne A, Haroche J, Donadieu J, et al. Revised classification of histiocytoses and neoplasms of the macrophage-dendritic cell lineages. Blood. 2016;127:2672–81.
Razanamahery J, Diamond EL, Cohen-Aubart F, Plate KH, Lourida G, Charlotte F, et al. Erdheim-Chester disease with concomitant Rosai-Dorfman like lesions: a distinct entity mainly driven by MAP2K1. Haematologica. 2020;105:e5–8.
Goyal G, Heaney ML, Collin M, Cohen-Aubart F, Vaglio A, Durham BH, et al. Erdheim-Chester disease: consensus recommendations for evaluation, diagnosis, and treatment in the molecular era. Blood. 2020;135:1929–45.
Cohen-Aubart F, Emile JF, Carrat F, Helias-Rodzewicz Z, Taly V, Charlotte F, et al. Phenotypes and survival in Erdheim-Chester disease: results from a 165-patient cohort. Am J Hematol. 2018;93:E114–7.
Chazal T, Pegoraro F, Manari G, Bettiol A, Maniscalco V, Gelain E, et al. Clinical phenotypes and long-term outcome of kidney involvement in Erdheim-Chester histiocytosis. Kidney Int. 2023;103:177–86.
Papo M, Razanamahéry J, Da Silva M, Hélias-Rodzewicz Z, Potapenko V, Bota S, et al. Clinical phenotype of adult-onset systemic histiocytosis harboring BRAF in-frame deletions. Haematologica. 2024;109:3048–52.
Kemps PG, Picarsic J, Durham BH, Hélias-Rodzewicz Z, Hiemcke-Jiwa L, van den Bos C, et al. ALK-positive histiocytosis: a new clinicopathologic spectrum highlighting neurologic involvement and responses to ALK inhibition. Blood. 2022;139:256–80.
Pegoraro F, Papo M, Cohen-Aubart F, Peyronel F, Lugli G, Trambusti I, et al. Long-term outcome and prognosis of mixed histiocytosis (Erdheim-Chester disease and Langerhans Cell Histiocytosis). EClinicalMedicine. 2024;73:102658.
Friedman JS, Durham BH, Reiner AS, Yabe M, Petrova-Drus K, Dogan A, et al. Mixed histiocytic neoplasms: a multicentre series revealing diverse somatic mutations and responses to targeted therapy. Br J Haematol. 2024;205:127–37.
McLachlan GJ. Cluster analysis and related techniques in medical research. Stat Methods Med Res. 1992;1:27–48.
van Rooden SM, Heiser WJ, Kok JN, Verbaan D, van Hilten JJ, Marinus J. The identification of Parkinson’s disease subtypes using cluster analysis: a systematic review. Mov Disord. 2010;25:969–78.
Moore WC, Meyers DA, Wenzel SE, Teague WG, Li H, Li X, et al. Identification of asthma phenotypes using cluster analysis in the Severe Asthma Research Program. Am J Respir Crit Care Med. 2010;181:315–23.
Burgel PR, Paillasseur JL, Caillaud D, Tillie-Leblond I, Chanez P, Escamilla R, et al. Clinical COPD phenotypes: a novel approach using principal component and cluster analyses. Eur Respir J. 2010;36:531–9.
Husson F, Josse J. Multiple correspondence analysis. In: Visualization and verbalization of data. Chapman and Hall/CRC; 2014.
Josse J, Husson F. Selecting the number of components in principal component analysis using cross-validation approximations. Computational Stat Data Anal. 2012;56:1869–79.
Josse J, Chavent M, Liquet B, Husson F. Handling missing values with regularized iterative multiple correspondence analysis. J Classif. 2012;29:91–116.
Murtagh F, Legendre P. Ward’s hierarchical agglomerative clustering method: which algorithms implement Ward’s criterion?. J Classif. 2014;31:274–95.
Steinley D. K-means clustering: a half-century synthesis. Brit J Math Stat Psychol. 2006;59:1–34.
Hennig C. Cluster-wise assessment of cluster stability. Computational Stat Data Anal. 2007;52:258–71.
Lê S, Josse J, Husson F. FactoMineR: an R package for multivariate analysis. J Stat Softw. 2008;25:1–18.
Diamond EL, Durham BH, Haroche J, Yao Z, Ma J, Parikh SA, et al. Diverse and targetable kinase alterations drive histiocytic neoplasms. Cancer Discov. 2016;6:154–65.
Durham BH, Lopez Rodrigo E, Picarsic J, Abramson D, Rotemberg V, De Munck S, et al. Activating mutations in CSF1R and additional receptor tyrosine kinases in histiocytic neoplasms. Nat Med. 2019;25:1839–42.
Berres ML, Lim KPH, Peters T, Price J, Takizawa H, Salmon H, et al. BRAF-V600E expression in precursor versus differentiated dendritic cells defines clinically distinct LCH risk groups. J Exp Med. 2014;211:669–83.
Milne P, Bigley V, Bacon CM, Néel A, McGovern N, Bomken S, et al. Hematopoietic origin of Langerhans cell histiocytosis and Erdheim-Chester disease in adults. Blood. 2017;130:167–75.
Durham BH, Roos-Weil D, Baillou C, Cohen-Aubart F, Yoshimi A, Miyara M, et al. Functional evidence for derivation of systemic histiocytic neoplasms from hematopoietic stem/progenitor cells. Blood. 2017;130:176–80.
Mass E, Jacome-Galarza CE, Blank T, Lazarov T, Durham BH, Ozkaya N, et al. A somatic mutation in erythro-myeloid progenitors causes neurodegenerative disease. Nature. 2017;549:389–93.
Gomez Perdiguero E, Klapproth K, Schulz C, Busch K, Azzoni E, Crozet L, et al. Tissue-resident macrophages originate from yolk-sac-derived erythro-myeloid progenitors. Nature. 2015;518:547–51.
Mass E, Ballesteros I, Farlik M, Halbritter F, Günther P, Crozet L, et al. Specification of tissue-resident macrophages during organogenesis. Science. 2016;353:aaf4238.
Acknowledgements
This work was funded in part by the 2021 Annual Grant of the ECD Global Alliance and by the “Current Research Annual Funding” of the Italian Ministry of Health.
Funding
This work was funded in part by the 2021 Annual Grant of the ECD Global Alliance (FP) and by the “Current Research Annual Funding” of the Italian Ministry of Health (AV). It was supported by the National Institutes of Health/National Cancer Institute (P30 CA008748), as well as the National Cancer Institute (R37CA259260; ELD). This was supported by the Frame Family Fund (ELD), the Joy Family West Foundation (ELD), and the Applebaum Foundation (ELD). ELD discloses unpaid editorial support from Pfizer Inc and serves on an advisory board for Opna Bio, all outside the submitted work.
Author information
Authors and Affiliations
Contributions
Tesi M contributed to conceptualization, data collection and curation, study management, statistical analysis, and writing of the manuscript draft, Pegoraro F to conceptualization, data collection and curation, study management, scientific investigation, critical review and editing of the manuscript, Vaglio A to conceptualization, study management, writing of the manuscript draft, scientific investigation and supervision, Haroche J to conceptualization, study management, scientific investigation and supervision, critical review and editing of the manuscript, Peyronel F to conceptualization and scientific investigation, Catamerò F to data collection and curation. Emile JF, Koster MJ, Goyal G, Collin M, Milne P, Boussouar S, Cohen-Aubart F, Papo M, Amoura Z, Estrada-Veras JI, O’Brien K, Razanamahery J, Goulabchand R, Idbaih A, de Menthon M, Gensous N, Aouba A, Ledoult E, Le Scornet T, Néel A, Go RS, Mazor RD, Campochiaro C, Dagna L, Diamond EL contributed by overseeing patient care at their respective institutions, collecting clinical data, and providing it for inclusion in this study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of Meyer Children’s Hospital IRCCS (approval date: June 9, 2020). All methods were carried out in accordance with relevant guidelines and regulations. Written informed consent was obtained from all participants for inclusion in the study. For participants whose identifiable images are published, written informed consent for publication was also obtained separately.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tesi, M., Pegoraro, F., Peyronel, F. et al. Cluster analysis reveals the clinical spectrum of Erdheim-Chester disease. Leukemia 39, 1987–1996 (2025). https://doi.org/10.1038/s41375-025-02656-w
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41375-025-02656-w