Abstract
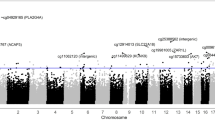
Despite numerous studies demonstrate that genetics and epigenetics factors play important roles on smoking behavior, our understanding of their functional relevance and coordinated regulation remains largely unknown. Here we present a multiomics study on smoking behavior for Chinese smoker population with the goal of not only identifying smoking-associated functional variants but also deciphering the pathogenesis and mechanism underlying smoking behavior in this under-studied ethnic population. After whole-genome sequencing analysis of 1329 Chinese Han male samples in discovery phase and OpenArray analysis of 3744 samples in replication phase, we discovered that three novel variants located near FOXP1 (rs7635815), and between DGCR6 and PRODH (rs796774020), and in ARVCF (rs148582811) were significantly associated with smoking behavior. Subsequently cis-mQTL and cis-eQTL analysis indicated that these variants correlated significantly with the differential methylation regions (DMRs) or differential expressed genes (DEGs) located in the regions where these variants present. Finally, our in silico multiomics analysis revealed several hub genes, like DRD2, PTPRD, FOXP1, COMT, CTNNAP2, to be synergistic regulated each other in the etiology of smoking.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The sequencing data have been deposited in the Genome Sequence Archive (GSA) database with Accession numbers of CRA011029 and HRA007273. Additional information for this study is available from the corresponding author upon reasonable request.
References
CDC. Current cigarette smoking among adults – United States, 2011. MMWR Morb Mortal Wkly Rep. 2012;61:889–94.
CDC. Racial/Ethnic disparities and geographic differences in lung cancer incidence --- 38 States and the District of Columbia, 1998-2006. MMWR Morb Mortal Wkly Rep. 2010;59:1434–8.
Ma Y, Wen L, Cui W, Yuan W, Yang Z, Jiang K, et al. Prevalence of cigarette smoking and nicotine dependence in men and women residing in two provinces in China. Front Psychiatry. 2017;8:254.
Zhang G, Zhan J, Fu H. Trends in smoking prevalence and intensity between 2010 and 2018: implications for tobacco control in China. Int J Environ Res Public Health 2022;19:670.
Zhang M, Yang L, Wang L, Jiang Y, Huang Z, Zhao Z, et al. Trends in smoking prevalence in urban and rural China, 2007 to 2018: Findings from 5 consecutive nationally representative cross-sectional surveys. PLoS Med. 2022;19:e1004064.
Li MD, Cheng R, Ma JZ, Swan GE. A meta-analysis of estimated genetic and environmental effects on smoking behavior in male and female adult twins. Addiction. 2003;98:23–31.
Yang J, Li MD. Converging findings from linkage and association analyses on susceptibility genes for smoking and other addictions. Mol Psychiatry. 2016;21:992–1008.
Liu M, Jiang Y, Wedow R, Li Y, Brazel DM, Chen F, et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat Genet. 2019;51:237–44.
Thorgeirsson TE, Geller F, Sulem P, Rafnar T, Wiste A, Magnusson KP, et al. A variant associated with nicotine dependence, lung cancer and peripheral arterial disease. Nature. 2008;452:638–42.
Saccone SF, Hinrichs AL, Saccone NL, Chase GA, Konvicka K, Madden PA, et al. Cholinergic nicotinic receptor genes implicated in a nicotine dependence association study targeting 348 candidate genes with 3713 SNPs. Hum Mol Genet. 2007;16:36–49.
Ng B, White CC, Klein HU, Sieberts SK, McCabe C, Patrick E, et al. An xQTL map integrates the genetic architecture of the human brain’s transcriptome and epigenome. Nat Neurosci. 2017;20:1418–26.
Schulz H, Ruppert AK, Herms S, Wolf C, Mirza-Schreiber N, Stegle O, et al. Genome-wide mapping of genetic determinants influencing DNA methylation and gene expression in human hippocampus. Nature Commun. 2017;8:1511.
Jaffe AE, Gao Y, Deep-Soboslay A, Tao R, Hyde TM, Weinberger DR, et al. Mapping DNA methylation across development, genotype and schizophrenia in the human frontal cortex. Nature Neurosci. 2016;19:40–47.
Pai S, Li P, Killinger B, Marshall L, Jia P, Liao J, et al. Differential methylation of enhancer at IGF2 is associated with abnormal dopamine synthesis in major psychosis. Nat Commun. 2019;10:2046.
Liu J, Carnero-Montoro E, van Dongen J, Lent S, Nedeljkovic I, Ligthart S, et al. An integrative cross-omics analysis of DNA methylation sites of glucose and insulin homeostasis. Nat Commun. 2019;10:2581.
Hancock DB, Wang JC, Gaddis NC, Levy JL, Saccone NL, Stitzel JA, et al. A multiancestry study identifies novel genetic associations with CHRNA5 methylation in human brain and risk of nicotine dependence. Hum Mol Genet. 2015;24:5940–54.
Shui IM, Wong CJ, Zhao S, Kolb S, Ebot EM, Geybels MS, et al. Prostate tumor DNA methylation is associated with cigarette smoking and adverse prostate cancer outcomes. Cancer. 2016;122:2168–77.
Alexandrov LB, Ju YS, Haase K, Van Loo P, Martincorena I, Nik-Zainal S, et al. Mutational signatures associated with tobacco smoking in human cancer. Science. 2016;354:618–22.
Guida F, Sandanger TM, Castagne R, Campanella G, Polidoro S, Palli D, et al. Dynamics of smoking-induced genome-wide methylation changes with time since smoking cessation. Hum Mol Genet. 2015;24:2349–59.
Joehanes R, Just AC, Marioni RE, Pilling LC, Reynolds LM, Mandaviya PR, et al. Epigenetic signatures of cigarette smoking. Circ Cardiovasc Genet. 2016;9:436–47.
Liu Q, Xu Y, Mao Y, Ma Y, Wang M, Han H, et al. Genetic and epigenetic analysis revealing variants in the NCAM1-TTC12-ANKK1-DRD2 cluster associated significantly with nicotine dependence in Chinese Han smokers. Nicotine Tob Res. 2020;22:1301–9.
Hannon E, Spiers H, Viana J, Pidsley R, Burrage J, Murphy TM, et al. Methylation QTLs in the developing brain and their enrichment in schizophrenia risk loci. Nat Neurosci. 2016;19:48–54.
McClay JL, Shabalin AA, Dozmorov MG, Adkins DE, Kumar G, Nerella S, et al. High density methylation QTL analysis in human blood via next-generation sequencing of the methylated genomic DNA fraction. Genome Biol. 2015;16:291.
Liu Q, Han H, Wang M, Yao Y, Wen L, Jiang K, et al. Association and cis-mQTL analysis of variants in CHRNA3-A5, CHRNA7, CHRNB2, and CHRNB4 in relation to nicotine dependence in a Chinese Han population. Transl Psychiatry. 2018;8:83.
Han H, Liu Q, Yang Z, Wang M, Ma Y, Cao L, et al. Association and cis-mQTL analysis of variants in serotonergic genes associated with nicotine dependence in Chinese Han smokers. Transl Psychiatry. 2018;8:243.
Chen J, Liu Q, Fan R, Han H, Yang Z, Cui W, et al. Demonstration of critical role of GRIN3A in nicotine dependence through both genetic association and molecular functional studies. Addict Biol. 2020;25:e12718.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i90.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Van der Auwera GA, Carneiro MO, Hartl C, Poplin R, Del Angel G, Levy-Moonshine A, et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinform. 2013;43:11 10 11–11 10 33.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Wu MC, Lee S, Cai T, Li Y, Boehnke M, Lin X. Rare-variant association testing for sequencing data with the sequence kernel association test. Am J Hum Genet. 2011;89:82–93.
Ionita-Laza I, Lee S, Makarov V, Buxbaum JD, Lin X. Sequence kernel association tests for the combined effect of rare and common variants. Am J Hum Genet. 2013;92:841–53.
Kechin A, Boyarskikh U, Kel A, Filipenko M. cutPrimers: a new tool for accurate cutting of primers from reads of targeted next generation sequencing. J Comput Biol. 2017;24:1138–43.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20.
Krueger F, Andrews SR. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics. 2011;27:1571–2.
Hansen KD, Langmead B, Irizarry RA. BSmooth: from whole genome bisulfite sequencing reads to differentially methylated regions. Genome Biol. 2012;13:R83.
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinform. 2012;13:86.
Teschendorff AE, Breeze CE, Zheng SC, Beck S. A comparison of reference-based algorithms for correcting cell-type heterogeneity in Epigenome-Wide Association Studies. BMC Bioinform. 2017;18:105.
Pedersen BS, Schwartz DA, Yang IV, Kechris KJ. Comb-p: software for combining, analyzing, grouping and correcting spatially correlated P-values. Bioinformatics. 2012;28:2986–8.
Roadmap Epigenomics C, Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, et al. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518:317–30.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60.
Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol. 2015;33:290–5.
Sahraeian SME, Mohiyuddin M, Sebra R, Tilgner H, Afshar PT, Au KF, et al. Gaining comprehensive biological insight into the transcriptome by performing a broad-spectrum RNA-seq analysis. Nat Commun. 2017;8:59.
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 2004;5:R80.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559.
Shabalin AA. Matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics. 2012;28:1353–8.
Zhao JH, Stacey D, Eriksson N, Macdonald-Dunlop E, Hedman AK, Kalnapenkis A, et al. Genetics of circulating inflammatory proteins identifies drivers of immune-mediated disease risk and therapeutic targets. Nat Immunol. 2023;24:1540–51.
Giambartolomei C, Vukcevic D, Schadt EE, Franke L, Hingorani AD, Wallace C, et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 2014;10:e1004383.
Holstege H, Hulsman M, Charbonnier C, Grenier-Boley B, Quenez O, Grozeva D, et al. Exome sequencing identifies rare damaging variants in ATP8B4 and ABCA1 as risk factors for Alzheimer’s disease. Nat Genet. 2022;54:1786–94.
Chen XD, Xiong DH, Yang TL, Pei YF, Guo YF, Li J, et al. ANKRD7 and CYTL1 are novel risk genes for alcohol drinking behavior. Chinese Med J. 2012;125:1127–34.
Kuo PH, Chuang LC, Su MH, Chen CH, Chen CH, Wu JY, et al. Genome-wide association study for autism spectrum disorder in Taiwanese Han population. PLoS ONE. 2015;10:e0138695.
Armstrong ND, Srinivasasainagendra V, Patki A, Tanner RM, Hidalgo BA, Tiwari HK, et al. Genetic contributors of incident stroke in 10,700 African Americans with hypertension: a meta-analysis from the genetics of hypertension associated treatments and reasons for geographic and racial differences in stroke studies. Front Genet. 2021;12:781451.
Ma Y, Yuan W, Jiang X, Cui WY, Li MD. Updated findings of the association and functional studies of DRD2/ANKK1 variants with addictions. Molecular Neurobiol. 2015;51:281–99.
Bidwell LC, McGeary JE, Gray JC, Palmer RH, Knopik VS, MacKillop J. NCAM1-TTC12-ANKK1-DRD2 variants and smoking motives as intermediate phenotypes for nicotine dependence. Psychopharmacology. 2015;232:1177–86.
Cox JW, Sherva RM, Lunetta KL, Johnson EC, Martin NG, Degenhardt L, et al. Genome-wide association study of opioid cessation. J Clin Med 2020;9:180.
Cao C, Chen J, Lyu C, Yu J, Zhao W, Wang Y, et al. Bioinformatics analysis of the effects of tobacco smoke on gene expression. PLoS ONE. 2015;10:e0143377.
Maurano MT, Humbert R, Rynes E, Thurman RE, Haugen E, Wang H, et al. Systematic localization of common disease-associated variation in regulatory DNA. Science. 2012;337:1190–5.
Consortium EP. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489:57–74.
Li MD, Yoon D, Lee JY, Han BG, Niu T, Payne TJ, et al. Associations of variants in CHRNA5/A3/B4 gene cluster with smoking behaviors in a Korean population. PLoS One. 2010;5:e12183.
Koon HB, Ippolito GC, Banham AH, Tucker PW. FOXP1: a potential therapeutic target in cancer. Expert Opin Ther Targets. 2007;11:955–65.
Kupers LK, Xu X, Jankipersadsing SA, Vaez A, la Bastide-van Gemert S, Scholtens S, et al. DNA methylation mediates the effect of maternal smoking during pregnancy on birthweight of the offspring. Int J Epidemiol. 2015;44:1224–37.
Witt SH, Frank J, Gilles M, Lang M, Treutlein J, Streit F, et al. Impact on birth weight of maternal smoking throughout pregnancy mediated by DNA methylation. BMC Genomics. 2018;19:290.
Rauschert S, Melton PE, Burdge G, Craig JM, Godfrey KM, Holbrook JD, et al. Maternal smoking during pregnancy induces persistent epigenetic changes into adolescence, independent of postnatal smoke exposure and is associated with cardiometabolic risk. Front Genet. 2019;10:770.
Richmond RC, Simpkin AJ, Woodward G, Gaunt TR, Lyttleton O, McArdle WL, et al. Prenatal exposure to maternal smoking and offspring DNA methylation across the lifecourse: findings from the Avon Longitudinal Study of Parents and Children (ALSPAC). Hum Mol Genet. 2015;24:2201–17.
Sanders AR, Rusu I, Duan J, Vander Molen JE, Hou C, Schwab SG, et al. Haplotypic association spanning the 22q11.21 genes COMT and ARVCF with schizophrenia. Mol Psychiatry. 2005;10:353–65.
Liu C, Kanazawa T, Tian Y, Mohamed Saini S, Mancuso S, Mostaid MS, et al. The schizophrenia genetics knowledgebase: a comprehensive update of findings from candidate gene studies. Transl Psychiatry. 2019;9:205.
Yang Z, Chen J, Han H, Wang Y, Shi X, Zhang B, et al. Single nucleotide polymorphisms rs148582811 regulates its host gene ARVCF expression to affect nicotine-associated hippocampus-dependent memory. iScience. 2023;26:108335.
Das Chakraborty R, Bernal AJ, Schoch K, Howard TD, Ip EH, Hooper SR, et al. Dysregulation of DGCR6 and DGCR6L: psychopathological outcomes in chromosome 22q11.2 deletion syndrome. Transl Psychiatry. 2012;2:e105.
Liu H, Heath SC, Sobin C, Roos JL, Galke BL, Blundell ML, et al. Genetic variation at the 22q11 PRODH2/DGCR6 locus presents an unusual pattern and increases susceptibility to schizophrenia. Proc Natl Acad Sci USA. 2002;99:3717–22.
Zhang J, Bai R, Li M, Ye H, Wu C, Wang C, et al. Excessive miR-25-3p maturation via N(6)-methyladenosine stimulated by cigarette smoke promotes pancreatic cancer progression. Nat Commun. 2019;10:1858.
Ma Y, Li MD. Establishment of a strong link between smoking and cancer pathogenesis through DNA methylation analysis. Sci Rep. 2017;7:1811.
Price JF, Mowbray PI, Lee AJ, Rumley A, Lowe GD, Fowkes FG. Relationship between smoking and cardiovascular risk factors in the development of peripheral arterial disease and coronary artery disease: Edinburgh Artery Study. Eur Heart J. 1999;20:344–53.
Jonas MA, Oates JA, Ockene JK, Hennekens CH. Statement on smoking and cardiovascular disease for health care professionals. American Heart Association. Circulation. 1992;86:1664–9.
Inouye M, Abraham G, Nelson CP, Wood AM, Sweeting MJ, Dudbridge F, et al. Genomic risk prediction of coronary artery disease in 480,000 adults: implications for primary prevention. J Am Coll Cardiol. 2018;72:1883–93.
Davies MN, Volta M, Pidsley R, Lunnon K, Dixit A, Lovestone S, et al. Functional annotation of the human brain methylome identifies tissue-specific epigenetic variation across brain and blood. Genome Biol. 2012;13:R43.
Kerkel K, Spadola A, Yuan E, Kosek J, Jiang L, Hod E, et al. Genomic surveys by methylation-sensitive SNP analysis identify sequence-dependent allele-specific DNA methylation. Nat Genet. 2008;40:904–8.
Acknowledgements
This study was supported in part by the National Natural Science Foundation of China (82271560), China Precision Medicine Initiative (2016YFC0906300), Open project of Joint Research Institute of Tobacco and Health of China (2021539200340045), and Research Center for Air Pollution and Health of Zhejiang University.
Author information
Authors and Affiliations
Contributions
MDL, QL, XS, YM, YM, HH, WY and ZY participated sample collections; QL, XS, YM, ZZ, JH, YM and ZY participated in data analysis; WY, JY, YG and ZY managed the project; MDL, QL, XS and ZY participated in paper writing and editing; MDL and ZY conceived the study and was involved in every step of it. All authors approved the paper as submitted.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, M.D., Liu, Q., Shi, X. et al. Integrative analysis of genetics, epigenetics and RNA expression data reveal three susceptibility loci for smoking behavior in Chinese Han population. Mol Psychiatry 29, 3516–3526 (2024). https://doi.org/10.1038/s41380-024-02599-1
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41380-024-02599-1
This article is cited by
-
Investigating the effect of Arvcf reveals an essential role on regulating the mesolimbic dopamine signaling-mediated nicotine reward
Communications Biology (2025)
-
Systematic dissection of pleiotropic loci and critical regulons in excitatory neurons and microglia relevant to neuropsychiatric and ocular diseases
Translational Psychiatry (2025)