Abstract
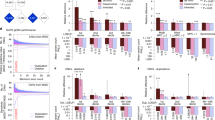
De novo variants adjacent to the canonical splicing sites or in the well-defined splicing-related regions are more likely to impair splicing but remain under-investigated in autism spectrum disorder (ASD). By analyzing large, recent ASD genome sequencing cohorts, we find a significant burden of de novo potential splicing-disrupting variants (PSDVs) in 5048 probands compared to 4090 unaffected siblings. We identified 55 genes with recurrent de novo PSDVs that were highly intolerant to variation. Forty-six of these genes have not been strongly implicated in ASD or other neurodevelopmental disorders previously, including GSK3B. Through international, multicenter collaborations, we assembled genotype and phenotype data for 15 individuals with GSK3B variants and identified common phenotypes including developmental delay, ASD, sleeping disturbance, and aggressive behavior. Using available single-cell transcriptomic data, we show that GSK3B is enriched in dorsal progenitors and intermediate forms of excitatory neurons in the developing brain. We showed that Gsk3b knockdown in mouse excitatory neurons interferes with dendrite arborization and spine maturation which could not be rescued by de novo missense variants identified from affected individuals. In summary, our findings suggest that PSDVs may play an important role in the genetic etiology of ASD and allow for the prioritization of new ASD candidate genes. Importantly, we show that genetic variation resulting in GSK3B loss-of-function can lead to a neurodevelopmental disorder with core features of ASD and developmental delay.
This is a preview of subscription content, access via your institution
Access options






Similar content being viewed by others
Data availability
The ES and GS data used in this study are available from the following resources. The VCF (Variant Call Format) files for SPARK ES generated by Deepvariant and GS data generated by GATK and SPARK phenotype data used in this study are available through SFARI and available to approved researchers at SFARI Base (https://base.sfari.org/) accession nos. SFARI_SPARK_ES_1, SFARI_ SPARK_ES_2, SFARI_SPARK_ES_3, SFARI_SPARK_ES_4, SFARI_SPARK_GS_1, SFARI_SPARK_GS_2, SFARI_SPARK_GS_3 and SFARI_SPARK_GS_4. All GATK VCF files for SSC GS data and SSC phenotype data are available by request from SFARI Base accession no. SFARI_SSC_GS.
Code availability
All software used in this study is publicly available. The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Lord C, Elsabbagh M, Baird G, Veenstra-Vanderweele J. Autism spectrum disorder. Lancet. 2018;392:508–20.
Hirota T, King BH. Autism spectrum disorder: a review. JAMA. 2023;329:157–68.
Lord C, Brugha TS, Charman T, Cusack J, Dumas G, Frazier T, et al. Autism spectrum disorder. Nat Rev Dis Primers. 2020;6:5.
Coe BP, Stessman HAF, Sulovari A, Geisheker MR, Bakken TE, Lake AM, et al. Neurodevelopmental disease genes implicated by de novo mutation and copy number variation morbidity. Nat Genet. 2019;51:106–16.
Matson JL, Shoemaker M. Intellectual disability and its relationship to autism spectrum disorders. Res Dev Disabil. 2009;30:1107–14.
Lai MC, Kassee C, Besney R, Bonato S, Hull L, Mandy W, et al. Prevalence of co-occurring mental health diagnoses in the autism population: a systematic review and meta-analysis. Lancet Psychiatry. 2019;6:819–29.
Iossifov I, O’Roak BJ, Sanders SJ, Ronemus M, Krumm N, Levy D, et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature. 2014;515:216–21.
Trost B, Thiruvahindrapuram B, Chan AJS, Engchuan W, Higginbotham EJ, Howe JL, et al. Genomic architecture of autism from comprehensive whole-genome sequence annotation. Cell. 2022;185:4409–27.e4418
Zhou X, Feliciano P, Shu C, Wang T, Astrovskaya I, Hall JB, et al. Integrating de novo and inherited variants in 42,607 autism cases identifies mutations in new moderate-risk genes. Nat Genet. 2022;54:1305–19.
Fu JM, Satterstrom FK, Peng M, Brand H, Collins RL, Dong S, et al. Rare coding variation provides insight into the genetic architecture and phenotypic context of autism. Nat Genet. 2022;54:1320–31.
Wang T, Zhao PA, Eichler EE. Rare variants and the oligogenic architecture of autism. Trends Genet. 2022;38:895–903.
Krumm N, Turner TN, Baker C, Vives L, Mohajeri K, Witherspoon K, et al. Excess of rare, inherited truncating mutations in autism. Nat Genet. 2015;47:582–8.
Wilfert AB, Turner TN, Murali SC, Hsieh P, Sulovari A, Wang T, et al. Recent ultra-rare inherited variants implicate new autism candidate risk genes. Nat Genet. 2021;53:1125–34.
Hamanaka K, Miyake N, Mizuguchi T, Miyatake S, Uchiyama Y, Tsuchida N, et al. Large-scale discovery of novel neurodevelopmental disorder-related genes through a unified analysis of single-nucleotide and copy number variants. Genome Med. 2022;14:40.
Ruzzo EK, Perez-Cano L, Jung JY, Wang LK, Kashef-Haghighi D, Hartl C, et al. Inherited and de novo genetic risk for autism impacts shared networks. Cell. 2019;178:850–66. e826
Sibley CR, Blazquez L, Ule J. Lessons from non-canonical splicing. Nat Rev Genet. 2016;17:407–21.
Chiang HL, Chen YT, Su JY, Lin HN, Yu CA, Hung YJ, et al. Mechanism and modeling of human disease-associated near-exon intronic variants that perturb RNA splicing. Nat Struct Mol Biol. 2022;29:1043–55.
Eilbeck K, Lewis SE, Mungall CJ, Yandell M, Stein L, Durbin R, et al. The Sequence Ontology: a tool for the unification of genome annotations. Genome Biol. 2005;6:R44.
Takata A, Ionita-Laza I, Gogos JA, Xu B, Karayiorgou M. De novo synonymous mutations in regulatory elements contribute to the genetic etiology of autism and schizophrenia. Neuron. 2016;89:940–7.
Lord J, Gallone G, Short PJ, McRae JF, Ironfield H, Wynn EH, et al. Pathogenicity and selective constraint on variation near splice sites. Genome Res. 2019;29:159–70.
pfeliciano@simonsfoundation.org SCEa, Consortium S. SPARK: A US cohort of 50,000 families to accelerate autism research. Neuron. 2018;97:488–93.
An JY, Lin K, Zhu L, Werling DM, Dong S, Brand H, et al. Genome-wide de novo risk score implicates promoter variation in autism spectrum disorder. Science. 2018;362:eaat6576.
Satterstrom FK, Kosmicki JA, Wang J, Breen MS, De Rubeis S, An JY, et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell. 2020;180:568–84. e523.
C Yuen RK, Merico D, Bookman M, L Howe J, Thiruvahindrapuram B, Patel RV, et al. Whole genome sequencing resource identifies 18 new candidate genes for autism spectrum disorder. Nat Neurosci. 2017;20:602–11.
Ng SB, Turner EH, Robertson PD, Flygare SD, Bigham AW, Lee C, et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature. 2009;461:272–6.
McLaren W, Gil L, Hunt SE, Riat HS, Ritchie GR, Thormann A, et al. The Ensembl Variant Effect Predictor. Genome Biol. 2016;17:122.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Rentzsch P, Schubach M, Shendure J, Kircher M. CADD-Splice-improving genome-wide variant effect prediction using deep learning-derived splice scores. Genome Med. 2021;13:31.
Cheng J, Nguyen TYD, Cygan KJ, Celik MH, Fairbrother WG, Avsec Z, et al. MMSplice: modular modeling improves the predictions of genetic variant effects on splicing. Genome Biol. 2019;20:48.
Zhang X, Li M, Lin H, Rao X, Feng W, Yang Y, et al. regSNPs-splicing: a tool for prioritizing synonymous single-nucleotide substitution. Hum Genet. 2017;136:1279–89.
Liu X, Wu C, Li C, Boerwinkle E. dbNSFP v3.0: a one-stop database of functional predictions and annotations for human nonsynonymous and splice-site SNVs. Hum Mutat. 2016;37:235–41.
Jaganathan K, Kyriazopoulou Panagiotopoulou S, McRae JF, Darbandi SF, Knowles D, Li YI, et al. Predicting splicing from primary sequence with deep learning. Cell. 2019;176:535–48. e524
Samocha KE, Robinson EB, Sanders SJ, Stevens C, Sabo A, McGrath LM, et al. A framework for the interpretation of de novo mutation in human disease. Nat Genet. 2014;46:944–50.
Giorgi FM, Ceraolo C, Mercatelli D. The R language: an engine for bioinformatics and data science. Life. 2022;12:648.
Ware JS, Samocha KE, Homsy J, Daly MJ. Interpreting de novo variation in human disease using denovolyzeR. Curr Protoc Hum Genet. 2015;87:7.
O’Roak BJ, Vives L, Fu W, Egertson JD, Stanaway IB, Phelps IG, et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science. 2012;338:1619–22.
He X, Sanders SJ, Liu L, De Rubeis S, Lim ET, Sutcliffe JS, et al. Integrated model of de novo and inherited genetic variants yields greater power to identify risk genes. PLoS Genet. 2013;9:e1003671.
Stein D, Kars ME, Wu Y, Bayrak CS, Stenson PD, Cooper DN, et al. Genome-wide prediction of pathogenic gain- and loss-of-function variants from ensemble learning of a diverse feature set. Genome Med. 2023;15:103.
Blakes AJM, Wai HA, Davies I, Moledina HE, Ruiz A, Thomas T, et al. A systematic analysis of splicing variants identifies new diagnoses in the 100,000 Genomes Project. Genome Med. 2022;14:79.
Kaplanis J, Samocha KE, Wiel L, Zhang Z, Arvai KJ, Eberhardt RY, et al. Evidence for 28 genetic disorders discovered by combining healthcare and research data. Nature. 2020;586:757–62.
Wang T, Kim CN, Bakken TE, Gillentine MA, Henning B, Mao Y, et al. Integrated gene analyses of de novo variants from 46,612 trios with autism and developmental disorders. Proc Natl Acad Sci USA. 2022;119:e2203491119.
Sobreira N, Schiettecatte F, Valle D, Hamosh A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum Mutat. 2015;36:928–30.
Speir ML, Bhaduri A, Markov NS, Moreno P, Nowakowski TJ, Papatheodorou I, et al. UCSC Cell Browser: visualize your single-cell data. Bioinformatics. 2021;37:4578–80.
Becht E, McInnes L, Healy J, Dutertre CA, Kwok IWH, Ng LG, et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat Biotechnol. 2018;37:38–44.
Trapnell C, Cacchiarelli D, Grimsby J, Pokharel P, Li S, Morse M, et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat Biotechnol. 2014;32:381–6.
Qiu X, Mao Q, Tang Y, Wang L, Chawla R, Pliner HA, et al. Reversed graph embedding resolves complex single-cell trajectories. Nat Methods. 2017;14:979–82.
Beaudoin GMJ, Lee S-H, Singh D, Yuan Y, Ng Y-G, Reichardt LF, et al. Culturing pyramidal neurons from the early postnatal mouse hippocampus and cortex. Nat Protoc. 2012;7:1741–54.
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alfoldi J, Wang Q, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:434–43.
Miller JA, Ding SL, Sunkin SM, Smith KA, Ng L, Szafer A, et al. Transcriptional landscape of the prenatal human brain. Nature. 2014;508:199–206.
Szklarczyk D, Kirsch R, Koutrouli M, Nastou K, Mehryary F, Hachilif R, et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 2023;51:D638–D646.
Thomas PD, Ebert D, Muruganujan A, Mushayahama T, Albou LP, Mi H. PANTHER: making genome-scale phylogenetics accessible to all. Protein Sci. 2022;31:8–22.
De Rubeis S, He X, Goldberg AP, Poultney CS, Samocha K, Cicek AE, et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature. 2014;515:209–15.
Golpich M, Amini E, Hemmati F, Ibrahim NM, Rahmani B, Mohamed Z, et al. Glycogen synthase kinase-3 beta (GSK-3beta) signaling: Implications for Parkinson’s disease. Pharmacol Res. 2015;97:16–26.
Lauretti E, Dincer O, Pratico D. Glycogen synthase kinase-3 signaling in Alzheimer’s disease. Biochim Biophys Acta Mol Cell Res. 2020;1867:118664.
Hernandez F, Lucas JJ, Avila J. GSK3 and tau: two convergence points in Alzheimer’s disease. J Alzheimers Dis. 2013;33:S141–144.
Ng PC, Henikoff S. SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003;31:3812–4.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
Hardt SE, Sadoshima J. Glycogen synthase kinase-3beta: a novel regulator of cardiac hypertrophy and development. Circ Res. 2002;90:1055–63.
Murphy E, Steenbergen C. Inhibition of GSK-3beta as a target for cardioprotection: the importance of timing, location, duration and degree of inhibition. Expert Opin Ther Targets. 2005;9:447–56.
Kerkela R, Woulfe K, Force T. Glycogen synthase kinase-3beta – actively inhibiting hypertrophy. Trends Cardiovasc Med. 2007;17:91–6.
Deng S, Dai G, Chen S, Nie Z, Zhou J, Fang H, et al. Dexamethasone induces osteoblast apoptosis through ROS-PI3K/AKT/GSK3beta signaling pathway. Biomed Pharmacother. 2019;110:602–8.
Wang FS, Ko JY, Weng LH, Yeh DW, Ke HJ, Wu SL. Inhibition of glycogen synthase kinase-3beta attenuates glucocorticoid-induced bone loss. Life Sci. 2009;85:685–92.
Gambardella A, Nagaraju CK, O’Shea PJ, Mohanty ST, Kottam L, Pilling J, et al. Glycogen synthase kinase-3alpha/beta inhibition promotes in vivo amplification of endogenous mesenchymal progenitors with osteogenic and adipogenic potential and their differentiation to the osteogenic lineage. J Bone Miner Res. 2011;26:811–21.
Clough BH, Zeitouni S, Krause U, Chaput CD, Cross LM, Gaharwar AK, et al. Rapid osteogenic enhancement of stem cells in human bone marrow using a glycogen-synthease-kinase-3-beta inhibitor improves osteogenic efficacy in vitro and in vivo. Stem Cells Transl Med. 2018;7:342–53.
Li J, Zhang T, Huang C, Xu M, Xie W, Pei Q, et al. Chemerin located in bone marrow promotes osteogenic differentiation and bone formation via Akt/Gsk3beta/beta-catenin axis in mice. J Cell Physiol. 2021;236:6042–54.
Khayachi A, Ase A, Liao C, Kamesh A, Kuhlmann N, Schorova L, et al. Chronic lithium treatment alters the excitatory/ inhibitory balance of synaptic networks and reduces mGluR5-PKC signalling in mouse cortical neurons. J Psychiatry Neurosci. 2021;46:E402–E414.
Martin L, Magnaudeix A, Esclaire F, Yardin C, Terro F. Inhibition of glycogen synthase kinase-3beta downregulates total tau proteins in cultured neurons and its reversal by the blockade of protein phosphatase-2A. Brain Res. 2009;1252:66–75.
Wu D, Pan W. GSK3: a multifaceted kinase in Wnt signaling. Trends Biochem Sci. 2010;35:161–8.
Kayumi S, Perez-Jurado LA, Palomares M, Rangu S, Sheppard SE, Chung WK, et al. Genomic and phenotypic characterization of 404 individuals with neurodevelopmental disorders caused by CTNNB1 variants. Genet Med. 2022;24:2351–66.
Kharbanda M, Pilz DT, Tomkins S, Chandler K, Saggar A, Fryer A, et al. Clinical features associated with CTNNB1 de novo loss of function mutations in ten individuals. Eur J Med Genet. 2017;60:130–5.
Liu C, Li Y, Semenov M, Han C, Baeg GH, Tan Y, et al. Control of beta-catenin phosphorylation/degradation by a dual-kinase mechanism. Cell. 2002;108:837–47.
Wolff M, Johannesen KM, Hedrich UBS, Masnada S, Rubboli G, Gardella E, et al. Genetic and phenotypic heterogeneity suggest therapeutic implications in SCN2A-related disorders. Brain. 2017;140:1316–36.
Hoyer J, Ekici AB, Endele S, Popp B, Zweier C, Wiesener A, et al. Haploinsufficiency of ARID1B, a member of the SWI/SNF-a chromatin-remodeling complex, is a frequent cause of intellectual disability. Am J Hum Genet. 2012;90:565–72.
Hamdan FF, Daoud H, Piton A, Gauthier J, Dobrzeniecka S, Krebs MO, et al. De novo SYNGAP1 mutations in nonsyndromic intellectual disability and autism. Biol Psychiatry. 2011;69:898–901.
Van Dijck A, Vulto-van Silfhout AT, Cappuyns E, van der Werf IM, Mancini GM, Tzschach A, et al. Clinical presentation of a complex neurodevelopmental disorder caused by mutations in ADNP. Biol Psychiatry. 2019;85:287–97.
Acknowledgements
We are grateful to the families involved in this study. We thank all of the families in SPARK, the SPARK clinical sites, and the SPARK staff. We thank all of the families at the participating SSC sites, as well as the principal investigators A. Beaudet, R. Bernier, J. Constantino, E. Cook, E. Fombonne, D. Geschwind, R. Goin-Ko- chel, E. Hanson, D. Grice, A. Klin, D. Ledbetter, C. Lord, C. Martin, D. Martin, R. Maxim, J. Miles, O. Ousley, K. Pelphrey, B. Peterson, J. Piggot, C. Saulnier, M. State, W. Stone, J. Sutcliffe, C. Walsh, Z. Warren, and E. Wijsman. We appreciate obtaining access to the phenotypic and genetic data on SFARI Base. Approved researchers can obtain the SSC population dataset described in this study (https://www.sfari.org/resource/simons-simplex-collection/) and the SPARK population dataset described in this study (https://www.sfari.org/resource/spark/) by applying at https://base.sfari.org. We thank Hartmut Engels for coordinating data collection and assistance.
Funding
This work was partially supported by the High-Performance Computing Center of Central South University. This work was supported in part by the Bioinformatics Center, Xiangya Hospital, Central South University. Funding: This study was supported by STI 2030-Major Project (no. 2021ZD0201704 to HG); National Natural Science Foundation of China (nos. 82222025 and 81871079 to HG; nos. 82130043, 82330035 and 82361138573 to KX); National Key Research and Development Program of China (no. 2021YFA0805200 to ZH and JT), Hunan Provincial grants (nos. 2023RC1020 and 2023SK2084 to HG; nos. 2021SK1010, 2021SK1010 and 2023SK2114 to KX; no. 2019JJ40408 to SZ), Jiangxi Province Key Research and Development Project (no. 20232BBG70023), and the France Genomic Medicine Plan 2025 (SeqOIA).
Author information
Authors and Affiliations
Contributions
HG, KX, ST, and QZ designed and conceived this study. ST and SL analyzed and interpreted the genomic data. QZ, RZ, YH, BY, and SZ designed and conducted the experiments and analyzed the data. FL, ZH, and JT helped with data interpretation. HG and KX supervised the work. ST, QZ, HG, and KX wrote and revised the manuscript. Other authors including CM, VP, SM, AD, SP, CP, MK, NS, JPT, CR, FT, CP, ADB, JM, XW, XT, SS, DU, JAM, HAS, MMM, AB, XL, SJ and PL contributed and interpreted the genetic and clinical data recruited from an international collaborative network. All authors commented on the manuscript and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
AB and MMM are employees of GeneDx, Inc. All of the remaining authors declare no conflict of interest.
Ethics approval and consent to participate
We strictly follow the Guidelines for Clinical Laboratory Biosafety. All of the experimental procedures involving animals were conducted in accordance with the Institutional Animal Care guidelines of Central South University. All cell lines were verified for authenticity. All patient consents were obtained from study participants or their parents or legal guardians, in line with local institutional review board (IRB) requirements at the time of collection. The IRB of Central South University approved this study (Registration number: IRB #2022-1-3).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tan, S., Zhang, Q., Zhan, R. et al. Monoallelic loss-of-function variants in GSK3B lead to autism and developmental delay. Mol Psychiatry 30, 1952–1965 (2025). https://doi.org/10.1038/s41380-024-02806-z
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41380-024-02806-z
This article is cited by
-
Early-life microbiota dysbiosis as a link between Autism Spectrum Disorder and Parkinson’s Disease
Molecular Psychiatry (2026)