Abstract
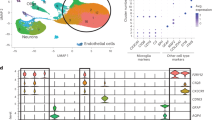
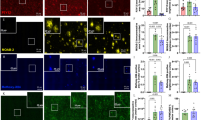
Microglia are central mediators of neuroinflammation in Alzheimer’s disease (AD), yet conserved microglial states and activation trajectories across AD mouse models remain incompletely defined. Here, we constructed a comprehensive Mouse Microglia Atlas (MoMicA) to resolve conserved subtypes, delineate dynamic trajectories, and identify key regulators associated with AD pathology. We integrated ten single-cell and single-nucleus RNA-sequencing datasets from major AD mouse models, yielding 84,764 microglia for harmonized clustering, co-expression network analysis, and pseudotime inference, complemented by immune staining. Across models, AD was characterized by contraction of homeostatic microglia and marked expansion of DAM. MoMicA further delineated refined homeostatic and disease-associated subpopulations, including different homeostatic microglia subclusters and a stepwise progression from homeostatic microglia through activated response and pre-disease-associated states to disease-associated microglia. Network analysis highlighted lipid metabolism and inflammation as dominant AD-related programs and identified Fabp5 as a central hub gene within a DAM-associated lipid module. Multiplex immunofluorescence confirmed that Fabp5 is induced in Clec7a-positive DAM around amyloid plaques in two amyloidosis models. MoMicA therefore provides a valuable resource for dissecting the mechanistic roles of microglia in the onset and progression of AD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The source single-cell transcriptomic data used in this study are publicly accessible through the following accession numbers: Figshare: 19706530; Gene Expression Omnibus (GEO): GSE127892, GSE140510, GSE142267, GSE147495, GSE153895, GSE175546, GSE190607, GSE206114 and GSE224398.
Code availability
Code is available from the corresponding author upon request.
References
GBD 2019 Dementia Forecasting Collaborators. Estimation of the global prevalence of dementia in 2019 and forecasted prevalence in 2050: an analysis for the Global Burden of Disease Study 2019. Lancet Public Health. 2022;7:e105–e25.
2022 Alzheimer’s disease facts and figures. Alzheimers Dement. 2022;18:700–89.
Gustavsson A, Norton N, Fast T, Frölich L, Georges J, Holzapfel D, et al. Global estimates on the number of persons across the Alzheimer’s disease continuum. Alzheimers Dement. 2023;19:658–70.
Frisoni GB, Aho E, Brayne C, Ciccarelli O, Dubois B, Fox NC, et al. Alzheimer’s disease outlook: controversies and future directions. Lancet. 2025;406:1424–42.
Zheng Q, Wang X. Alzheimer’s disease: insights into pathology, molecular mechanisms, and therapy. Protein Cell. 2025;16:83–120.
Scheltens P, De Strooper B, Kivipelto M, Holstege H, Chételat G, Teunissen CE, et al. Alzheimer’s disease. Lancet. 2021;397:1577–90.
Taddei RN. Synapse vulnerability and resilience underlying Alzheimer’s disease. EBioMedicine. 2025;112:105557.
Jucker M, Walker LC. Alzheimer’s disease: From immunotherapy to immunoprevention. Cell. 2023;186:4260–70.
Fox NC, Belder C, Ballard C, Kales HC, Mummery C, Caramelli P, et al. Treatment for Alzheimer’s disease. Lancet. 2025;406:1408–23.
Courade JP, Zetterberg H, Höglinger GU, Dewachter I. The evolving landscape of Alzheimer’s disease therapy: From Aβ to tau. Cell. 2025;188:7337–54.
Leng F, Edison P. Neuroinflammation and microglial activation in Alzheimer disease: where do we go from here? Nat Rev Neurol. 2021;17:157–72.
Jack CR Jr, Andrews JS, Beach TG, Buracchio T, Dunn B, Graf A, et al. Revised criteria for diagnosis and staging of Alzheimer’s disease: Alzheimer’s association workgroup. Alzheimers Dement. 2024;20:5143–69.
Zhu W, Wang Y, Qin M, Zhao Q, Feng L, Li M. Insights into biomarkers of Alzheimer’s disease: from core markers to emerging directions. Aging Dis. 2026;17. https://doi.org/10.14336/AD.2025.0761.
Sharma R, Mei A, Mathew V, Kashpur O, Wallingford MC. Interaction of extraembryonic microglia and neonatal brain development. Exp Neurol. 2022;351:113986.
Depp C, Doman JL, Hingerl M, Xia J, Stevens B. Microglia transcriptional states and their functional significance: context drives diversity. Immunity. 2025;58:1052–67.
Paolicelli RC, Sierra A, Stevens B, Tremblay ME, Aguzzi A, Ajami B, et al. Microglia states and nomenclature: A field at its crossroads. Neuron. 2022;110:3458–83.
Kunkle BW, Grenier-Boley B, Sims R, Bis JC, Damotte V, Naj AC, et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat Genet. 2019;51:414–30.
Jorfi M, Maaser-Hecker A, Tanzi RE. The neuroimmune axis of Alzheimer’s disease. Genome Med. 2023;15:6.
van Olst L, Simonton B, Edwards AJ, Forsyth AV, Boles J, Jamshidi P, et al. Microglial mechanisms drive amyloid-β clearance in immunized patients with Alzheimer’s disease. Nat Med. 2025;31:1604–16.
Hwang B, Lee JH, Bang D. Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp Mol Med. 2018;50:1–14.
Mathys H, Davila-Velderrain J, Peng Z, Gao F, Mohammadi S, Young JZ, et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature. 2019;570:332–7.
Ofengeim D, Giagtzoglou N, Huh D, Zou C, Yuan J. Single-cell RNA sequencing: unraveling the brain one cell at a time. Trends Mol Med. 2017;23:563–76.
Fumagalli L, Nazlie Mohebiany A, Premereur J, Polanco Miquel P, Bijnens B, Van de Walle P, et al. Microglia heterogeneity, modeling and cell-state annotation in development and neurodegeneration. Nat Neurosci. 2025;28:1381–92.
Pettas S, Karagianni K, Kanata E, Chatziefstathiou A, Christoudia N, Xanthopoulos K, et al. Profiling microglia through single-cell RNA sequencing over the course of development, aging, and disease. Cells. 2022;11:2383.
Keren-Shaul H, Spinrad A, Weiner A, Matcovitch-Natan O, Dvir-Szternfeld R, Ulland TK, et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell. 2017;169:1276–90.e17
Martins-Ferreira R, Calafell-Segura J, Leal B, Rodríguez-Ubreva J, Martínez-Saez E, Mereu E, et al. The Human microglia atlas (HuMicA) unravels changes in disease-associated microglia subsets across neurodegenerative conditions. Nat Commun. 2025;16:739.
Dhandapani R, Neri M, Bernhard M, Brzak I, Schweizer T, Rudin S, et al. Sustained Trem2 stabilization accelerates microglia heterogeneity and Aβ pathology in a mouse model of Alzheimer’s disease. Cell Rep. 2022;39:110883.
Sala Frigerio C, Wolfs L, Fattorelli N, Thrupp N, Voytyuk I, Schmidt I, et al. The major risk factors for Alzheimer’s disease: age, sex, and genes modulate the microglia response to Aβ plaques. Cell Rep. 2019;27:1293–306.e6
Zhou Y, Song WM, Andhey PS, Swain A, Levy T, Miller KR, et al. Human and mouse single-nucleus transcriptomics reveal TREM2-dependent and TREM2-independent cellular responses in Alzheimer’s disease. Nat Med. 2020;26:131–42.
Sierksma A, Lu A, Mancuso R, Fattorelli N, Thrupp N, Salta E, et al. Novel Alzheimer risk genes determine the microglia response to amyloid-β but not to TAU pathology. EMBO Mol Med. 2020;12:e10606.
Lau SF, Chen C, Fu WY, Qu JY, Cheung TH, Fu AKY, et al. IL-33-PU.1 transcriptome reprogramming drives functional state transition and clearance activity of microglia in alzheimer’s disease. Cell Rep. 2020;31:107530.
Lee SH, Meilandt WJ, Xie L, Gandham VD, Ngu H, Barck KH, et al. Trem2 restrains the enhancement of tau accumulation and neurodegeneration by β-amyloid pathology. Neuron. 2021;109:1283–301.e6.
Kim DW, Tu KJ, Wei A, Lau AJ, Gonzalez-Gil A, Cao T, et al. Amyloid-beta and tau pathologies act synergistically to induce novel disease stage-specific microglia subtypes. Mol Neurodegener. 2022;17:83.
Karaahmet B, Le L, Mendes MS, Majewska AK, O’Banion MK. Repopulated microglia induce expression of Cxcl13 with differential changes in Tau phosphorylation but do not impact amyloid pathology. J Neuroinflammation. 2022;19:173.
Drieu A, Du S, Storck SE, Rustenhoven J, Papadopoulos Z, Dykstra T, et al. Parenchymal border macrophages regulate the flow dynamics of the cerebrospinal fluid. Nature. 2022;611:585–93.
Park H, Cho B, Kim H, Saito T, Saido TC, Won KJ, et al. Single-cell RNA-sequencing identifies disease-associated oligodendrocytes in male APP NL-G-F and 5XFAD mice. Nat Commun. 2023;14:802.
Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol. 2018;36:411–20.
McGinnis CS, Murrow LM, Gartner ZJ. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 2019;8:329–37.e4
Korsunsky I, Millard N, Fan J, Slowikowski K, Zhang F, Wei K, et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods. 2019;16:1289–96.
Chen Y, Colonna M. Microglia in Alzheimer’s disease at single-cell level. Are there common patterns in humans and mice? J Exp Med. 2021;218:e20202717.
Xu S, Hu E, Cai Y, Xie Z, Luo X, Zhan L, et al. Using clusterProfiler to characterize multiomics data. Nat Protoc. 2024;19:3292–320.
Morabito S, Reese F, Rahimzadeh N, Miyoshi E, Swarup V. hdWGCNA identifies co-expression networks in high-dimensional transcriptomics data. Cell Rep Methods. 2023;3:100498.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Wu X, Liu M, Zhang X, Pan X, Cui X, Jin J, et al. Elucidating microglial heterogeneity and functions in Alzheimer’s disease using single-cell analysis and convolutional neural network disease model construction. Sci Rep. 2024;14:17271.
Oakley H, Cole SL, Logan S, Maus E, Shao P, Craft J, et al. Intraneuronal beta-amyloid aggregates, neurodegeneration, and neuron loss in transgenic mice with five familial Alzheimer’s disease mutations: potential factors in amyloid plaque formation. J Neurosci. 2006;26:10129–40.
Sturchler-Pierrat C, Abramowski D, Duke M, Wiederhold KH, Mistl C, Rothacher S, et al. Two amyloid precursor protein transgenic mouse models with Alzheimer disease-like pathology. Proc Natl Acad Sci USA. 1997;94:13287–92.
Yoshiyama Y, Higuchi M, Zhang B, Huang SM, Iwata N, Saido TC, et al. Synapse loss and microglial activation precede tangles in a P301S tauopathy mouse model. Neuron. 2007;53:337–51.
Oddo S, Caccamo A, Shepherd JD, Murphy MP, Golde TE, Kayed R, et al. Triple-transgenic model of Alzheimer’s disease with plaques and tangles: intracellular Abeta and synaptic dysfunction. Neuron. 2003;39:409–21.
Hickman S, Izzy S, Sen P, Morsett L, El Khoury J. Microglia in neurodegeneration. Nat Neurosci. 2018;21:1359–69.
Olah M, Menon V, Habib N, Taga MF, Ma Y, Yung CJ, et al. Single cell RNA sequencing of human microglia uncovers a subset associated with Alzheimer’s disease. Nat Commun. 2020;11:6129.
Rangaraju S, Dammer EB, Raza SA, Rathakrishnan P, Xiao H, Gao T, et al. Identification and therapeutic modulation of a pro-inflammatory subset of disease-associated-microglia in Alzheimer’s disease. Mol Neurodegener. 2018;13:24.
Zeng H, Huang J, Zhou H, Meilandt WJ, Dejanovic B, Zhou Y, et al. Integrative in situ mapping of single-cell transcriptional states and tissue histopathology in a mouse model of Alzheimer’s disease. Nat Neurosci. 2023;26:430–46.
Baik SH, Kang S, Lee W, Choi H, Chung S, Kim JI, et al. A breakdown in metabolic reprogramming causes microglia dysfunction in Alzheimer’s disease. Cell Metab. 2019;30:493–507.e6.
Sun N, Victor MB, Park YP, Xiong X, Scannail AN, Leary N, et al. Human microglial state dynamics in Alzheimer’s disease progression. Cell. 2023;186:4386–403.e29.
Matera A, Compagnion A-C, Pedicone C, Kotah JM, Ivanov A, Monsorno K, et al. Microglial lipid phosphatase SHIP1 limits complement-mediated synaptic pruning in the healthy developing hippocampus. Immunity. 2024;58:197–217.e13
Chou V, Pearse RV, Aylward AJ, Ashour N, Taga M, Terzioglu G, et al. INPP5D regulates inflammasome activation in human microglia. Nat Commun. 2023;14:7552.
Peng W, Xie Y, Liao C, Bai Y, Wang H, Li C. Spatiotemporal patterns of gliosis and neuroinflammation in presenilin 1/2 conditional double knockout mice. Front Aging Neurosci. 2022;14:966153.
Wang C, Fan L, Khawaja RR, Liu B, Zhan L, Kodama L, et al. Microglial NF-κB drives tau spreading and toxicity in a mouse model of tauopathy. Nat Commun. 2022;13:1969.
Lan Y, Zhang X, Liu S, Guo C, Jin Y, Li H, et al. Fate mapping of Spp1 expression reveals age-dependent plasticity of disease-associated microglia-like cells after brain injury. Immunity. 2024;57:349–63.e9
March-Diaz R, Lara-Ureña N, Romero-Molina C, Heras-Garvin A, Ortega-de San Luis C, Alvarez-Vergara MI, et al. Hypoxia compromises the mitochondrial metabolism of Alzheimer’s disease microglia via HIF1. Nat Aging. 2021;1:385–99.
Hou Y, Wei D, Zhang Z, Guo H, Li S, Zhang J, et al. FABP5 controls macrophage alternative activation and allergic asthma by selectively programming long-chain unsaturated fatty acid metabolism. Cell Rep. 2022;41:111668.
Acknowledgements
This work was supported by the National Key R&D Program of China (STI2030-Major Projects 2021ZD0202400), Natural Science Foundation of Chongqing, China (CSTB2025NSCQ-GPX0386), the National Key R&D Program of China (ST12030-Major Projects 2021zD0200600), the National Natural Science Foundation Project of China (82201688, 82571748, 82471545, 82401784, 32400850, 82401523, 82501836, 82501469, 82501837), National Reserve Talent Project in the Health and Wellness Sector of Chongqing (HBRC202410, HBRC202417), China Postdoctoral Science Foundation (2024MD754023, 2025MD774171, 2025T180580), the Key Project of the Natural Science Foundation of Chongqing (Chongqing Science and Technology Development Foundation) under Grant No. 2024NSCQ-KJFZZDX0005, and the New Chongqing Youth Innovation Talent Project (Life and Health) under Grant No.2024NSCQ-qncxX0029.
Author information
Authors and Affiliations
Contributions
Designed the experiments: PZ. Performed the experiments: PL and TFS. Analyzed the data: JY, HRL, WWM, XYZ, RHX, JW, YH, YFL, MHY, JPZ, XMT, XDS, and HSZ. Drafted the paper: PL and TFS. Revised the paper for intellectual content: PZ and MLW.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The original studies of single-cell transcriptomic data had obtained the necessary ethical approval for sample collection. This study obtained ethical clearance from the Ethical Committee of Chongqing Medical University (Chongqing, China; IACUC-CQMU-2024-0710).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, P., Sun, T., Yang, J. et al. Comprehensive single-cell transcriptomic atlas of microglia in Alzheimer’s disease mouse models. Mol Psychiatry (2026). https://doi.org/10.1038/s41380-026-03529-z
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41380-026-03529-z