Abstract
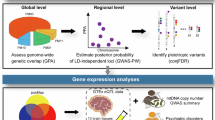
Psychiatric disorders, including bipolar disorder (BD), major depressive disorder (MDD), and schizophrenia (SCZ), share substantial genetic overlap. We conducted a cross-ancestry multivariate genome-wide association study (GWAS) integrating European and East Asian populations to uncover shared genetic underpinnings. Our analyses identified 403 loci associated with shared polygenic liability to psychiatric disorders, including 88 novel regions. Cross-ancestry fine-mapping highlighted robust shared signals, notably at VRK2 (rs7596038), consistently significant across ancestries. Gene prioritization revealed 90 high-confidence candidate genes enriched in neurodevelopmental pathways. Single-nucleus RNA sequencing implicated excitatory neurons and astrocytes as key cellular contexts, emphasizing NCAM1–FGFR1 and NEGR1–NEGR1 signaling pathways. Mendelian randomization analyses provided causal evidence linking shared genetic liability to structural brain alterations, particularly in regions crucial for emotion and cognition. Polygenic risk scores derived from shared genetic liability substantially enhanced predictive accuracy for BD and SCZ, demonstrating strong trans-ancestry validity. These results advance understanding of shared genetic architecture in psychiatric disorders, highlighting potential therapeutic targets and emphasizing the critical importance of diverse ancestry studies in precision psychiatry.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The summary statistics of BD, MDD, and SCZ in two ancestries are available here: https://pgc.unc.edu/for-researchers/download-results/. The summary statistics of eQTL are available from GTEx: https://yanglab.westlake.edu.cn/software/smr/#eQTLsummarydata. The snRNA-seq data is available from https://www.ncbi.nlm.nih.gov/geo/. The gene specificity data per cell type for enrichment analysis is available from https://www.nature.com/articles/s41467-024-55611-1#Sec50. The summary statistics of IDPs are available from here: https://www.ebi.ac.uk/gwas/. The reference panel of two ancestries are available from 1000 Genomes Project: https://www.internationalgenome.org/data/. The UK Biobank data are available to any bona fide researcher following application: https://www.ukbiobank.ac.uk/enable-your-research/apply-for-access. The data of CLASS-BD are available from the corresponding author upon reasonable request.
Code availability
Packages including ‘LAVA’ (version 1.0), ‘GenomicSEM’ (version 0.05), ‘MeSuSiE’ (version 1.0), ‘TwoSampleMR’ (version 0.6.6) and ‘survival’ (version 3.8-3) in R version 4.3.3 were used to perform local genetic correlation, gSEM model, cross-ancestry fine-mapping, Mendelian randomization, and survival analysis. Seurat (version 5.3.0), CellChat (version 2.1.0) in R were used to perform snRNA-seq data processing and cell-cell communication analysis. MiXeR (version 1.2) was used to estimate genetic overlapping. METAL was used to perform GWAS meta-analysis. LDSC (version 1.0.1) were used to perform genetic correlation analysis and enrichment analysis. mBAT (GCTA, version 1.94.1), FUMA, SMR (version 1.4.0), TWAS, PoPS, and PsyOPS were used to identify the pleiotropic genes. DBSLMM (version 1.0), PRS-CSx and PLINK2 were used to estimate in- and cross-ancestry PRS.
References
GBD 2019 Mental Disorders Collaborators. Global, regional, and national burden of 12 mental disorders in 204 countries and territories, 1990-2019: A systematic analysis for the Global Burden of Disease Study 2019. Lancet Psychiatry. 2022;9:137–50.
The L. Ensuring care for people with depression. Lancet. 2022;399:885.
Casella CB, Kousoulis AA, Kohrt BA, Bantjes J, Kieling C, Cuijpers P, et al. Data gaps in prevalence rates of mental health conditions around the world: a retrospective analysis of nationally representative data. Lancet Glob Health. 2025;13:e879–e887.
Li HJ, Zhang C, Hui L, Zhou DS, Li Y, Zhang CY, et al. Novel risk loci associated with genetic risk for bipolar disorder among Han Chinese individuals: A genome-wide association study and meta-analysis. JAMA Psychiatry. 2021;78:320–30.
Liu J, Li S, Li X, Li W, Yang Y, Guo S, et al. Genome-wide association study followed by trans-ancestry meta-analysis identify 17 new risk loci for schizophrenia. BMC Med. 2021;19:177.
Li W, Chen R, Feng L, Dang X, Liu J, Chen T, et al. Genome-wide meta-analysis, functional genomics and integrative analyses implicate new risk genes and therapeutic targets for anxiety disorders. Nat Hum Behav. 2024;8:361–79.
Jung JY, Ahn Y, Park JW, Jung K, Kim S, Lim S, et al. Polygenic overlap between subjective well-being and psychiatric disorders and cross-ancestry validation. Nat Hum Behav. 2025;9:1272–82.
Lam M, Chen CY, Li Z, Martin AR, Bryois J, Ma X, et al. Comparative genetic architectures of schizophrenia in East Asian and European populations. Nat Genet. 2019;51:1670–8.
Wu Y, Cao H, Baranova A, Huang H, Li S, Cai L, et al. Multi-trait analysis for genome-wide association study of five psychiatric disorders. Transl Psychiatry. 2020;10:209.
Ferreira MA, Purcell SM. A multivariate test of association. Bioinformatics. 2009;25:132–3.
Kim S, Xing EP. Statistical estimation of correlated genome associations to a quantitative trait network. PLoS Genet. 2009;5:e1000587.
Shao K, Luo Z, Huang P, Yang S. ProteoNexus: an integrative database to characterize genetic architecture, estimate mediation effects, and construct and evaluate prediction models of the plasma proteome. Nucleic Acids Res. 2026;54:D1222–d1233.
Turley P, Walters RK, Maghzian O, Okbay A, Lee JJ, Fontana MA, et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat Genet. 2018;50:229–37.
Grotzinger AD, Rhemtulla M, de Vlaming R, Ritchie SJ, Mallard TT, Hill WD, et al. Genomic structural equation modelling provides insights into the multivariate genetic architecture of complex traits. Nat Hum Behav. 2019;3:513–25.
Ray D, Chatterjee N. A powerful method for pleiotropic analysis under composite null hypothesis identifies novel shared loci between type 2 diabetes and prostate cancer. PLoS Genet. 2020;16:e1009218.
Mallard TT, Linnér RK, Grotzinger AD, Sanchez-Roige S, Seidlitz J, Okbay A, et al. Multivariate GWAS of psychiatric disorders and their cardinal symptoms reveal two dimensions of cross-cutting genetic liabilities. Cell Genom. 2022;2:100140.
Harkness KL, Stewart JG. Stress and diathesis–stress models in depression. APA handbook of depression: Classification, co-occurring conditions, and etiological processes. 1. Washington, DC, US: American Psychological Association; 2026. p. 441–58.
Walker EF, Diforio D. Schizophrenia: A neural diathesis-stress model. Psychol Rev. 1997;104:667–85.
Colodro-Conde L, Couvy-Duchesne B, Zhu G, Coventry WL, Byrne EM, Gordon S, et al. A direct test of the diathesis-stress model for depression. Mol Psychiatry. 2018;23:1590–6.
Levey DF, Galimberti M, Deak JD, Wendt FR, Bhattacharya A, Koller D, et al. Multi-ancestry genome-wide association study of cannabis use disorder yields insight into disease biology and public health implications. Nat Genet. 2023;55:2094–103.
Friligkou E, Løkhammer S, Cabrera-Mendoza B, Shen J, He J, Deiana G, et al. Gene discovery and biological insights into anxiety disorders from a large-scale multi-ancestry genome-wide association study. Nat Genet. 2024;56:2036–45.
Karlsson Linnér R, Mallard TT, Barr PB, Sanchez-Roige S, Madole JW, Driver MN, et al. Author Correction: Multivariate analysis of 1.5 million people identifies genetic associations with traits related to self-regulation and addiction. Nat Neurosci. 2025;28:213.
Akimova ET, Wolfram T, Ding X, Tropf FC, Mills MC. Polygenic prediction of occupational status GWAS elucidates genetic and environmental interplay in intergenerational transmission, careers and health in UK Biobank. Nat Hum Behav. 2025;9:391–405.
Ruscio J, Roche B. Determining the number of factors to retain in an exploratory factor analysis using comparison data of known factorial structure. Psychol Assess. 2012;24:282–92.
Kaiser HF. The application of electronic computers to factor analysis. Educ Psychol Meas. 1960;20:141–51.
Brown BC, Ye CJ, Price AL, Zaitlen N. Transethnic genetic-correlation estimates from summary statistics. Am J Hum Genet. 2016;99:76–88.
Cao X, Jiang M, Guan Y, Li S, Duan C, Gong Y, et al. Trans-ancestry GWAS identifies 59 loci and improves risk prediction and fine-mapping for kidney stone disease. Nat Commun. 2025;16:3473.
Li Y, Dang X, Chen R, Teng Z, Wang J, Li S, et al. Cross-ancestry genome-wide association study and systems-level integrative analyses implicate new risk genes and therapeutic targets for depression. Nat Hum Behav. 2025;9:806–23.
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet. 2014;46:310–5.
Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012;22:1790–7.
Kolberg L, Raudvere U, Kuzmin I, Vilo J, Peterson H. gprofiler2 – an R package for gene list functional enrichment analysis and namespace conversion toolset g:Profiler. F1000Res. 2020;9:ELIXIR-709.
Fröhlich AS, Gerstner N, Gagliardi M, Ködel M, Yusupov N, Matosin N, et al. Single-nucleus transcriptomic profiling of human orbitofrontal cortex reveals convergent effects of aging and psychiatric disease. Nat Neurosci. 2024;27:2021–32.
Gerstner N, Fröhlich AS, Matosin N, Gagliardi M, Cruceanu C, Ködel M, et al. Contrasting genetic predisposition and diagnosis in psychiatric disorders: A multi-omic single-nucleus analysis of the human OFC. Sci Adv. 2025;11:eadq2290.
Zhang YQ, Zhang Q, Yang Y, Yu LL, Fan NL, Wu Y, et al. Elevated NEGR1 in brain induces anxiety or depression-like phenotypes and synaptic dysfunction. Mol Psychiatry. 2025;30:4627–40.
Richards AL, Cardno A, Harold G, Craddock NJ, Di Florio A, Jones L, et al. Genetic liabilities differentiating bipolar disorder, schizophrenia, and major depressive disorder, and phenotypic heterogeneity in bipolar disorder. JAMA Psychiatry. 2022;79:1032–9.
Grotzinger AD, Mallard TT, Akingbuwa WA, Ip HF, Adams MJ, Lewis CM, et al. Genetic architecture of 11 major psychiatric disorders at biobehavioral, functional genomic and molecular genetic levels of analysis. Nat Genet. 2022;54:548–59.
Ahangari M, Kirkpatrick R, Nguyen T-H, Gillespie N, Consortium ISG, Kendler KS, et al. Examining the source of increased bipolar disorder and major depressive disorder common risk variation burden in multiplex schizophrenia families. Schizophr. 2022;8:106.
Zerbin‐Rüdin E, Kendler KS. Ernst Rüdin (1874-1952) and his genealogic-demographic department in Munich (1917-1986): an introduction to their family studies of schizophrenia. Am J Med Genet. 1996;67:332–7.
ENCODE Project Consortium, A. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489:57–74.
Li M, Yue W. VRK2, a candidate gene for psychiatric and neurological disorders. Mol Neuropsychiatry. 2018;4:119–33.
Steinberg S, de Jong S, Andreassen OA, Werge T, Børglum AD, Mors O, et al. Common variants at VRK2 and TCF4 conferring risk of schizophrenia. Hum Mol Genet. 2011;20:4076–81.
Xiao X, Luo XJ, Chang H, Liu Z, Li M. Evaluation of European schizophrenia GWAS Loci in Asian populations via comprehensive meta-analyses. Mol Neurobiol. 2017;54:4071–80.
Chang H, Zhang C, Xiao X, Pu X, Liu Z, Wu L, et al. Further evidence of VRK2 rs2312147 associated with schizophrenia. World J Biol Psychiatry. 2016;17:457–66.
Hyde CL, Nagle MW, Tian C, Chen X, Paciga SA, Wendland JR, et al. Identification of 15 genetic loci associated with risk of major depression in individuals of European descent. Nat Genet. 2016;48:1031–6.
Stahl EA, Breen G, Forstner AJ, McQuillin A, Ripke S, Trubetskoy V, et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat Genet. 2019;51:793–803.
Nagel M, Jansen PR, Stringer S, Watanabe K, de Leeuw CA, Bryois J, et al. Meta-analysis of genome-wide association studies for neuroticism in 449,484 individuals identifies novel genetic loci and pathways. Nat Genet. 2018;50:920–7.
Howes OD, Onwordi EC. The synaptic hypothesis of schizophrenia version III: A master mechanism. Mol Psychiatry. 2023;28:1843–56.
Pardiñas AF, Holmans P, Pocklington AJ, Escott-Price V, Ripke S, Carrera N, et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat Genet. 2018;50:381–9.
Zhou J, Ma C, Wang K, Li X, Zhang H, Chen J, et al. Rare and common variants analysis of the EMB gene in patients with schizophrenia. BMC Psychiatry. 2020;20:135.
Newton S, Aguilar C, Bunton-Stasyshyn RK, Flook M, Stewart M, Marcotti W, et al. Absence of Embigin accelerates hearing loss and causes sub-viability, brain and heart defects in C57BL/6N mice due to interaction with Cdh23(ahl). iScience. 2023;26:108056.
Wilson MC, Meredith D, Fox JE, Manoharan C, Davies AJ, Halestrap AP. Basigin (CD147) is the target for organomercurial inhibition of monocarboxylate transporter isoforms 1 and 4: the ancillary protein for the insensitive MCT2 is EMBIGIN (gp70). J Biol Chem. 2005;280:27213–21.
Bledsoe X, Gamazon ER. A transcriptomic atlas of the human brain reveals genetically determined aspects of neuropsychiatric health. Am J Hum Genet. 2024;111:1559–72.
Kiselyov VV, Skladchikova G, Hinsby AM, Jensen PH, Kulahin N, Soroka V, et al. Structural basis for a direct interaction between FGFR1 and NCAM and evidence for a regulatory role of ATP. Structure. 2003;11:691–701.
Stark C, Breitkreutz BJ, Reguly T, Boucher L, Breitkreutz A, Tyers M. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 2006;34:D535–539.
Životić M, Tampe B, Müller G, MÃMüller C, Lipkovski A, Xu X, et al. Modulation of NCAM/FGFR1 signaling suppresses EMT program in human proximal tubular epithelial cells. PLoS ONE. 2018;13:e0206786.
Shukrun R, Golan H, Caspi R, Pode-Shakked N, Pleniceanu O, Vax E, et al. NCAM1/FGF module serves as a putative pleuropulmonary blastoma therapeutic target. Oncogenesis. 2019;8:48.
Habicher J, Sanvido I, Buehler A, Sartori S, Piccoli G, Carl M. The risk genes for neuropsychiatric disorders negr1 and opcml are expressed throughout zebrafish brain development. Genes (Basel). 2024;15:363.
Pischedda F, Szczurkowska J, Daniela Cirnaru M, Cancedda L, Piccoli G The role of Negr1 in cortical development via NCAM-FGFR2 signaling. LID - P38. (2193-1801 (Electronic)).
Lipsky RK, Garrett ME, Dennis MF, Hauser MA, Beckham JC, Ashley-Koch AE, et al. Impact of traumatic life events and polygenic risk scores for major depression and posttraumatic stress disorder on Iraq/Afghanistan Veterans. J Psychiatr Res. 2023;158:15–19.
Thorp JG, Gerring ZF, Colodro-Conde L, Byrne EM, Medland SE, Middeldorp CM, et al. The association between trauma exposure, polygenic risk and individual depression symptoms. Psychiatry Res. 2023;321:115101.
Vilhjálmsson BJ, Yang J, Finucane HK, Gusev A, Lindström S, Ripke S, et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am J Hum Genet. 2015;97:576–92.
Ge T, Chen CY, Ni Y, Feng YA, Smoller JW. Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat Commun. 2019;10:1776.
Zhao Z, Fritsche LG, Smith JA, Mukherjee B, Lee S. The construction of cross-population polygenic risk scores using transfer learning. Am J Hum Genet. 2022;109:1998–2008.
Meng X, Navoly G, Giannakopoulou O, Levey DF, Koller D, Pathak GA, et al. Multi-ancestry genome-wide association study of major depression aids locus discovery, fine mapping, gene prioritization and causal inference. Nat Genet. 2024;56:222–33.
Guo X, Feng Y, Ji X, Jia N, Maimaiti A, Lai J, et al. Shared genetic architecture and bidirectional clinical risks within the psycho-metabolic nexus. EBioMedicine. 2025;111:105530.
O’Connell KS, Koromina M, van der Veen T, Boltz T, David FS, Yang JMK, et al. Genomics yields biological and phenotypic insights into bipolar disorder. Nature. 2025;639:968–75.
Palmer LJ. UK Biobank: Bank on it. Lancet. 2007;369:1980–2.
Cao C, Zhang S, Wang J, Tian M, Ji X, Huang D, et al. PGS-Depot: a comprehensive resource for polygenic scores constructed by summary statistics based methods. Nucleic Acids Res. 2024;52:D963–d971.
Yang S, Zhou X. Accurate and scalable construction of polygenic scores in large biobank data sets. Am J Hum Genet. 2020;106:679–93.
Yang S, Ye X, Ji X, Li Z, Tian M, Huang P, et al. PGSFusion streamlines polygenic score construction and epidemiological applications in biobank-scale cohorts. Genome Med. 2025;17:77.
Li Z, Lai J, Zhang P, Ding J, Jiang J, Liu C, et al. Multi-omics analyses of serum metabolome, gut microbiome and brain function reveal dysregulated microbiota-gut-brain axis in bipolar depression. Mol Psychiatry. 2022;27:4123–35.
Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Cong PK, Bai WY, Li JC, Yang MY, Khederzadeh S, Gai SR, et al. Genomic analyses of 10,376 individuals in the Westlake BioBank for Chinese (WBBC) pilot project. Nat Commun. 2022;13:2939.
Greenberg A, Reynolds K, McNulty MT, Sampson MG, Kang HM, Lee D Accurate cross-platform GWAS analysis via two-stage imputation. medRxiv: 2024.2004.2019.24306081 [Preprint] 2024. https://www.medrxiv.org/content/10.1101/2024.04.19.24306081v1.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Frei O, Holland D, Smeland OB, Shadrin AA, Fan CC, Maeland S, et al. Bivariate causal mixture model quantifies polygenic overlap between complex traits beyond genetic correlation. Nat Commun. 2019;10:2417.
Holland D, Frei O, Desikan R, Fan CC, Shadrin AA, Smeland OB, et al. Beyond SNP heritability: Polygenicity and discoverability of phenotypes estimated with a univariate Gaussian mixture model. PLoS Genet. 2020;16:e1008612.
Liu M, Wang L, Zhang Y, Dong H, Wang C, Chen Y, et al. Investigating the shared genetic architecture between depression and subcortical volumes. Nat Commun. 2024;15:7647.
Werme J, van der Sluis S, Posthuma D, de Leeuw CA. An integrated framework for local genetic correlation analysis. Nat Genet. 2022;54:274–82.
Bulik-Sullivan BK, Loh PR, Finucane HK, Ripke S, Yang J, Patterson N, et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:291–5.
Schoeler T, Baldwin J, Allegrini A, Barkhuizen W, McQuillin A, Pirastu N, et al. Novel biological insights into the common heritable liability to substance involvement: A multivariate genome-wide association study. Biol Psychiatry. 2023;93:524–35.
Willer CJ, Li Y, Abecasis GR. METAL: Fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
Gao B, Zhou X. MESuSiE enables scalable and powerful multi-ancestry fine-mapping of causal variants in genome-wide association studies. Nat Genet. 2024;56:170–9.
Li A, Liu S, Bakshi A, Jiang L, Chen W, Zheng Z, et al. mBAT-combo: A more powerful test to detect gene-trait associations from GWAS data. Am J Hum Genet. 2023;110:30–43.
Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Gusev A, Ko A, Shi H, Bhatia G, Chung W, Penninx BW, et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet. 2016;48:245–52.
Zhu Z, Zhang F, Hu H, Bakshi A, Robinson MR, Powell JE, et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet. 2016;48:481–7.
Weeks EM, Ulirsch JC, Cheng NY, Trippe BL, Fine RS, Miao J, et al. Leveraging polygenic enrichments of gene features to predict genes underlying complex traits and diseases. Nat Genet. 2023;55:1267–76.
Wainberg M, Merico D, Keller MC, Fauman EB, Tripathy SJ. Predicting causal genes from psychiatric genome-wide association studies using high-level etiological knowledge. Mol Psychiatry. 2022;27:3095–106.
Jin S, Guerrero-Juarez CF, Zhang L, Chang I, Ramos R, Kuan CH, et al. Inference and analysis of cell-cell communication using CellChat. Nat Commun. 2021;12:1088.
Finucane HK, Bulik-Sullivan B, Gusev A, Trynka G, Reshef Y, Loh PR, et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat Genet. 2015;47:1228–35.
Bryois J, Skene NG, Hansen TF, Kogelman LJA, Watson HJ, Liu Z, et al. Genetic identification of cell types underlying brain complex traits yields insights into the etiology of Parkinson’s disease. Nat Genet. 2020;52:482–93.
Yao S, Harder A, Darki F, Chang YW, Li A, Nikouei K, et al. Connecting genomic results for psychiatric disorders to human brain cell types and regions reveals convergence with functional connectivity. Nat Commun. 2025;16:395.
Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, et al. The MR-Base platform supports systematic causal inference across the human phenome. eLife. 2018;7:e34408.
Fu J, Zhang Q, Wang J, Wang M, Zhang B, Zhu W, et al. Cross-ancestry genome-wide association studies of brain imaging phenotypes. Nat Genet. 2024;56:1110–20.
Ruan Y, Lin YF, Feng YA, Chen CY, Lam M, Guo Z, et al. Improving polygenic prediction in ancestrally diverse populations. Nat Genet. 2022;54:573–80.
Yang S, Zhou X. PGS-server: accuracy, robustness and transferability of polygenic score methods for biobank scale studies. Brief Bioinform. 2022;23:bbac039.
Acknowledgements
We thank the GWAS Catalog, the Psychiatric Genomics Consortium, 1000 Genomes Project, and UK Biobank (Application 144904) for access to genome-wide association study data, and all research participants who provided DNA samples for these studies.
Funding
The study was supported by funding from the National Natural Science Foundation of China (82571735, 82173585), Key R&D Program of Zhejiang Province (2024C03098, 2025C02109), and The National Key Research and Development Program of China (2023YFC2506200, 2023YFC2506203).
Author information
Authors and Affiliations
Contributions
S.H. and S.Y. conceived and designed the study. Y.F., N.J., and P.H. analyzed the data. Y.F., N.J., and P.H. interpreted the findings. Y.F., S.Y., and S.H. drafted or substantively revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Ethical approval of UK Biobank study was granted by the National Health Service National Research Ethics Service (reference 11/NW/0382). Ethical approval of Chinese Longitudinal and Systematic Study of Bipolar Disorder was granted by the Medical Research Ethics Committee, and all participants provided written informed consent (Approval number: 2017-397). Written informed consent was secured from all participants and their legal guardians before they were enrolled in the study. It was a single-center study conducted at the First Affiliated Hospital, Zhejiang University School of Medicine (Clinical trial registration number: NCT05480150). All methods were performed in accordance with the relevant guidelines and regulations.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feng, Y., Jia, N., Huang, P. et al. Cross-ancestry genetic architecture reveals shared biological pathways of major psychiatric disorders. Mol Psychiatry (2026). https://doi.org/10.1038/s41380-026-03541-3
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41380-026-03541-3