Abstract
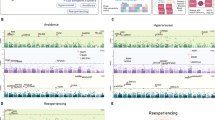
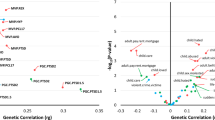
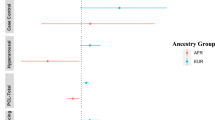
Biomarkers can potentially improve the diagnosis, monitoring, and treatment of posttraumatic stress disorder (PTSD). However, PTSD biomarkers that are scalable and easily integrated into real-world clinical settings have not been identified. The analysis was conducted between June to November 2024 using genomic samples and laboratory test results recorded in the Mass General Brigham (MGB) Health System. The analysis included 23,743 European ancestry participants from the nested MGB Biobank study. The first exposure was polygenic risk score (PRS) for PTSD, calculated using the largest available European ancestry genome-wide association study (GWAS), employing a Bayesian polygenic scoring method. The second exposure was a clinical diagnosis of PTSD, determined by the presence of two or more instances of PTSD-related diagnostic codes in the longitudinal electronic health records (EHR). The primary outcomes were the inverse normal quantile transformed, median lab values of 241 laboratory traits with non-zero h2SNP estimates. Sixteen unique laboratory traits across the cardiometabolic, hematologic, hepatic, and immune systems were implicated in both genomic and phenotypic lab-wide association scans (LabWAS). Two-sample Mendelian randomization analyses provided evidence of potential unidirectional causal effects of PTSD liability on hepatic (decreased albumin and total bilirubin), cardiometabolic (decreased HDL cholesterol and increased VLDL cholesterol), and hematologic (decreased mean platelet volume) markers. These findings demonstrate the potential of a triangulation approach to uncover scalable and clinically relevant biomarkers for PTSD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
MGB Biobank data were accessed under a protocol approved by the MGB Institutional Review Board for the current study (# 2019P003696) and are not publicly available due to restrictions on the data.
References
Katrinli S, Oliveira NCS, Felger JC, Michopoulos V, Smith AK. The role of the immune system in posttraumatic stress disorder. Transl Psychiatry. 2022;12:313.
Gottesman O, Kuivaniemi H, Tromp G, Faucett WA, Li R, Manolio TA, et al. The Electronic Medical Records and Genomics (eMERGE) network: past, present, and future. Genet Med. 2013;15:761–71.
Dennis JK, Sealock JM, Straub P, Lee YH, Hucks D, Actkins K, et al. Clinical laboratory test-wide association scan of polygenic scores identifies biomarkers of complex disease. Genome Med. 2021;13:6.
Sealock JM, Lee YH, Moscati A, Venkatesh S, Voloudakis G, Straub P, et al. Use of the PsycheMERGE network to investigate the association between depression polygenic scores and white blood cell count. JAMA Psychiatry. 2021;78:1365–74.
Wendt FR, Pathak GA, Singh K, Stein MB, Koenen KC, Krystal JH, et al. Sex-specific genetic and transcriptomic liability to neuroticism. Biol Psychiatry. 2023;93:243–52.
Pathak GA, Singh K, Choi KW, Fang Y, Kouakou MR, Lee YH, et al. Genetic liability to posttraumatic stress disorder symptoms and its association with cardiometabolic and respiratory outcomes. JAMA Psychiatry. 2024;81:34–44.
Linnstaedt SD, Zannas AS, McLean SA, Koenen KC, Ressler KJ. Literature review and methodological considerations for understanding circulating risk biomarkers following trauma exposure. Mol Psychiatry. 2020;25:1986–99.
Boutin NT, Schecter SB, Perez EF, Tchamitchian NS, Cerretani XR, Gainer VS, et al. The evolution of a large Biobank at mass general brigham. J Pers Med. 2022;12:1323.
Ge T, Chen C-Y, Ni Y, Feng Y-CA, Smoller JW. Polygenic prediction via bayesian regression and continuous shrinkage priors. Nat Commun. 2019;10:1776.
Nievergelt CM, Maihofer AX, Atkinson EG, Chen C-Y, Choi KW, Coleman JRI, et al. Genome-wide association analyses identify 95 risk loci and provide insights into the neurobiology of post-traumatic stress disorder. Nat Genet. 2024;56:792–808.
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4:7.
Denny JC, Ritchie MD, Basford MA, Pulley JM, Bastarache L, Brown-Gentry K, et al. PheWAS: demonstrating the feasibility of a phenome-wide scan to discover gene-disease associations. Bioinformatics. 2010;26:1205–10.
Wei W-Q, Teixeira PL, Mo H, Cronin RM, Warner JL, Denny JC. Combining billing codes, clinical notes, and medications from electronic health records provides superior phenotyping performance. J Am Med Inform Assoc. 2016;23:e20–7.
Zheutlin AB, Dennis J, Karlsson Linnér R, Moscati A, Restrepo N, Straub P, et al. Penetrance and pleiotropy of polygenic risk scores for schizophrenia in 106,160 patients across four health care systems. Am J Psychiatry. 2019;176:846–55.
Yang J, Lee SH, Goddard ME, Visscher PM. GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet. 2011;88:76–82.
Kim GS, Smith AK, Xue F, Michopoulos V, Lori A, Armstrong DL, et al. Methylomic profiles reveal sex-specific differences in leukocyte composition associated with post-traumatic stress disorder. Brain Behav Immun. 2019;81:280–91.
Ponomareva OY, Ressler KJ. Genomic factors underlying sex differences in trauma-related disorders. Neurobiol Stress. 2021;14:100330.
Department of Veterans Affairs and Department of Defense. VA/DOD clinical practice guideline for the management of posttraumatic stress disorder and acute stress disorder. 2023. https://www.healthquality.va.gov/guidelines/MH/ptsd/.
Gramaglia C, Gambaro E, Bartolomei G, Camera P, Chiarelli-Serra M, Lorenzini L, et al. Increased risk of metabolic syndrome in antidepressants users: a mini review. Front Psychiatry. 2018;9:621 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6279880/.
Grundy A, Cotterchio M, Kirsh VA, Kreiger N. Associations between anxiety, depression, antidepressant medication, obesity and weight gain among Canadian women. PLoS One. 2014;9:e99780.
Graham SE, Clarke SL, Wu K-HH, Kanoni S, Zajac GJM, Ramdas S, et al. The power of genetic diversity in genome-wide association studies of lipids. Nature. 2021;600:675–9.
Richardson TG, Leyden GM, Wang Q, Bell JA, Elsworth B, Davey Smith G, et al. Characterising metabolomic signatures of lipid-modifying therapies through drug target mendelian randomisation. PLoS Biol. 2022;20:e3001547.
Lagou V, Jiang L, Ulrich A, Zudina L, González KSG, Balkhiyarova Z, et al. GWAS of random glucose in 476,326 individuals provide insights into diabetes pathophysiology, complications and treatment stratification. Nat Genet. 2023;55:1448–61.
Hoffmann TJ, Theusch E, Haldar T, Ranatunga DK, Jorgenson E, Medina MW, et al. A large electronic-health-record-based genome-wide study of serum lipids. Nat Genet. 2018;50:401–13.
Sakaue S, Kanai M, Tanigawa Y, Karjalainen J, Kurki M, Koshiba S, et al. A cross-population atlas of genetic associations for 220 human phenotypes. Nat Genet. 2021;53:1415–24.
Revez JA, Lin T, Qiao Z, Xue A, Holtz Y, Zhu Z, et al. Genome-wide association study identifies 143 loci associated with 25 hydroxyvitamin D concentration. Nat Commun. 2020;11:1647.
Pazoki R, Evangelou E, Mosen-Ansorena D, Pinto RC, Karaman I, Blakeley P, et al. GWAS for urinary sodium and potassium excretion highlights pathways shared with cardiovascular traits. Nat Commun. 2019;10:3653.
Chen M-H, Raffield LM, Mousas A, Sakaue S, Huffman JE, Moscati A, et al. Trans-ethnic and ancestry-specific blood-cell genetics in 746,667 individuals from 5 global populations. Cell. 2020;182:1198–1213.e14.
Vuckovic D, Bao EL, Akbari P, Lareau CA, Mousas A, Jiang T, et al. The polygenic and monogenic basis of blood traits and diseases. Cell. 2020;182:1214–1231.e11.
Sinnott-Armstrong N, Tanigawa Y, Amar D, Mars N, Benner C, Aguirre M, et al. Genetics of 35 blood and urine biomarkers in the UK Biobank. Nat Genet. 2021;53:185–94.
Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife. 2018;7:e34408 https://elifesciences.org/articles/34408/figures.
Davey Smith G, Hemani G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum Mol Genet. 2014;23:R89–98.
Smith GD, Ebrahim S. “Mendelian randomization”: can genetic epidemiology contribute to understanding environmental determinants of disease? Int J Epidemiol. 2003;32:1–22.
Glymour MM, Tchetgen Tchetgen EJ, Robins JM. Credible mendelian randomization studies: approaches for evaluating the instrumental variable assumptions. Am J Epidemiol. 2012;175:332–9.
Swanson SA, Hernán MA. Commentary. Epidemiology. 2013;24:370–4.
Labrecque J, Swanson SA. Understanding the assumptions underlying instrumental variable analyses: a brief review of falsification strategies and related tools. Curr Epidemiol Rep. 2018;5:214–20.
Sheehan NA, Didelez V. Epidemiology, genetic epidemiology and mendelian randomisation: more need than ever to attend to detail. Hum Genet. 2020;139:121–36.
Davies NM, Smith GD, Windmeijer F, Martin RM. Issues in the reporting and conduct of instrumental variable studies: a systematic review. Epidemiology. 2013;24:363–9.
Islam SN, Ahammed T, Anjum A, Albalawi O, Uddin MJ. Reporting methodological issues of the mendelian randomization studies in health and medical research: a systematic review. BMC Med Res Methodol. 2022;22:21.
Muhie S, Gautam A, Yang R, Misganaw B, Daigle BJ Jr, Mellon SH, et al. Molecular signatures of post-traumatic stress disorder in war-zone-exposed veteran and active-duty soldiers. Cell Rep Med. 2023;4:101045.
Michopoulos V, Beurel E, Gould F, Dhabhar FS, Schultebraucks K, Galatzer-Levy I, et al. Association of prospective risk for chronic PTSD symptoms with low TNFα and IFNγ concentrations in the immediate aftermath of trauma exposure. Am J Psychiatry. 2020;177:58–65.
Schultebraucks K, Shalev AY, Michopoulos V, Grudzen CR, Shin S-M, Stevens JS, et al. A validated predictive algorithm of post-traumatic stress course following emergency department admission after a traumatic stressor. Nat Med. 2020;26:1084–8.
Lawn RB, Murchland AR, Kim Y, Chibnik LB, Tworoger SS, Rimm EB, et al. Trauma, psychological distress and markers of systemic inflammation among US women: a longitudinal study. Psychoneuroendocrinology. 2022;145:105915.
Weiss T, Skelton K, Phifer J, Jovanovic T, Gillespie CF, Smith A, et al. Posttraumatic stress disorder is a risk factor for metabolic syndrome in an impoverished urban population. Gen Hosp Psychiatry. 2011;33:135–42.
Roberts AL, Agnew-Blais JC, Spiegelman D, Kubzansky LD, Mason SM, Galea S, et al. Posttraumatic stress disorder and incidence of type 2 diabetes mellitus in a sample of women: a 22-year longitudinal study: A 22-year longitudinal study. JAMA Psychiatry. 2015;72:203–10.
Blessing EM, Reus V, Mellon SH, Wolkowitz OM, Flory JD, Bierer L, et al. Biological predictors of insulin resistance associated with posttraumatic stress disorder in young military veterans. Psychoneuroendocrinology. 2017;82:91–7.
Mellon SH, Bersani FS, Lindqvist D, Hammamieh R, Donohue D, Dean K, et al. Metabolomic analysis of male combat veterans with post traumatic stress disorder. PLoS One. 2019;14:e0213839.
Sepúlveda-Lizcano L, Arenas-Villamizar VV, Jaimes-Duarte EB, García-Pacheco H, Paredes CS, Bermúdez V, et al. Metabolic adverse effects of psychotropic drug therapy: a systematic review. Eur J Investig Health Psychol Educ. 2023;13:1505–20.
O’Donnell CJ, Schwartz Longacre L, Cohen BE, Fayad ZA, Gillespie CF, Liberzon I, et al. Posttraumatic stress disorder and cardiovascular disease: state of the science, knowledge gaps, and research opportunities. JAMA Cardiol. 2021;6:1207–16.
Shaffer LR, Kaplan DE, Taddei TH, Mahmud N. The association between mental illness and all-cause mortality in patients with cirrhosis: a veterans affairs retrospective cohort study. Hepatol Commun. 2023;7:e0129.
Pathak GA, Singh K, Wendt FR, Fleming TW, Overstreet C, Koller D, et al. Genetically regulated multi-omics study for symptom clusters of posttraumatic stress disorder highlights pleiotropy with hematologic and cardio-metabolic traits. Mol Psychiatry. 2022;27:1394–404.
Dean KR, Hammamieh R, Mellon SH, Abu-Amara D, Flory JD, Guffanti G, et al. Multi-omic biomarker identification and validation for diagnosing warzone-related post-traumatic stress disorder. Mol Psychiatry. 2020;25:3337–49.
Jamilian H, Amirani E, Milajerdi A, Kolahdooz F, Mirzaei H, Zaroudi M, et al. The effects of vitamin D supplementation on mental health, and biomarkers of inflammation and oxidative stress in patients with psychiatric disorders: A systematic review and meta-analysis of randomized controlled trials. Prog Neuropsychopharmacol Biol Psychiatry. 2019;94:109651.
Terock J, Hannemann A, Van der Auwera S, Janowitz D, Spitzer C, Bonk S, et al. Posttraumatic stress disorder is associated with reduced vitamin D levels and functional polymorphisms of the vitamin D binding-protein in a population-based sample. Prog Neuropsychopharmacol Biol Psychiatry. 2020;96:109760.
Konzok J, Baumeister S-E, Winkler TW, Leitzmann MF, Baurecht H. Effect of 25-hydroxyvitamin D levels on the internalising dimension as a transdiagnostic risk factor: mendelian randomisation study. Br J Psychiatry. 2023;222:257–63.
Yang S, Wynn GH, Ursano RJ. A clinician’s guide to PTSD biomarkers and their potential future use. Focus. 2018;16:143–52.
Michopoulos V, Norrholm SD, Jovanovic T. Diagnostic biomarkers for posttraumatic stress disorder: promising horizons from translational neuroscience research. Biol Psychiatry. 2015;78:344–53.
Somvanshi PR, Mellon SH, Flory JD, Abu-Amara D, PTSD Systems Biology Consortium, Wolkowitz OM, et al. Mechanistic inferences on metabolic dysfunction in posttraumatic stress disorder from an integrated model and multiomic analysis: role of glucocorticoid receptor sensitivity. Am J Physiol Endocrinol Metab. 2019;317:E879–98.
Daskalakis NP, Iatrou A, Chatzinakos C, Jajoo A, Snijders C, Wylie D, et al. Systems biology dissection of PTSD and MDD across brain regions, cell types, and blood. Science. 2024;384:eadh3707.
Siegel CE, Laska EM, Lin Z, Xu M, Abu-Amara D, Jeffers MK, et al. Utilization of machine learning for identifying symptom severity military-related PTSD subtypes and their biological correlates. Transl Psychiatry. 2021;11:227.
Chen ZS, Kulkarni PP, Galatzer-Levy IR, Bigio B, Nasca C, Zhang Y. Modern views of machine learning for precision psychiatry. Patterns (N Y). 2022;3:100602.
Stein MB, Chen C-Y, Ursano RJ, Cai T, Gelernter J, Heeringa SG, et al. Genome-wide association studies of posttraumatic stress disorder in 2 cohorts of US Army soldiers. JAMA Psychiatry. 2016;73:695–704.
Nievergelt CM, Maihofer AX, Klengel T, Atkinson EG, Chen C-Y, Choi KW, et al. International meta-analysis of PTSD genome-wide association studies identifies sex- and ancestry-specific genetic risk loci. Nat Commun. 2019;10:4558.
Lawlor DA, Harbord RM, Sterne JAC, Timpson N, Davey Smith G. Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat Med. 2008;27:1133–63.
Lawlor DA. Commentary: two-sample mendelian randomization: opportunities and challenges. Int J Epidemiol. 2016;45:908–15.
DeViva JC, Sheerin CM, Southwick SM, Roy AM, Pietrzak RH, Harpaz-Rotem I. Correlates of VA mental health treatment utilization among OEF/OIF/OND veterans: resilience, stigma, social support, personality, and beliefs about treatment. Psychol Trauma. 2016;8:310–8.
Weathers FW DSM-5 Diagnostic Criteria for PTSD. Nemeroff CB, Marmar CR, editors. Oxford University Press; 2018.
Lee YH, Thaweethai T, Sheu Y-H, Feng Y-CA, Karlson EW, Ge T, et al. Impact of selection bias on polygenic risk score estimates in healthcare settings. Psychol Med. 2023;53:7435–45.
Burgess S, Davies NM, Thompson SG. Bias due to participant overlap in two-sample Mendelian randomization. Genet Epidemiol. 2016;40:597–608.
Seaman SR, White IR. Review of inverse probability weighting for dealing with missing data. Stat Methods Med Res. 2013;22:278–95.
Kang HJ, Yoon S, Lyoo IK. Peripheral biomarker candidates of posttraumatic stress disorder. Exp Neurobiol. 2015;24:186–96.
Daskalakis NP, Yehuda R, Diamond DM. Animal models in translational studies of PTSD. Psychoneuroendocrinology. 2013;38:1895–911.
Acknowledgements
This study would not be possible without the contributions of Mass General Brigham (MGB) patients and Biobank participants. We would also like to thank the research coordinators and the MGB Biobank study for their tremendous effort in participant recruitment and sample collection. Lastly, we would like to acknowledge the MGB Research Patient Data Registry (RPDR) team for their work maintaining the enterprise research patient data warehouse.
Funding
Financial support for the Psychiatric Genomics Consortium (PGC) PTSD Working Group was provided by the Cohen Veterans Bioscience, Stanley Center for Psychiatric Research at the Broad Institute, One Mind, and the National Institute of Mental Health (NIMH; R01MH106595, R01MH124847, R01MH124851, R01MH118233). JHL is individually supported by National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) K23 DK131345. YHL and NS are supported by the Broad Trauma Initiative. ALED is supported by NIMH T32 MH 017119.
Author information
Authors and Affiliations
Contributions
Conceptualization and design of the project: YHL, NS, ALR, KCK, YZ, ALED; Phenotypic data extraction and preprocessing: YHL, YZ; Genomic data extraction and preprocessing: TG, YAF; Genome-wide analyses: CMN, AXM; Polygenic risk scoring: YZ; Laboratory test-wide scan analyses: YHL, YZ; Interpretation of results, writing, and editing of the manuscript: YHL, NS, ALR, YZ, ALED, JHL, JDT, TG, KCK, JWS; Critical revision and supervision: YHL, NS, ALR, JWS, KCK.
Corresponding authors
Ethics declarations
Ethics declarations
JWS is a member of the Scientific Advisory Board of Sensorium Therapeutics (with options), has received a consulting fee from Data Driven, Inc., and has received grant support from Biogen, Inc. KCK receives consulting fees from the US Department of Justice and Covington Burling LLP and has royalties from Guilford Press and Oxford University Press. All other authors report no biomedical financial interests or potential conflicts of interest.
Ethics approval and consent to participate
MGB Biobank participants are enrolled through a broad-based consent process by research coordinators at health system practices, public hospital locations, dual consent as part of collaborating studies, or electronically through Patient Gateway, the MGB patient portal. All participants provided informed consent prior to enrollment, agreeing to contribute a blood sample linked to their electronic health record. The seven-page consent form (available upon request) covers key provisions, including the use of samples for any type of research, recontact by Biobank staff as needed, and the return of medically actionable findings.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, Y.H., Zhang, Y., Espinosa Dice, A.L. et al. Towards scalable biomarker discovery in posttraumatic stress disorder: triangulating genomic and phenotypic evidence from a health system biobank. Mol Psychiatry (2026). https://doi.org/10.1038/s41380-026-03553-z
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41380-026-03553-z