Abstract
Genetics can inform biologically relevant drug development and repurposing, which may improve patient care. Here, we leverage the genetics of psychiatric disorders to prioritize potential drug targets and compounds. We used the genome-wide association studies of four psychiatric disorders [attention deficit hyperactivity disorder (ADHD), bipolar disorder, depression, and schizophrenia] and genes encoding drug targets. We conducted drug enrichment analyses incorporating the novel and biologically specific GSA-MiXeR tool. We conducted multiple molecular trait analyses using large-scale transcriptomic and proteomic datasets sampled from brain and blood tissue. This included the novel use of the UK Biobank proteomic data for a proteome-wide association study of psychiatric disorders. With the accumulated evidence, we prioritize potential drug targets and compounds for each disorder. We reveal candidate drug targets associated with a single or multiple disorders that implicate glutamate signaling. Drug prioritization indicated genetic support for psychotropic medications, including several top-ranked antipsychotics for schizophrenia. We also observed genetic support for commonly used psychotropics for psychiatric treatment (e.g., clozapine, duloxetine, and lithium). Revealed opportunities for drug repurposing included cholinergic drugs for ADHD, estrogen modulators for depression, and matrix metalloproteinases for ADHD and depression. Our findings indicate the genetic liability to schizophrenia is associated with reduced brain and blood expression of CYP2D6, a gene encoding a metabolizer of drugs and neurotransmitters, suggesting a genetic risk for poor drug response and altered neurotransmission. Our extensive analyses highlight the utility of genetics for informing drug development and repurposing for psychiatric disorders, providing novel opportunities for improving patient outcomes.

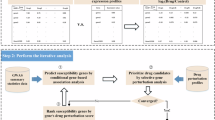
Depicted is the series of analyses conducted to generate a list of prioritized drug targets and compounds. First pairings of genome-wide association study (GWAS) traits with drugs are generated using enrichment analyses. Next, a series of molecular trait analyses is conducted to generate and rank a list of potential drug targets for each GWAS trait. Finally, enrichment and molecular trait results are combined to generate a ranked list of prioritized drugs for each GWAS trait based on supporting genetic evidence. ADHD = Attention deficit hyperactivity disorder, BIP = Bipolar disorder, DEP = Depression, SCZ = Schizophrenia, DBP = Diastolic blood pressure, T2D = Type 2 diabetes, RNA = ribonucleic acid, XWAS = both transcriptome and proteome-wide association studies, MR = Mendelian randomization, coloc = colocalization.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 13 print issues and online access
$259.00 per year
only $19.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All GWAS summary statistics and molecular trait data included in this study are publicly available. We generated PWAS weights using proteomic data from the UK Biobank and have shared those weights on figshare [https://doi.org/10.6084/m9.figshare.31487881]. The GWAS summary statistics from the Million Veteran Program were downloaded from the dbGaP website, under accession phs001672.
References
Minikel EV, Painter JL, Dong CC, Nelson MR. Refining the impact of genetic evidence on clinical success. Nature. 2024;629:624–9.
Nelson MR, Tipney H, Painter JL, Shen J, Nicoletti P, Shen Y, et al. The support of human genetic evidence for approved drug indications. Nat Genet. 2015;47:856–60.
Linden M, Schermuly-Haupt M-L. Definition, assessment and rate of psychotherapy side effects. World Psychiatry. 2014;13:306–09.
Correll CU, Detraux J, De Lepeleire J, De Hert M. Effects of antipsychotics, antidepressants and mood stabilizers on risk for physical diseases in people with schizophrenia, depression and bipolar disorder. World Psychiatry. 2015;14:119–36.
Solovieff N, Cotsapas C, Lee PH, Purcell SM, Smoller JW. Pleiotropy in complex traits: challenges and strategies. Nat Rev Genet. 2013;14:483–95.
Bell N, Uffelmann E, Walree E Van, Leeuw C de, Posthuma D. Using genome-wide association results to identify drug repurposing candidates. 2022.
Mullins N, Forstner AJ, O’Connell KS, Coombes B, Coleman JRI, Qiao Z, et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat Genet. 2021;53:817–29.
O’Connell KS, Koromina M, Veen T van der, Boltz T, David FS, Yang JM, et al. Genomics yields biological and phenotypic insights into bipolar disorder. Nature. 2024:639:968–75.
Frei O, Hindley G, Shadrin AA, van der Meer D, Akdeniz BC, Hagen E, et al. Improved functional mapping of complex trait heritability with GSA-MiXeR implicates biologically specific gene sets. Nat Genet. 2024;56:1310–18.
Reay WR, Cairns MJ. Advancing the use of genome-wide association studies for drug repurposing. Nat Rev Genet. 2021;22:658–71.
Grotzinger AD, Fuente J de la, Davies G, Nivard MG, Tucker-Drob EM. Transcriptome-wide and stratified genomic structural equation modeling identify neurobiological pathways shared across diverse cognitive traits. Nat Commun. 2021;13:6280.
Wingo TS, Liu Y, Gerasimov ES, Gockley J, Logsdon BA, Duong DM, et al. Brain proteome-wide association study implicates novel proteins in depression pathogenesis. Nat Neurosci. 2021;24:810–17.
Li X, Shen A, Zhao Y, Xia J. Mendelian randomization using the druggable genome reveals genetically supported drug targets for psychiatric disorders. Schizophr Bull. 2023;49:1305–15.
Liu J, Cheng Y, Li M, Zhang Z, Li T, Luo X-J. Genome-wide Mendelian randomization identifies actionable novel drug targets for psychiatric disorders. Neuropsychopharmacol. 2023;48:270–80.
Deng Y-T, Ou Y-N, Wu B-S, Yang Y-X, Jiang Y, Huang Y-Y, et al. Identifying causal genes for depression via integration of the proteome and transcriptome from the brain and blood. Mol Psychiatry. 2022;27:2849–57.
Lu T, Forgetta V, Greenwood CMT, Zhou S, Richards JB. Circulating proteins influencing psychiatric disease: a mendelian randomization study. Biol Psychiatry. 2023;93:82–91.
Namba S, Konuma T, Wu K-H, Zhou W, Okada Y. A practical guideline of genomics-driven drug discovery in the era of global biobank meta-analysis. Cell Genom. 2022;2:100190.
Sadler MC, Auwerx C, Deelen P, Kutalik Z. Multi-layered genetic approaches to identify approved drug targets. Cell Genom. 2023;3:100341.
Reay WR, Geaghan MP, Atkins JR, Carr VJ, Green MJ, Cairns MJ. Genetics-informed precision treatment formulation in schizophrenia and bipolar disorder. Am J Hum Genet. 2022;109:1620–37.
Koscielny G, An P, Carvalho-Silva D, Cham JA, Fumis L, Gasparyan R, et al. Open targets: a platform for therapeutic target identification and validation. Nucleic Acids Res. 2017;45:D985–94.
Koch E, Kauppi K, Chen C-H. Candidates for drug repurposing to address the cognitive symptoms in schizophrenia. Prog Neuro Psychopharmacol Biol Psychiatry. 2023;120:110637.
Demontis D, Walters GB, Athanasiadis G, Walters R, Therrien K, Nielsen TT, et al. Genome-wide analyses of ADHD identify 27 risk loci, refine the genetic architecture and implicate several cognitive domains. Nat Genet. 2023;55:198–208.
Als TD, Kurki MI, Grove J, Voloudakis G, Therrien K, Tasanko E, et al. Depression pathophysiology, risk prediction of recurrence and comorbid psychiatric disorders using genome-wide analyses. Nat Med. 2023;29:1832–44.
Trubetskoy V, Pardiñas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:502–8.
Evangelou E, Warren HR, Mosen-Ansorena D, Mifsud B, Pazoki R, Gao H, et al. Genetic analysis of over 1 million people identifies 535 new loci associated with blood pressure traits. Nat Genet. 2018;50:1412–25.
Mahajan A, Spracklen CN, Zhang W, Ng MCY, Petty LE, Kitajima H, et al. Multi-ancestry genetic study of type 2 diabetes highlights the power of diverse populations for discovery and translation. Nat Genet. 2022;54:560–72.
Giri A, Hellwege JN, Keaton JM, Park J, Qiu C, Warren HR, et al. Trans-ethnic association study of blood pressure determinants in over 750,000 individuals. Nat Genet. 2019;51:51–62.
Vujkovic M, Keaton JM, Lynch JA, Miller DR, Zhou J, Tcheandjieu C, et al. Discovery of 318 new risk loci for type 2 diabetes and related vascular outcomes among 1.4 million participants in a multi-ancestry meta-analysis. Nat Genet. 2020;52:680–91.
Cannon M, Stevenson J, Stahl K, Basu R, Coffman A, Kiwala S, et al. DGIdb 5.0: rebuilding the drug–gene interaction database for precision medicine and drug discovery platforms. Nucleic Acids Res. 2024;52:D1227–35.
Cheng W, Parker N, Karadag N, Koch E, Hindley G, Icick R, et al. The relationship between cannabis use, schizophrenia, and bipolar disorder: a genetically informed study. Lancet Psychiatry. 2023;10:441–51.
Bipolar Disorder Working Group of the Psychiatric Genomics Consortium 23andMe Research TeamO’Connell KS, Koromina M, Forstner AJ, McQuillin A, Florio AD, et al. Genetic diversity enhances gene discovery for bipolar disorder. 2023.
De Leeuw CA, Mooij JM, Heskes T, Posthuma D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput Biol. 2015;11: e1004219.
Gusev A, Ko A, Shi H, Bhatia G, Chung W, Penninx BWJH, et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet. 2016;48:245–52.
Gandal MJ, Zhang P, Hadjimichael E, Walker RL, Chen C, Liu S, et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science. 2018;362:eaat8127.
Ardlie KG, DeLuca DS, Segrè AV, Sullivan TJ, Young TR, Gelfand ET, et al. The genotype-tissue expression (GTEx) pilot analysis: Multitissue gene regulation in humans. Science. 2015;348:648–60.
Wingo AP, Liu Y, Gerasimov ES, Vattathil SM, Liu J, Cutler DJ, et al. Sex differences in brain protein expression and disease. Nat Med. 2023;29:2224–32.
Zhang J, Dutta D, Köttgen A, Tin A, Schlosser P, Grams ME, et al. Plasma proteome analyses in individuals of European and African ancestry identify cis-pQTLs and models for proteome-wide association studies. Nat Genet. 2022;54:593–602.
Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, et al. The MR-Base platform supports systematic causal inference across the human phenome. eLife. 2018;7:e34408.
de Klein N, Tsai EA, Vochteloo M, Baird D, Huang Y, Chen C-Y, et al. Brain expression quantitative trait locus and network analyses reveal downstream effects and putative drivers for brain-related diseases. Nat Genet. 2023;55:377–88.
Wingo AP, Liu Y, Gerasimov ES, Gockley J, Logsdon BA, Duong DM, et al. Integrating human brain proteomes with genome-wide association data implicates new proteins in Alzheimer’s disease pathogenesis. Nat Genet. 2021;53:143–6.
Võsa U, Claringbould A, Westra H-J, Bonder MJ, Deelen P, Zeng B, et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat Genet. 2021;53:1300–10.
Sun BB, Chiou J, Traylor M, Benner C, Hsu Y-H, Richardson TG, et al. Plasma proteomic associations with genetics and health in the UK Biobank. Nature. 2023;622:329–338.
Giambartolomei C, Vukcevic D, Schadt EE, Franke L, Hingorani AD, Wallace C, et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLOS Genet. 2014;10:e1004383.
Subramanian A, Narayan R, Corsello SM, Peck DD, Natoli TE, Lu X, et al. A next-generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell. 2017;171:1437–52.e17.
Enache OM, Lahr DL, Natoli TE, Litichevskiy L, Wadden D, Flynn C, et al. The GCTx format and cmap{Py, R, M, J} packages: resources for optimized storage and integrated traversal of annotated dense matrices. Bioinformatics. 2019;35:1427–9.
Lee PH, Anttila V, Won H, Feng Y-CA, Rosenthal J, Zhu Z, et al. Genomic relationships, novel loci, and pleiotropic mechanisms across eight psychiatric disorders. Cell. 2019;179:1469–82.e11.
Smeland OB, Wang Y, Frei O, Li W, Hibar DP, Franke B, et al. Genetic overlap between schizophrenia and volumes of hippocampus, putamen, and intracranial volume indicates shared molecular genetic mechanisms. Schizophr Bull. 2018;44:854–64.
Vosberg DE, Leyton M, Flores C. The Netrin-1/DCC guidance system: dopamine pathway maturation and psychiatric disorders emerging in adolescence. Mol Psychiatry. 2020;25:297–307.
Parker N, Cheng W, Hindley GFL, Parekh P, Shadrin AA, Maximov II, et al. Psychiatric disorders and brain white matter exhibit genetic overlap implicating developmental and neural cell biology. Mol Psychiatry. 2023;28:4924–32.
Repple J, Gruber M, Mauritz M, de Lange SC, Winter NR, Opel N, et al. Shared and specific patterns of structural brain connectivity across affective and psychotic disorders. Biol Psychiatry. 2023;93:178–86.
Javitt DC. Glutamate as a therapeutic target in psychiatric disorders. Mol Psychiatry. 2004;9:984–97.
Leucht S, Siafis S, McGrath JJ, McGorry P, Howes OD, Tamminga C, et al. Schizophrenia. Nat Rev Dis Primers. 2025;11:83.
Howard DM, Adams MJ, Shirali M, Clarke T-K, Marioni RE, Davies G, et al. Genome-wide association study of depression phenotypes in UK Biobank identifies variants in excitatory synaptic pathways. Nat Commun. 2018;9:1470.
Wingo AP, Liu Y, Vattathil SM, Gerasimov ES, Mei Z, Ravindran SP, et al. Multiancestry brain pQTL fine-mapping and integration with genome-wide association studies of 21 neurologic and psychiatric conditions. Nat Genet. 2025;57:2156–65.
Quintero J, Gutiérrez-Casares JR, Álamo C. Molecular characterisation of the mechanism of action of stimulant drugs lisdexamfetamine and methylphenidate on ADHD neurobiology: a review. Neurol Ther. 2022;11:1489–1517.
Potter AS, Schaubhut G, Shipman M. Targeting the nicotinic cholinergic system to treat attention-deficit/hyperactivity disorder: rationale and progress to date. CNS Drugs. 2014;28:1103–13.
Kaczmarek KT, Protokowicz K, Kaczmarek L. Matrix metalloproteinase-9: a magic drug target in neuropsychiatry? J Neurochem. 2024;168:1842–53.
Adams MJ, Streit F, Meng X, Awasthi S, Adey BN, Choi KW, et al. Trans-ancestry genome-wide study of depression identifies 697 associations implicating cell types and pharmacotherapies. Cell. 2025;188:640–52.e9.
de Novaes Soares C, Almeida OP, Joffe H, Cohen LS. Efficacy of estradiol for the treatment of depressive disorders in perimenopausal women: a double-blind, randomized, placebo-controlled trial. Arch Gen Psychiatry. 2001;58:529–34.
Lorsch ZS, Loh Y-HE, Purushothaman I, Walker DM, Parise EM, Salery M, et al. Estrogen receptor α drives pro-resilient transcription in mouse models of depression. Nat Commun. 2018;9:1116.
Stanikova D, Luck T, Bae YJ, Thiery J, Ceglarek U, Engel C, et al. Increased estrogen levels can be associated with depression in males. Psychoneuroendocrinology. 2018;87:196–203.
Carrier N, Saland SK, Duclot F, He H, Mercer R, Kabbaj M. The anxiolytic and antidepressant-like effects of testosterone and estrogen in gonadectomized male rats. Biol Psychiatry. 2015;78:259–69.
Albert KM, Newhouse PA. Estrogen, stress, and depression: cognitive and biological interactions. Annu Rev Clin Psychol. 2019;15:399–423.
Bertilsson L, Dahl M-L, Dalén P, Al-Shurbaji A. Molecular genetics of CYP2D6: Clinical relevance with focus on psychotropic drugs. Br J Clin Pharmacol. 2002;53:111–22.
Ma L, Shcherbina A, Chetty S. Variations and expression features of CYP2D6 contribute to schizophrenia risk. Mol Psychiatry. 2021;26:2605–15.
Wang X, Li J, Dong G, Yue J. The endogenous substrates of brain CYP2D. Eur J Pharmacol. 2014;724:211–8.
Milosavljević, Bukvić F, Pavlović N, Miljević Z, Pešić V, Molden E, et al. Association of CYP2C19 and CYP2D6 poor and intermediate metabolizer status with antidepressant and antipsychotic exposure. JAMA Psychiatry. 2021;78:1–11.
Neville MJ, Johnstone EC, Walton RT. Identification and characterization of ANKK1: a novel kinase gene closely linked to DRD2 on chromosome band 11q23.1. Hum Mutat. 2004;23:540–5.
Suzuki A, Mihara K, Kondo T, Tanaka O, Nagashima U, Otani K, et al. The relationship between dopamine D2 receptor polymorphism at the Taq1 A locus and therapeutic response to nemonapride, a selective dopamine antagonist, in schizophrenic patients. Pharmacogenetics Genom. 2000;10:335.
Meltzer HY. The role of serotonin in antipsychotic drug action. Neuropsychopharmacol. 1999;21:106–15.
Cipriani A, Saunders K, Attenburrow M-J, Stefaniak J, Panchal P, Stockton S, et al. A systematic review of calcium channel antagonists in bipolar disorder and some considerations for their future development. Mol Psychiatry. 2016;21:1324–32.
Manzoni C, Kia DA, Vandrovcova J, Hardy J, Wood NW, Lewis PA, et al. Genome, transcriptome and proteome: the rise of omics data and their integration in biomedical sciences. Brief Bioinforma. 2018;19:286–302.
Qi T, Wu Y, Zeng J, Zhang F, Xue A, Jiang L, et al. Identifying gene targets for brain-related traits using transcriptomic and methylomic data from blood. Nat Commun. 2018;9:2282.
Acknowledgements
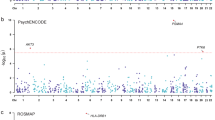
We thank all participants, staff, and providers of the publicly available data used in this study, including individuals from the UKB, psychiatric genomics consortium, Million Veterans Program, PsychENCODE, ARIC, MetaBrain, ROS/MAP, eQTLGen, and GTEx.
Funding
Funding was provided by the US National Institutes of Health (grants U24DA041123, R01AG076838, U24DA055330, OT2HL161847 and 5R01MH124839-02), the Research Council of Norway (grants 296030, 273291, 273446, 300309, 324252, 326813, 334920), the South-East Regional Health Authority (grant 2022–073), EEA-RO-NO-2018–0573, EEA-RO-NO-2018–0535, EU’s Horizon 2020 Research and Innovation Programme (grant 847776; CoMorMent and 964874 REALMENT), Novo Nordisk Foundation (grant NNF23OC0099658), the Marie Skłodowska-Curie Actions Grant 801133 (Scientia fellowship). This work was performed on Services for sensitive data (TSD), University of Oslo, Norway, with resources provided by UNINETT Sigma2—the National Infrastructure for High Performance Computing and Data Storage in Norway (NS9666S).
Author information
Authors and Affiliations
Contributions
NP and OBS conceived of the study, and EK, AAS, GFLH, SS, PJ, KSO, and OAA were involved in study design. NP, EK, AAS, TSW, and APW were involved in data acquisition and processing. NP, EK, AAS, and JF were involved in data analysis. NP had access to all data and drafted the initial manuscript. All authors contributed to data interpretation and editing of the manuscript and accepted responsibility to submit the manuscript for publication.
Corresponding authors
Ethics declarations
Competing interests
OAA has received speaker fees from Lundbeck, Janssen, Otsuka, and Sunovion and is a consultant to Cortechs.ai. and Precision Health. AMD is a Founding Director and holds equity in CorTechs Labs, Inc. (DBA Cortechs.ai), Precision Pro, Inc., and Precision Health and Wellness, Inc. Dr. Dale is the President and a Board of Trustees member of the J. Craig Venter Institute (JCVI) and holds an appointment as Professor II at University of Oslo in Norway. OF is a consultant to Precision Health. TSW is a co-founder of revXon, which is disclosed and approved by the University of California, Davis, per its conflict-of-interest policy. All other authors report no biomedical financial interests or potential conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Parker, N., Koch, E., Shadrin, A.A. et al. Leveraging the genetics of psychiatric disorders to prioritize potential drug targets and compounds. Neuropsychopharmacol. (2026). https://doi.org/10.1038/s41386-026-02380-8
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41386-026-02380-8