Abstract
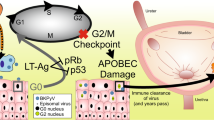
Chronic BK polyomavirus (BKPyV) infection is recognized as a potential oncogenic factor of urothelial carcinoma (UC) in renal transplant recipients. Recent studies have reported a positive correlation among BKPyV integration, persistent overexpression of viral large T antigen (TAg), and malignancy, yet little is known about the specific integration mechanisms and the impacts of viral integration. Here, we performed whole-genome sequencing (WGS) and viral capture-based sequencing on high-grade immunohistochemically TAg-positive UCs in two renal transplant recipients. A total of 181 integration sites, including the three found by WGS, were identified by viral capture-based sequencing, indicating its enhanced sensitivity and ability in identifying low-read integration sites in subpopulations of the tumor cells. The microhomologies between human and BKPyV genomes were significantly enriched in the flanking regions of 84.5% the integration sites, with a median length of 7 bp. Notably, 75 human genes formed fusion sequences due to viral insertional integration. Among them, the expression of 15 genes were statistically associated with UC based on GEO2R expression analysis. Our results indicated a multisite and multifragment linear integration pattern and a potential microhomology or nonhomologous end joining integration mechanism at the single-nucleotide level. We put forward a potential selection mechanism driven by immunity and centered on viral integration in the carcinogenesis of BKPyV.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Fu F, Deng W, Yu S, Liu Y, Yu L, Liu R, et al. Occurrence and regression of BK polyomavirus associated carcinoma: a clinical and next-generation sequencing study. Clin Sci. 2018;132:1753–63.
Rosales BM, De La Mata N, Vajdic CM, Kelly PJ, Wyburn K, Webster AC. Cancer mortality in kidney transplant recipients: an Australian and New Zealand population-based cohort study, 1980–2013. Int J Cancer. 2020;146:2703–11.
O’Neill JP, Sexton DJ, O’Leary E, O’Kelly P, Murray S, Deady S, et al. Post-transplant malignancy in solid organ transplant recipients in Ireland The Irish Transplant Cancer Group. Clin Transplant. 2019;33:e13669.
Awan AA, Niu J, Pan JS, Erickson KF, Mandayam S, Winkelmayer WC, et al. Trends in the CAuses of Death among Kidney Transplant Recipients in the United States (1996–2014). AM J Nephrol. 2018;48:472–81.
Papadimitriou JC, Randhawa P, Rinaldo CH, Drachenberg CB, Alexiev B, Hirsch HH. BK polyomavirus infection and renourinary tumorigenesis. AM J Transplant. 2016;16:398–406.
Starrett GJ, Buck CB. The case for BK polyomavirus as a cause of bladder cancer. Curr Opin Virol. 2019;39:8–15.
Saribas AS, Coric P, Hamazaspyan A, Davis W, Axman R, White MK, et al. Emerging from the unknown: structural and functional features of agnoprotein of polyomaviruses. J Cell Physiol. 2016;231:2115–27.
Houben R, Angermeyer S, Haferkamp S, Aue A, Goebeler M, Schrama D, et al. Characterization of functional domains in the Merkel cell polyoma virus Large T antigen. Int J Cancer. 2015;136:E290–300.
Hesbacher S, Pfitzer L, Wiedorfer K, Angermeyer S, Borst A, Haferkamp S, et al. RB1 is the crucial target of the Merkel cell polyomavirus Large T antigen in Merkel cell carcinoma cells. Oncotarget. 2016;7:32956–68.
Elfadawy N, Yamada M, Sarabu N. Management of BK polyomavirus infection in kidney and kidney-pancreas transplant recipients: a review article. Infect Dis Clin North AM. 2018;32:599–613.
Kenan DJ, Mieczkowski PA, Latulippe E, Cote I, Singh HK, Nickeleit V. BK polyomavirus genomic integration and large T antigen expression: evolving paradigms in human oncogenesis. AM J Transplant. 2017;17:1674–80.
Kenan DJ, Mieczkowski PA, Burger-Calderon R, Singh HK, Nickeleit V. The oncogenic potential of BK-polyomavirus is linked to viral integration into the human genome. J Pathol. 2015;237:379–89.
Muller DC, Ramo M, Naegele K, Ribi S, Wetterauer C, Perrina V, et al. Donor-derived, metastatic urothelial cancer after kidney transplantation associated with a potentially oncogenic BK polyomavirus. J Pathol. 2018;244:265–70.
Wang YC, Liu YN, Deng WF, Fu FX, Yan SS, Yang HW, et al. Viral integration in BK polyomavirus-associated urothelial carcinoma in renal transplant recipients: multistage carcinogenesis revealed by next-generation virome capture sequencing. Oncogene. 2020;39:5734–42.
Egli A, Infanti L, Dumoulin A, Buser A, Samaridis J, Stebler C, et al. Prevalence of polyomavirus BK and JC infection and replication in 400 healthy blood donors. J Infect Dis. 2009;199:837–46.
Kean JM, Rao S, Wang M, Garcea RL. Seroepidemiology of human polyomaviruses. Plos Pathog. 2009;5:e1000363.
Pinto M, Dobson S. BK and JC virus: a review. J Infect. 2014;68:S2–8.
Trofe-Clark J, Sawinski D. BK and other polyomaviruses in kidney transplantation. Semin Nephrol. 2016;36:372–85.
Lee YJ, Zheng J, Kolitsopoulos Y, Chung D, Amigues I, Son T, et al. Relationship of BK polyoma virus (BKV) in the urine with hemorrhagic cystitis and renal function in recipients of T Cell-depleted peripheral blood and cord blood stem cell transplantations. Biol Blood Marrow Transplant. 2014;20:1204–10.
Nickeleit V, Singh HK, Kenan DJ, Mieczkowski PA. The two-faced nature of BK polyomavirus: lytic infection or non-lytic large-T-positive carcinoma. J Pathol. 2018;246:7–11.
Roberts IS, Besarani D, Mason P, Turner G, Friend PJ, Newton R. Polyoma virus infection and urothelial carcinoma of the bladder following renal transplantation. BRJ Cancer. 2008;99:1383–6.
Gupta G, Kuppachi S, Kalil RS, Buck CB, Lynch CF, Engels EA. Treatment for presumed BK polyomavirus nephropathy and risk of urinary tract cancers among kidney transplant recipients in the United States. AM J Transplant. 2018;18:245–52.
Starrett GJ, Buck CB. The case for BK polyomavirus as a cause of bladder cancer. Curr Opin Virol. 2019;39:8–15.
Li W, Zeng X, Lee NP, Liu X, Chen S, Guo B, et al. HIVID: an efficient method to detect HBV integration using low coverage sequencing. Genomics. 2013;102:338–44.
Groves IJ, Coleman N. Human papillomavirus genome integration in squamous carcinogenesis: what have next-generation sequencing studies taught us? J Pathol. 2018;245:9–18.
Morrical SW. DNA-pairing and annealing processes in homologous recombination and homology-directed repair. Cold Spring Harb Perspect Biol. 2015;7:a16444.
Ottaviani D, LeCain M, Sheer D. The role of microhomology in genomic structural variation. Trends Genet. 2014;30:85–94.
Hu Z, Zhu D, Wang W, Li W, Jia W, Zeng X, et al. Genome-wide profiling of HPV integration in cervical cancer identifies clustered genomic hot spots and a potential microhomology-mediated integration mechanism. Nat Genet. 2015;47:158–63.
Yi K, Ju YS. Patterns and mechanisms of structural variations in human cancer. Exp Mol Med. 2018;50:98.
Zhou X, Workeneh B, Hu Z, Li R. Effect of immunosuppression on the human mesangial cell cycle. Mol Med Rep. 2015;11:910–6.
Zhang F, Khajavi M, Connolly AM, Towne CF, Batish SD, Lupski JR. The DNA replication FoSTeS/MMBIR mechanism can generate genomic, genic and exonic complex rearrangements in humans. Nat Genet. 2009;41:849–53.
Chang H, Pannunzio NR, Adachi N, Lieber MR. Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat Rev Mol Cell Biol. 2017;18:495–506.
Lieber MR. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem. 2010;79:181–211.
Michel OR, Wolff DJ, Schandl CA, Drabkin HA. Urothelial carcinoma of donor origin in a kidney transplant patient. J Immunother Cancer. 2016;4:63.
Sirohi D, Vaske C, Sanborn Z, Smith SC, Don MD, Lindsey KG, et al. Polyoma virus-associated carcinomas of the urologic tract: a clinicopathologic and molecular study. Mod Pathol. 2018;31:1429–41.
Cangemi M, Montico B, Fae DA, Steffan A, Dolcetti R. Dissecting the multiplicity of immune effects of immunosuppressive drugs to better predict the risk of de novo malignancies in solid organ transplant patients. Front Oncol. 2019;9:160.
Billups K, Neal J, Salyer J. Immunosuppressant-driven de novo malignant neoplasms after solid-organ transplant. Prog Transplant. 2015;25:182–8.
McCabe MT, Low JA, Imperiale MJ, Day ML. Human polyomavirus BKV transcriptionally activates DNA methyltransferase 1 through the pRb/E2F pathway. Oncogene. 2006;25:2727–35.
Acknowledgements
This work was funded by the National Natural Science Foundation of China (81500573), the Science Foundation of Guangdong Province (2020A1515010674), the Science and Technology Planning Project of Guangzhou (201803010109), the President Funding of Nanfang Hospital (2018B009, 2018C003) and College Students’ Innovative Entrepreneurial Training Plan Program (201812121148, X202012121239, 202012121046).
Author information
Authors and Affiliations
Contributions
CLW and YM designed and directed the study. YJ, YZ, WFD, and YCW wrote the manuscript. RJL, YNL, NE, and JG performed sample preparation, sequencing and statistical analysis and validation. MHL, RML, and BWG performed experimental work and analyzed the data. YCH, JFM, JPZ, and NL took charge for data visualization. HQN, JX, MLB, and RBC revised the manuscript. All authors read and approved the final version.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jin, Y., Zhou, Y., Deng, W. et al. Genome-wide profiling of BK polyomavirus integration in bladder cancer of kidney transplant recipients reveals mechanisms of the integration at the nucleotide level. Oncogene 40, 46–54 (2021). https://doi.org/10.1038/s41388-020-01502-w
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-020-01502-w