Abstract
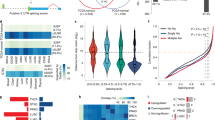
Cancer-related genes have evolved specific genetic and genomic features to favor tumor suppression. Previously we reported that tumor suppressor genes (TSGs) acquired high promoter CpG dinucleotide frequencies during evolution to maintain high expression in normal tissues and resist cancer-specific downregulation. In this study, we investigated whether 3′untranslated regions (3′UTRs) of TSGs have evolved specific features to carry out similar functions. We found that 3′UTRs of TSGs, especially those involved in multiple histological types and pediatric cancers, are longer than those of non-cancer genes. 3′UTRs of TSGs also exhibit higher density of binding sites for RNA-binding proteins (RBPs), particularly those having high affinities to C-rich motifs. Both longer 3′UTR length and RBP binding sites enrichment are correlated with higher gene expression in normal tissues across tissue types. Moreover, both features together with the correlated N6-methyladenosine modification and the extent of protein-protein interactions are positively associated with the ability of TSGs to resist cancer-specific downregulation. These results were successfully validated with independent datasets. Collectively, these findings indicate that TSGs have evolved longer 3′UTR with increased propensity to RBP binding, N6-methyladenosine modification and protein-protein interactions for optimizing their tumor-suppressing functions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Processed gene expression data were obtained from the firebrowse database (http://firebrowse.org/), GTEx Portal (https://gtexportal.org) and Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/).
Code availability
Computer codes used in this study are available at the GitHub page: https://github.com/Hd0909/Evolution-of-3UTR.
References
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA, Kinzler KW. Cancer genome landscapes. Science. 2013;340:1546–58.
Thomas MA. Evolutionary dynamics of oncogenes and tumor suppressor genes: higher intensities of purifying selection than other genes. Mol Biol Evol. 2003;20:964–8.
Wu WKK, Li X, Wang X, Dai RZW, Cheng ASL, Wang MHT, et al. Oncogenes without a neighboring tumor-suppressor gene are more prone to amplification. Mol Biol Evol. 2017;34:903–7.
Wang X, Li X, Zhang L, Wong SH, Wang MHT, Tse G, et al. Oncogenes expand during evolution to withstand somatic amplification. Ann Oncol. 2018;29:2254–60.
Huang D, Wang X, Liu Y, Huang Z, Hu X, Hu W. et al. Multi-omic analysis suggests tumor suppressor genes evolved specific promoter features to optimize cancer resistance. Brief Bioinform. 2021;22:bbab040.
Dolnik A, Engelmann JC, Scharfenberger-Schmeer M, Mauch J, Kelkenberg-Schade S, Haldemann B, et al. Commonly altered genomic regions in acute myeloid leukemia are enriched for somatic mutations involved in chromatin remodeling and splicing. Blood. 2012;120:e83–92.
Ren G-X, Guo X-P, Sun Y-C. Regulatory 3′ untranslated regions of bacterial mRNAs. Front Microbiol. 2017;8:1276.
Mayr C. Regulation by 3′-untranslated regions. Annu Rev Genet. 2017;51:171–94.
Xie X, Lu J, Kulbokas EJ, Golub TR, Mootha V, Lindblad-Toh K, et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature. 2005;434:338–45.
Siepel A, Bejerano G, Pedersen JS, Hinrichs AS, Hou M, Rosenbloom K, et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 2005;15:1034–50.
Matoulkova E, Michalova E, Vojtesek B, Hrstka R. The role of the 3′ untranslated region in post-transcriptional regulation of protein expression in mammalian cells. RNA Biol. 2012;9:563–76.
Runge S, Nielsen FC, Nielsen J, Lykke-Andersen J, Wewer UM, Christiansen J. H19 RNA binds four molecules of insulin-like growth factor II mRNA-binding protein. J Biol Chem. 2000;275:29562–9.
Wu L, Fan J, Belasco JG. MicroRNAs direct rapid deadenylation of mRNA. 2006;103:4034–9.
Gealy C, Denson M, Humphreys C, McSharry B, Wilkinson G, Caswell R. Posttranscriptional suppression of interleukin-6 production by human cytomegalovirus. J Virol. 2005;79:472–85.
Linker K, Pautz A, Fechir M, Hubrich T, Greeve J, Kleinert H. Involvement of KSRP in the post-transcriptional regulation of human iNOS expression-complex interplay of KSRP with TTP and HuR. Nucleic Acids Res. 2005;33:4813–27.
Zhong L, Liao D, Zhang M, Zeng C, Li X, Zhang R, et al. YTHDF2 suppresses cell proliferation and growth via destabilizing the EGFR mRNA in hepatocellular carcinoma. Cancer Lett. 2019;442:252–61.
Yin H, Wang G, Ma L, Yi SV, Zhang Z. What Signatures dominantly associate with gene age? Genome Biol Evol. 2016;8:3083–9.
Liberzon A, Subramanian A, Pinchback R, Thorvaldsdóttir H, Tamayo P, Mesirov JP. Molecular signatures database (MSigDB) 3.0. Bioinformatics. 2011;27:1739–40.
Yu H, Wang J, Sheng Q, Liu Q, Shyr Y. BeRBP: binding estimation for human RNA-binding proteins. Nucleic Acids Res. 2019;47:e26.
Bailey TL, Elkan C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol. 1994;2:28–36.
Grant CE, Bailey TL, Noble WS. FIMO: scanning for occurrences of a given motif. Bioinformatics. 2011;27:1017–8.
Gupta S, Stamatoyannopoulos JA, Bailey TL, Noble WS. Quantifying similarity between motifs. Genome Biol. 2007;8:R24.
Alkan SA, Martincic K, Milcarek C. The hnRNPs F and H2 bind to similar sequences to influence gene expression. Biochem J. 2006;393:361–71.
Yabe-Wada T, Philpott CC, Onai N. PCBP2 post-transcriptionally regulates sortilin expression by binding to a C-rich element in its 3′ UTR. FEBS Open Bio. 2020;10:407–13.
Stark A, Brennecke J, Bushati N, Russell RB, Cohen SM. Animal MicroRNAs confer robustness to gene expression and have a significant impact on 3' UTR evolution. Cell. 2005;123:1133–46.
Zhang C, Fu J, Zhou Y. A review in research progress concerning m6A methylation and immunoregulation. Front Immunol. 2019;10:922.
Huang H, Weng H, Deng X, Chen J. RNA modifications in cancer: functions, mechanisms, and therapeutic implications. Annu Rev Cancer Biol. 2020;4:221–40.
Liu S, Zhu A, He C, Chen M. REPIC: a database for exploring the N 6-methyladenosine methylome. Genome Biol. 2020;21:100.
Xiao S, Cao S, Huang Q, Xia L, Deng M, Yang M, et al. The RNA N6-methyladenosine modification landscape of human fetal tissues. Nat Cell Biol. 2019;21:651–61.
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell. 2012;149:1635–46.
Bailey MH, Tokheim C, Porta-Pardo E, Sengupta S, Bertrand D, Weerasinghe A, et al. Comprehensive characterization of cancer driver genes and mutations. Cell. 2018;173:371–85.
Aguet F, Brown AA, Castel SE, Davis JR, He Y, Jo B, et al. Genetic effects on gene expression across human tissues. Nature. 2017;550:204–13.
Edgar R, Domrachev M, Lash AE. Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002;30:207–10.
Yin H, Li M, Xia L, He C, Zhang Z. Computational determination of gene age and characterization of evolutionary dynamics in human. Brief Bioinform. 2019;20:2141–9.
Vishnoi A, Kryazhimskiy S, Bazykin GA, Hannenhalli S, Plotkin JB. Young proteins experience more variable selection pressures than old proteins. Genome Res. 2010;20:1574–81.
Albà MM, Castresana J. Inverse relationship between evolutionary rate and age of mammalian genes. Mol Biol Evol. 2005;22:598–606.
Warnefors M, Eyre-Walker A. The accumulation of gene regulation through time. Genome Biol Evol. 2011;3:667–73.
Zhao Y, Chen Y, Jin M, Wang J. The crosstalk between m6A RNA methylation and other epigenetic regulators: a novel perspective in epigenetic remodeling. Theranostics. 2021;11:4549–66.
Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, et al. Recognition of RNA N 6 -methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20:285–95.
Zhang Z, Chen LQ, Zhao YL, Yang CG, Roundtree IA, Zhang Z, et al. Single-base mapping of m6A by an antibody-independent method. Sci Adv. 2019;5:250–3.
Lawrence MS, Stojanov P, Polak P, Kryukov GV, Cibulskis K, Sivachenko A, et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature. 2013;499:214–8.
Zhao M, Kim P, Mitra R, Zhao J, Zhao Z. TSGene 2.0: an updated literature-based knowledgebase for tumor suppressor genes. Nucleic Acids Res. 2016;44:D1023–31.
Lever J, Zhao EY, Grewal J, Jones MR, Jones SJM. CancerMine: a literature-mined resource for drivers, oncogenes and tumor suppressors in cancer. Nat Methods. 2019;16:505–7.
Dweep H, Sticht C, Pandey P, Gretz N. MiRWalk—database: prediction of possible miRNA binding sites by ‘ walking’ the genes of three genomes. J Biomed Inf. 2011;44:839–47.
von Mering C, Jensen LJ, Snel B, Hooper SD, Krupp M, Foglierini M, et al. STRING: Known and predicted protein-protein associations, integrated and transferred across organisms. Nucleic Acids Res. 2005;33:D433–7.
Smit AFA, Hubley R, Green P. RepeatMasker Open-4.0. 2013–2015. http://www.repeatmasker.org.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7.
Chen CL, Rappailles A, Duquenne L, Huvet M, Guilbaud G, Farinelli L, et al. Impact of replication timing on non-CpG and CpG substitution rates in mammalian genomes. Genome Res. 2010;20:447–57.
Lieberman-Aiden E, Van Berkum NL, Williams L, Imakaev M, Ragoczy T, Telling A, et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science. 2009;326:289–93.
Brouwer-Visser J, Cheng WY, Bauer-Mehren A, Maisel D, Lechner K, Andersson E, et al. Regulatory T-cell genes drive altered immune microenvironment in adult solid cancers and allow for immune contextual patient subtyping. Cancer Epidemiol Biomark Prev. 2018;27:103–12.
Guo M, Tomoshige K, Meister M, Muley T, Fukazawa T, Tsuchiya T, et al. Gene signature driving invasive mucinous adenocarcinoma of the lung. EMBO Mol Med. 2017;9:462–81.
Gerlinger M, Horswell S, Larkin J, Rowan AJ, Salm MP, Varela I, et al. Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat Genet. 2014;46:225–33.
Acknowledgements
The results shown here are in part based upon data generated by TCGA Research Network: https://www.cancer.gov/tcga.
Funding
This work was supported by the Shenzhen Science and Technology Programme (JCYJ20180508161604382 awarded to WKKW), Shenzhen Science and Technology Innovation Commission, and National Natural Science Foundation of China (NSFC; 82103245).
Author information
Authors and Affiliations
Contributions
WKKW, HC and MTVC conceived and supervised the study. DH and XW performed bioinformatic analysis. DH, HC and WKKW wrote the manuscript. The remaining authors analyzed the data and assisted in editing. All authors approved the final version to be published and agree to be accountable for all aspects of the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Huang, D., Wang, X., Huang, Z. et al. 3′untranslated regions of tumor suppressor genes evolved specific features to favor cancer resistance. Oncogene 41, 3278–3288 (2022). https://doi.org/10.1038/s41388-022-02343-5
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-022-02343-5