Abstract
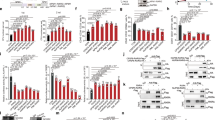
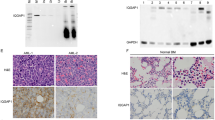
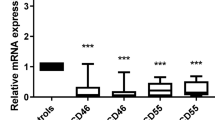
Rho GTPase-activating protein 4 (ARHGAP4) is an important Rho family GTPase-activating protein that is strongly associated with the onset and progression of some tumors. We found that ARHGAP4 mRNA and protein are overexpressed in human acute myeloid leukemia (AML) patients and are associated with a poor prognosis. ARHGAP4 knockdown significantly impairs viability and colony formation capacity and induces apoptosis in AML cells. Further results demonstrate that ARHGAP4 deletion impairs AML progression in vivo. Interestingly, DRAM1 signaling is significantly activated in AML cells with ARHGAP4 knockdown. Our results also indicated that ARHGAP4 might function in AML cells by binding with p53 to inhibit DRAM1. Moreover, knockdown of DRAM1 rescues the defects of ARHGAP4 in AML cells. This newly described role of the ARHGAP4/DRAM1 axis in regulating AML progression may have important therapeutic implications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The RNA-seq data discussed in this publication have been deposited in BioProject database with the accession code PRJNA956986. The datasets obtained and analyzed during the current study were made available from the corresponding authors through request.
References
Nechiporuk T, Kurtz SE, Nikolova O, Liu TT, Jones CL, D’Alessandro A, et al. The TP53 Apoptotic Network Is a Primary Mediator of Resistance to BCL2 Inhibition in AML Cells. Cancer Discov. 2019;9:910–25.
Duy C, Li M, Teater M, Meydan C, Garrett-Bakelman FE, Lee TC, et al. Chemotherapy Induces Senescence-Like Resilient Cells Capable of Initiating AML Recurrence. Cancer Discov. 2021;11:1542–61.
Gil-Sierra MD, Briceno-Casado MP, Sierra-Sanchez JF. Ivosidenib and Azacitidine in IDH1-Mutated AML. N Engl J Med. 2022;386:2535–6.
Freeman SD, Valk P. Transplant in older adults with AML: genomic wheat and chaff. Blood. 2022;139:3459–61.
Qiu KY, Liao XY, Liu Y, Huang K, Li Y, Fang JP, et al. Poor outcome of pediatric patients with acute myeloid leukemia harboring high FLT3/ITD allelic ratios. Nat Commun. 2022;13:3679.
Lyu Y, Lou J, Yang Y, Feng J, Hao Y, Huang S, et al. Dysfunction of the WT1-MEG3 signaling promotes AML leukemogenesis via p53-dependent and -independent pathways. Leukemia. 2017;31:2543–51.
He XX, Wan JB, Yang XN, Zhang XZ, Huang D, Li X, et al. Bone marrow niche ATP levels determine leukemia-initiating cell activity via P2X7 in leukemic models. J Clin Investig. 2021;131:e140242.
Crosas-Molist E, Samain R, Kohlhammer L, Orgaz JL, George SL, Maiques O, et al. Rho GTPase signaling in cancer progression and dissemination. Physiol Rev. 2022;102:455–510.
Liu R, Wang Y, Li BX, Wang H, Guan F, Tan ZQ, et al. Screening differentially expressed proteins from co-cultured hematopoietic cells and bone marrow-derived stromal cells by quantitative proteomics (SILAC) method. Clin Proteom. 2019;16:32.
Shen Y, Xu L, Ning Z, Liu L, Lin J, Chen H, et al. ARHGAP4 regulates the cell migration and invasion of pancreatic cancer by the HDAC2/beta-catenin signaling pathway. Carcinogenesis. 2019;40:1405–14.
Kang N, Matsui TS, Liu S, Fujiwara S, Deguchi S. Comprehensive analysis on the whole Rho-GAP family reveals that ARHGAP4 suppresses EMT in epithelial cells under negative regulation by Septin9. FASEB J. 2020;34:8326–40.
Shen Y, Chen G, Gao H, Li Y, Zhuang L, Meng Z, et al. miR-939-5p Contributes to the Migration and Invasion of Pancreatic Cancer by Targeting ARHGAP4. Onco Targets Ther. 2020;13:389–99.
Kang N, Matsui TS, Liu S, Deguchi S. ARHGAP4-SEPT2-SEPT9 complex enables both up- and down-modulation of integrin-mediated focal adhesions, cell migration, and invasion. Mol Biol Cell. 2021;32:ar28.
Bai Y, Chen YB, Kong XD. Contiguous 22.1-kb deletion embracing AVPR2 and ARHGAP4 genes at novel breakpoints leads to nephrogenic diabetes insipidus in a Chinese pedigree. Bmc Nephrol. 2018;19:26.
Geng J, Zhang R, Yuan X, Xu H, Zhu Z, Wang X, et al. DRAM1 plays a tumor suppressor role in NSCLC cells by promoting lysosomal degradation of EGFR. Cell Death Dis. 2020;11:768.
Lu T, Zhu Z, Wu J, She H, Han R, Xu H, et al. DRAM1 regulates autophagy and cell proliferation via inhibition of the phosphoinositide 3-kinase-Akt-mTOR-ribosomal protein S6 pathway. Cell Commun Signal. 2019;17:28.
Hu W, Chen S, Thorne RF, Wu M. TP53, TP53 Target Genes (DRAM, TIGAR), and Autophagy. Adv Exp Med Biol. 2019;1206:127–49.
Takahashi M, Kakudo Y, Takahashi S, Sakamoto Y, Kato S, Ishioka C. Overexpression of DRAM enhances p53-dependent apoptosis. Cancer Med. 2013;2:1–10.
Meijer AH, van der Vaart M. DRAM1 promotes the targeting of mycobacteria to selective autophagy. Autophagy. 2014;10:2389–91.
van der Vaart M, Korbee CJ, Lamers GE, Tengeler AC, Hosseini R, Haks MC, et al. The DNA damage-regulated autophagy modulator DRAM1 links mycobacterial recognition via TLR-MYD88 to autophagic defense [corrected]. Cell Host Microbe. 2014;15:753–67.
Nagata M, Arakawa S, Yamaguchi H, Torii S, Endo H, Tsujioka M, et al. Dram1 regulates DNA damage-induced alternative autophagy. Cell Stress. 2018;2:55–65.
Yu MQ, Jiang YG, Feng QL, Ouyang YA, Gan J. DRAM1 Protects Neuroblastoma Cells from Oxygen-Glucose Deprivation/Reperfusion-Induced Injury via Autophagy. Int J Mol Sci. 2014;15:19253–64.
Zhang Y, Li X, Li Y, Li Y, Wang Y, Zhu L, et al. DNA Damage-Regulated Autophagy Modulator 1 (DRAM1) Mediates Autophagy and Apoptosis of Intestinal Epithelial Cells in Inflammatory Bowel Disease. Dig Dis Sci. 2021;66:3375–90.
Chen C, Liang QY, Chen HK, Wu PF, Feng ZY, Ma XM, et al. DRAM1 regulates the migration and invasion of hepatoblastoma cells via autophagy-EMT pathway. Oncol Lett. 2018;16:2427–33.
Tan J, Zhang J, Wang MK, Wang YF, Dong MZ, Ma XF, et al. DRAM1 increases the secretion of PKM2-enriched EVs from hepatocytes to promote macrophage activation and disease progression in ALD. Mol Ther-Nucl Acids. 2022;27:375–89.
Feng WL, Yang X, Wang LN, Wang R, Yang FF, Wang H, et al. P2X7 promotes the progression of MLL-AF9 induced acute myeloid leukemia by upregulation of Pbx3. Haematologica. 2021;106:1278–89.
Zhang R, Varela M, Forn-Cuni G, Torraca V, van der Vaart M, Meijer AH. Deficiency in the autophagy modulator Dram1 exacerbates pyroptotic cell death of Mycobacteria-infected macrophages. Cell Death Dis. 2020;11:277.
Zeisig BB, Fung TK, Zarowiecki M, Tsai CT, Luo HC, Stanojevic B, et al. Functional reconstruction of human AML reveals stem cell origin and vulnerability of treatment-resistant MLL-rearranged leukemia. Sci Transl Med. 2021;13:582.
Curtiss BM, VanCampen J, Macaraeg J, Kong GL, Taherinasab A, Tsuchiya M, et al. PU.1 and MYC transcriptional network defines synergistic drug responses to KIT and LSD1 inhibition in acute myeloid leukemia. Leukemia. 2022;36:1781–93.
Stranahan AW, Berezniuk I, Chakraborty S, Feller F, Khalaj M, Park CY. Leukotrienes promote stem cell self-renewal and chemoresistance in acute myeloid leukemia. Leukemia. 2022;36:1575–84.
Leick MB, Silva H, Scarfo I, Larson R, Choi BD, Bouffard AA, et al. Non-cleavable hinge enhances avidity and expansion of CAR-T cells for acute myeloid leukemia. Cancer Cell. 2022;40:494–508.e495.
Shen Y, Chen G, Zhuang L, Xu L, Lin J, Liu L. ARHGAP4 mediates the Warburg effect in pancreatic cancer through the mTOR and HIF-1alpha signaling pathways. Onco Targets Ther. 2019;12:5003–12.
Katoh M, Katoh M. Characterization of human ARHGAP10 gene in silico. Int J Oncol. 2004;25:1201–6.
Shen YH, Chen G, Zhuang LP, Xu LT, Lin JH, Liu LM. ARHGAP4 mediates the Warburg effect in pancreatic cancer through the mTOR and HIF-1 alpha signaling pathways. Oncotargets Ther. 2019;12:5003–12.
Zhang Z, Lu Y, Qi Y, Xu Y, Wang S, Chen F, et al. CDK19 regulates the proliferation of hematopoietic stem cells and acute myeloid leukemia cells by suppressing p53-mediated transcription of p21. Leukemia. 2022;36:956–69.
Barthet VJA, Mrschtik M, Kania E, McEwan DG, Croft D, O'Prey J, et al. DRAM-4 and DRAM-5 are compensatory regulators of autophagy and cell survival in nutrient-deprived conditions. FEBS J. 2022;289:3752–69.
Acknowledgements
We thank Prof. Jinyong Wang for gifting CD45.1 mice, Yang Liu and Haiying Ran for technical support in flow cytometry.
Funding
This work was supported by grants from the Scientific Research Project of PLA (2018XLC306); Science Foundations of Army Medical Center (No.2019CXJSB020); The Key Program of the Natural Science Foundation of China (No. 81930090).
Author information
Authors and Affiliations
Contributions
YQ, MH and CH designed and performed the experiments, analyzed the data and wrote the paper; FC, YS and MZ contributed to animal experiments and data analysis; JZ, MC and SW contributed to the in vitro experiments; HX and ZZ provided human samples; JW and HG contributed to the initial experimental design and discussed the manuscript; JW and DZ conceived and supervised the study, analyzed the data, and wrote and revised the manuscript. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qi, Y., Hu, M., Han, C. et al. ARHGAP4 promotes leukemogenesis in acute myeloid leukemia by inhibiting DRAM1 signaling. Oncogene 42, 2547–2557 (2023). https://doi.org/10.1038/s41388-023-02770-y
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-023-02770-y
This article is cited by
-
E2F expression profiling-based subtypes in head and neck squamous cell carcinoma: clinical relevance, prognostic implications, and personalized therapy
World Journal of Surgical Oncology (2025)
-
DRAM1 enhances the proliferation and metastasis of gastric cancer through the PI3K/AKT/mTOR signaling pathway and energy metabolism
Scientific Reports (2025)
-
Multiomic single cell sequencing identifies stemlike nature of mixed phenotype acute leukemia
Nature Communications (2024)