Abstract
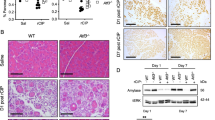
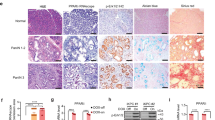
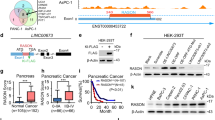
Oncogenic mutations in KRAS are present in ~95% of patients diagnosed with pancreatic ductal adenocarcinoma (PDAC) and are considered the initiating event of pancreatic intraepithelial neoplasia (PanIN) precursor lesions. While it is well established that KRAS mutations drive the activation of oncogenic kinase cascades during pancreatic oncogenesis, the effects of oncogenic KRAS signaling on regulation of phosphatases during this process is not fully appreciated. Protein Phosphatase 2A (PP2A) has been implicated in suppressing KRAS-driven cellular transformation and low PP2A activity is observed in PDAC cells compared to non-transformed cells, suggesting that suppression of PP2A activity is an important step in the overall development of PDAC. In the current study, we demonstrate that KRASG12D induces the expression of an endogenous inhibitor of PP2A activity, Cancerous Inhibitor of PP2A (CIP2A), and phosphorylation of the PP2A substrate, c-MYC. Consistent with these findings, KRASG12D sequestered the specific PP2A subunit responsible for c-MYC degradation, B56α, away from the active PP2A holoenzyme in a CIP2A-dependent manner. During PDAC initiation in vivo, knockout of B56α promoted KRASG12D tumorigenesis by accelerating acinar-to-ductal metaplasia (ADM) and the formation of PanIN lesions. The process of ADM was attenuated ex vivo in response to pharmacological re-activation of PP2A utilizing direct small molecule activators of PP2A (SMAPs). Together, our results suggest that suppression of PP2A-B56α through KRAS signaling can promote the MYC-driven initiation of pancreatic tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Data from the current study is available from the corresponding author upon reasonable request. RNA- and ATAC-seq have been deposited in GEO repository under: GSE250101.
References
Hobbs GA, Der CJ, Rossman KL. RAS isoforms and mutations in cancer at a glance. J Cell Sci. 2016;129:1287–92.
Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023;73:17–48.
Bryant KL, Mancias JD, Kimmelman AC, Der CJ. KRAS: feeding pancreatic cancer proliferation. Trends Biochem Sci. 2014;39:91–100.
Hingorani SR, Petricoin EF, Maitra A, Rajapakse V, King C, Jacobetz MA. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell. 2003;4:437–450.
Collins MA, Bednar F, Zhang Y, Brisset J-C, Galbán S, Galbán CJ, et al. Oncogenic Kras is required for both the initiation and maintenance of pancreatic cancer in mice. J Clin Invest. 2012;122:639–53.
Storz P. Acinar cell plasticity and development of pancreatic ductal adenocarcinoma. Nat Rev Gastroenterol. 2017;14:296–304.
Collado M, Gil J, Efeyan A, Guerra C, Schuhmacher AJ, Barradas M, et al. Senescence in premalignant tumours. Nature. 2005;436:642.
Sablina AA, Hector M, Colpaert N, Hahn WC. Identification of PP2A complexes and pathways involved in cell transformation. Cancer Res. 2010;70:10474–84.
Sangodkar J, Farrington CC, McClinch K, Galsky MD, Kastrinsky DB, Narla G. All roads lead to PP2A: exploiting the therapeutic potential of this phosphatase. FEBS J. 2016;283:1004–24.
Hydbring P, Larsson L-G. Cdk2: a key regulator of the senescence control function of Myc. Aging Albany 2010;2:244–50.
Karnoub AE, Weinberg RA. Ras oncogenes: split personalities. Nat Rev Mol Cell Biol. 2008;9:517–31.
Letourneux C, Rocher G, Porteu F. B56-containing PP2A dephosphorylate ERK and their activity is controlled by the early gene IEX-1 and ERK. EMBO J. 2006;25:727–38.
Lin SS, Bassik MC, Suh H, Nishino M, Arroyo JD, Hahn WC, et al. PP2A regulates BCL-2 phosphorylation and proteasome-mediated degradation at the endoplasmic reticulum*. J Biol Chem. 2006;281:23003–12.
Farrell AS, Allen-Petersen B, Daniel CJ, Wang X, Wang Z, Rodriguez S, et al. Targeting inhibitors of the tumor suppressor PP2A for the treatment of pancreatic cancer. Mol Cancer Res. 2014;12:924–39.
Janghorban M, Langer EM, Wang X, Zachman D, Daniel CJ, Hooper J, et al. The tumor suppressor phosphatase PP2A-B56α regulates stemness and promotes the initiation of malignancies in a novel murine model. PLoS ONE. 2017;12:e0188910.
Kauko O, Laajala TD, Jumppanen M, Hintsanen P, Suni V, Haapaniemi P, et al. Label-free quantitative phosphoproteomics with novel pairwise abundance normalization reveals synergistic RAS and CIP2A signaling. Sci Rep. 2015;5:13099.
Chang W-H, Liu Y, Hammes EA, Bryant KL, Cerione RA, Antonyak MA. Oncogenic RAS promotes MYC protein stability by upregulating the expression of the inhibitor of apoptosis protein family member Survivin. J Biol Chem. 2023;299:102842.
Wu Y, Gu T-T, Zheng P-S. CIP2A cooperates with H-Ras to promote epithelial–mesenchymal transition in cervical-cancer progression. Cancer Lett. 2015;356:646–55.
Arnold HK, Sears RC. Protein phosphatase 2A regulatory subunit B56α associates with c-Myc and negatively regulates c-Myc accumulation. Mol Cell Biol. 2006;26:2832–44.
Farrell AS, Sears RC. MYC Degradation. CSH Perspect Med. 2014;4:a014365.
Wang J, Okkeri J, Pavic K, Wang Z, Kauko O, Halonen T, et al. Oncoprotein CIP2A is stabilized via interaction with tumor suppressor PP2A/B56. EMBO Rep. 2017;18:437–50.
Sears R, Leone G, DeGregori J, Nevins JR. Ras enhances Myc protein stability. Mol Cell. 1999;3:169–79.
Seeling JM, Miller JR, Gil R, Moon RT, White R, Virshup DM. Regulation of β-Catenin signaling by the B56 subunit of protein phosphatase 2A. Science. 1999;283:2089–91.
Martinez AKF, Storz P. Mimicking and manipulating pancreatic acinar-to-ductal metaplasia in 3-dimensional cell culture. J Vis Exp. (144), e59096, https://doi.org/10.3791/59096 (2019).
Shi G, DiRenzo D, Qu C, Barney D, Miley D, Konieczny SF. Maintenance of acinar cell organization is critical to preventing Kras-induced acinar-ductal metaplasia. Oncogene. 2013;32:1950–8.
Tinsley SL, Allen-Petersen BL. PP2A and cancer epigenetics: a therapeutic opportunity waiting to happen. NAR Cancer. 2022;4:zcac002-.
Muthalagu N, Monteverde T, Raffo-Iraolagoitia X, Wiesheu R, Whyte D, Hedley A, et al. Repression of the Type I interferon pathway underlies MYC- and KRAS-dependent evasion of NK and B cells in pancreatic ductal adenocarcinoma. Cancer Discov. 2020;10:872–87.
Aakula A, Sharma M, Tabaro F, Nätkin R, Kamila J, Honkanen H, et al. RAS and PP2A activities converge on epigenetic gene regulation. Life Sci Alliance. 2023;6:e202301928.
Vogt M, Stankovic ND, Garcia YC, Hofstetter J, Schneider K, Kuybu F. et al. Targeting MYC effector functions in pancreatic cancer by inhibiting the ATPase RUVBL1/2. Gut. 2024;73:1509–1528.
Thomas D, Radhakrishnan P. Tumor-stromal crosstalk in pancreatic cancer and tissue fibrosis. Mol Cancer. 2019;18.
Leonard D, Huang W, Izadmehr S, O'Connor CM, Wiredja DD, Wang Z. Selective PP2A enhancement through biased heterotrimer stabilization. Cell. 2020;181:688–701.
Kemp SB, Cheng N, Markosyan N, Sor R, Kim I-K, Hallin J, et al. Efficacy of a small molecule inhibitor of KrasG12D in immunocompetent models of pancreatic cancer. Cancer Discov. 2022;13:298–311.
Hingorani SR, Wang L, Multani AS, Combs C, Deramaudt TB, Hruban RH, et al. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell. 2005;7:469–83.
Hosein AN, Dougan SK, Aguirre AJ, Maitra A. Translational advances in pancreatic ductal adenocarcinoma therapy. Nat Cancer. 2022;3:272–86.
Sodir NM, Kortlever RM, Barthet VJA, Campos T, Pellegrinet L, Kupczak S. et al. Myc instructs and maintains pancreatic adenocarcinoma phenotype. Cancer Discov. 2020;10(4):588–607.
Khanna A, Okkeri J, Bilgen T, Tiirikka T, Vihinen M, Visakorpi T, et al. ETS1 mediates MEK1/2-dependent overexpression of cancerous inhibitor of protein phosphatase 2A (CIP2A) in human cancer cells. PLoS ONE. 2011;6:e17979.
Sears R, Nuckolls F, Haura E, Taya Y, Tamai K, Nevins JR. Multiple Ras-dependent phosphorylation pathways regulate Myc protein stability. Genes Dev. 2000;14:2501–14.
Pavic K, Gupta N, Omella JD, Derua R, Aakula A, Huhtaniemi R, et al. Structural mechanism for inhibition of PP2A-B56α and oncogenicity by CIP2A. Nat Commun. 2023;14:1143.
Cristóbal I, Torrejón B, Madoz-Gúrpide J, Rojo F, García-Foncillas J. PP2A plays a key role in inflammation and cancer through tristetraprolin activation. Ann Rheum Dis. 2017;76:e11.
McHugh WM, Russell WW, Fleszar AJ, Rodenhouse PE, Rietberg SP, Sun L, et al. Protein phosphatase 2A activation attenuates inflammation in murine models of acute lung injury. Am J Physiol-Lung Cell Mol Physiol. 2016;311:L903–12.
Clark AR, Ohlmeyer M. Protein phosphatase 2A as a therapeutic target in inflammation and neurodegeneration. Pharmacol Ther. 2019;201:181–201.
Sangodkar J, Perl A, Tohme R, Kiselar J, Kastrinsky DB, Zaware N, et al. Activation of tumor suppressor protein PP2A inhibits KRAS-driven tumor growth. J Clin Investig. 2017;127:2081–90.
Tohmé R, Izadmehr S, Gandhe S, Tabaro G, Vallabhaneni S, Thomas A, et al. Direct activation of PP2A for the treatment of tyrosine kinase inhibitor–resistant lung adenocarcinoma. JCI Insight. 2019;4:e125693.
Kauko O, O’Connor CM, Kulesskiy E, Sangodkar J, Aakula A, Izadmehr S, et al. PP2A inhibition is a druggable MEK inhibitor resistance mechanism in KRAS-mutant lung cancer cells. Sci Transl Med. 2018;10:eaaq1093.
McClinch K, Avelar RA, Callejas D, Izadmehr S, Wiredja D, Perl A, et al. Small-Molecule Activators of Protein Phosphatase 2A for the Treatment of Castration-Resistant Prostate Cancer. Cancer Res. 2018;78:2065–80.
Avelar RA, Armstrong AJ, Carvette G, Gupta R, Puleo N, Colina JA, et al. Small molecule mediated stabilization of PP2A modulates the Homologous Recombination pathway and potentiates DNA damage-induced cell death. Mol Cancer Ther. 2023;22:599–615.
Allen-Petersen BL, Risom T, Feng Z, Wang Z, Jenny ZP, Thoma MC, et al. Activation of PP2A and inhibition of mTOR synergistically reduce MYC signaling and decrease tumor growth in pancreatic ductal adenocarcinoma. Cancer Res. 2018;79:209–19.
Strickler JH, Satake H, George TJ, Yaeger R, Hollebecque A, Garrido-Laguna I, et al. Sotorasib in KRAS p.G12C–mutated advanced pancreatic cancer. N Engl J Med. 2022;388:33–43.
Zhang J, Darman L, Hassan MS, Holzen UV, Awasthi N. Targeting KRAS for the potential treatment of pancreatic ductal adenocarcinoma: Recent advancements provide hope (Review). Oncol Rep. 2023;50:206.
Federico AD, Ricciotti I, Favorito V, Michelina SV, Scaparone P, Metro G, et al. Resistance to KRAS G12C inhibition in non-small cell lung cancer. Curr Oncol Rep. 2023;25:1017–29.
Aiello NM, Rhim AD, Stanger BZ. Orthotopic injection of pancreatic cancer cells. Cold Spring Harb Protoc. 2016;2016:pdb.prot078360.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–90.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Liao Y, Smyth GK, Shi W. The subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 2013;41:e108–e108.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnetJ. 2011;17:10–2.
Li H, Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 2009;25:1754–60.
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008;9:R137.
Ramírez F, Ryan DP, Grüning B, Bhardwaj V, Kilpert F, Richter AS, et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 2016;44:W160–5.
Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26:841–2.
Yu G, Wang L-G, He Q-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics. 2015;31:2382–3.
Heberle H, Meirelles GV, Silva FR, da, Telles GP, Minghim R. InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinform. 2015;16:169.
Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016;44:W90–7.
Daniel CJ, Sun X-X, Chen Y, Zhang X, Dai M-S, Sears RC. The Myc gene, methods and protocols. Methods Mol Biol. 2021;2318:69–85.
Acknowledgements
We would like to thank the Pasca di Magliano lab (University of Michigan) for providing cell lines and historical histology slides for use; the Sears lab (OHSU) for providing cell lines, antibodies, mouse models, and technical support; the Ogris lab for providing antibodies; and finally, the Narla lab (University of Michigan) for providing antibodies, DT061 stocks, and technical support for experiments. We would like to thank the Purdue Institute for Cancer Research (NIH grant P30 CA023168), the Purdue Histology Core, the Collaborative Core for Cancer Bioinformatics (C3B), and the Purdue Genomics Facility for their contributions to the data produced in this publication. We would also like to thank all the members of the BAP lab for editing of the manuscript and other helpful suggestions. G.N. acknowledges the support of the Rogel Cancer Center and is a Rogel Scholar. RC Sears was supported by NCI U01 CA224012, R01 CA186241, and DoD PA210068 and the Brenden-Colson Center Foundation. S. L. Tinsley was supported by the Frederick N. Andrews Fellowship and the SIRG grant administered through the Institute for Cancer Research. R. A. Shelley and ERD. Chianis were supported by the Institute for Cancer Research Summer Undergraduate Research Experience and support from the Purdue University Institute for Cancer Research, P30CA023168. B. L. Allen-Petersen was supported by the NIH NCI 1K22CA237620-01A1, Pancreatic Cancer Action Network 22-20-ALLE, Concern Foundation, and Ross-Lynn Scholars Research Grant.
Author information
Authors and Affiliations
Contributions
BLA-P, RCS, and SLT designed experiments. SLT, ERDC, RAS, GKM, GB, MCT, IAE, CJD, and BLA-P performed and analyzed experiments. GN and MPdM provided key resources. AD, HK, NAL, and ECD performed bioinformatic analysis. LCSA and LS performed image analysis. BLA-P and RCS secured funds and provided supervision. BLA-P and SLT wrote and revised the manuscript. All authors reviewed and finalized the manuscript.
Corresponding author
Ethics declarations
Competing interests
G.N. receives research support from RAPPTA Therapeutics, and has an equity interest and receives consulting fees from RAPPTA Therapeutics. R.C.S serves on the scientific advisory board for RAPPTA and Larkspur, and receives sponsored research support from Cardiff Oncology and AstraZeneca. L.S. receives sponsored research support from Eli Lilly and Company. All other authors have no potential conflicts of interest.
Ethics approval and consent to participate
All methods were performed in accordance with relevant guidelines and regulations. All animal studies were performed in compliance with Purdue University (West Lafayette, IN) animal use guidelines after approval by the Purdue Institutional Animal Care and Use Committee (PACUC #1910001962).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tinsley, S.L., Chianis, E.R.D., Shelley, R.A. et al. KRAS-mediated upregulation of CIP2A promotes suppression of PP2A-B56α to initiate pancreatic cancer development. Oncogene 43, 3673–3687 (2024). https://doi.org/10.1038/s41388-024-03196-w
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-024-03196-w
This article is cited by
-
Enhancing KRAS G12D inhibitor sensitivity in pancreatic cancer through SHP2/PI3K pathway
Medical Oncology (2025)