Abstract
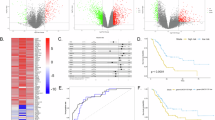
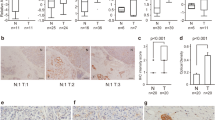
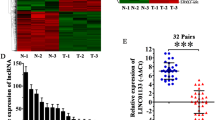
Pancreatic ductal adenocarcinoma (PDAC) is associated with a high mortality rate and short survival time. Long noncoding RNAs (lncRNAs) play a significant role in the progression of PDAC. However, prognostic lncRNAs associated with overall survival (OS) in patients with PDAC remain elusive. RNA sequencing was used to identify differential lncRNA expression between short-term and long-term PDAC patients. We identified a novel lncRNA (ENSG00000253924), termed ST18-AS1 (ST18-associated lncRNA), that is highly expressed in the tissues of long-term PDAC patients. High ST18-AS1 expression was correlated with favorable clinical outcomes, and the upregulation of ST18-AS1 expression in PDAC cell lines suppressed cell proliferation and promoted apoptosis both in vivo and in vitro. The key downstream target regulated by ST18-AS1 was Suppression of tumorigenicity 18 (ST18). We also found that ST18 expression was lower in PDAC tissues compared to non-tumorous adjacent tissues (NATs) and that higher ST18 expression was correlated with better clinical outcomes. Accordingly, the forced expression of ST18 inhibited proliferation and promoted apoptosis in tumor cells. Mechanistic studies showed that ST18-AS1 maintained the stability of ST18 mRNA by binding to Fused in sarcoma (FUS) and anchoring FUS in the cytoplasm. Overall, we identified ST18-AS1 as a novel biomarker that inhibits PDAC cell proliferation and promotes PDAC cell apoptosis through ST18. Targeting ST18-AS1/ST18 may be a potential therapeutic strategy for treating PDAC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The datasets in this study are available from the corresponding author on reasonable request.
References
Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023;73:17–48.
Bengtsson A, Andersson R, Ansari D. The actual 5-year survivors of pancreatic ductal adenocarcinoma based on real-world data. Sci Rep. 2020;10:16425.
Blackford AL, Canto MI, Klein AP, Hruban RH, Goggins M. Recent trends in the incidence and survival of stage 1A pancreatic cancer: a surveillance, epidemiology, and end results analysis. J Natl Cancer Inst. 2020;112:1162–9.
Stoffel EM, Brand RE, Goggins M. Pancreatic cancer: changing epidemiology and new approaches to risk assessment, early detection, and prevention. Gastroenterology. 2023;164:752–65.
Wahler IL, Damanakis A, Grosse Hokamp N, Bruns C, Schmidt T. Therapy of Locally Advanced and Oligometastatic Pancreatic Adenocarcinoma. Cancers (Basel). 2023;15:5881.
Kleeff J, Korc M, Apte M, La Vecchia C, Johnson CD, Biankin AV, et al. Pancreatic cancer. Nat Rev Dis Prim. 2016;2:16022.
Strobel O, Neoptolemos J, Jager D, Buchler MW. Optimizing the outcomes of pancreatic cancer surgery. Nat Rev Clin Oncol. 2019;16:11–26.
Bhandare MS, Gupta V, Chaudhari V, Nandy K, Ostwal V, Ramaswamy A, et al. Differential impact of incrementally elevated CA 19-9 levels on prognosis of resected pancreatic ductal adenocarcinoma. HPB (Oxford). 2024;26:1237–47.
Luo G, Jin K, Deng S, Cheng H, Fan Z, Gong Y, et al. Roles of CA19-9 in pancreatic cancer: biomarker, predictor and promoter. Biochim Biophys Acta Rev Cancer. 2021;1875:188409.
Statello L, Guo CJ, Chen LL, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22:96–118.
Mattick JS, Amaral PP, Carninci P, Carpenter S, Chang HY, Chen LL, et al. Long non-coding RNAs: definitions, functions, challenges and recommendations. Nat Rev Mol Cell Biol. 2023;24:430–47.
Matsui M, Corey DR. Non-coding RNAs as drug targets. Nat Rev Drug Discov. 2017;16:167–79.
Huang X, Zhi X, Gao Y, Ta N, Jiang H, Zheng J. LncRNAs in pancreatic cancer. Oncotarget. 2016;7:57379–90.
Mas AM, Huarte M. Long noncoding RNA signatures as cancer biomarkers. J Clin Oncol. 2023;41:3059–62.
Gao J, Zheng Y, Li L, Lu M, Chen X, Wang Y, et al. Integrated transcriptomics and epigenomics reveal chamber-specific and species-specific characteristics of human and mouse hearts. PLoS Biol. 2021;19:e3001229.
Jiang P, Li Y, Yang X, Zhou J, Wei P. Clinical Value of Differential lncRNA Expressions in Diagnosis of Giant Cell Tumor of Bone and Tumor Recurrence. Clin Lab. 2020;66:1381–9.
He J, Li F, Zhou Y, Hou X, Liu S, Li X, et al. LncRNA XLOC_006390 promotes pancreatic carcinogenesis and glutamate metabolism by stabilizing c-Myc. Cancer Lett. 2020;469:419–28.
Shi X, Guo X, Li X, Wang M, Qin R. Loss of Linc01060 induces pancreatic cancer progression through vinculin-mediated focal adhesion turnover. Cancer Lett. 2018;433:76–85.
Winkle M, El-Daly SM, Fabbri M, Calin GA. Noncoding RNA therapeutics - challenges and potential solutions. Nat Rev Drug Discov. 2021;20:629–51.
Nemeth K, Bayraktar R, Ferracin M, Calin GA. Non-coding RNAs in disease: from mechanisms to therapeutics. Nat Rev Genet. 2024;25:211–32.
Yee KS, Yu VC. Isolation and characterization of a novel member of the neural zinc finger factor/myelin transcription factor family with transcriptional repression activity. J Biol Chem. 1998;273:5366–74.
Jiang Y, Yu VC, Buchholz F, O’Connell S, Rhodes SJ, Candeloro C, et al. A novel family of Cys-Cys, His-Cys zinc finger transcription factors expressed in developing nervous system and pituitary gland. J Biol Chem. 1996;271:10723–30.
Jandrig B, Seitz S, Hinzmann B, Arnold W, Micheel B, Koelble K, et al. ST18 is a breast cancer tumor suppressor gene at human chromosome 8q11.2. Oncogene. 2004;23:9295–302.
Yang J, Siqueira MF, Behl Y, Alikhani M, Graves DT. The transcription factor ST18 regulates proapoptotic and proinflammatory gene expression in fibroblasts. FASEB J. 2008;22:3956–67.
Henry C, Close AF, Buteau J. A critical role for the neural zinc factor ST18 in pancreatic beta-cell apoptosis. J Biol Chem. 2014;289:8413–9.
Li Y, Kong R, Chen H, Zhao Z, Li L, Li J, et al. Overexpression of KLF5 is associated with poor survival and G1/S progression in pancreatic cancer. Aging (Albany NY). 2019;11:5035–57.
Wang S, Chen J, Li P, Chen Y. LINC01133 can induce acquired ferroptosis resistance by enhancing the FSP1 mRNA stability through forming the LINC01133-FUS-FSP1 complex. Cell Death Dis. 2023;14:767.
Liu C, Zhang Y, She X, Fan L, Li P, Feng J, et al. A cytoplasmic long noncoding RNA LINC00470 as a new AKT activator to mediate glioblastoma cell autophagy. J Hematol Oncol. 2018;11:77.
Xiong XD, Ren X, Cai MY, Yang JW, Liu X, Yang JM. Long non-coding RNAs: An emerging powerhouse in the battle between life and death of tumor cells. Drug Resist Updat. 2016;26:28–42.
Qi P, Du X. The long non-coding RNAs, a new cancer diagnostic and therapeutic gold mine. Mod Pathol. 2013;26:155–65.
Choudhari R, Sedano MJ, Harrison AL, Subramani R, Lin KY, Ramos EI, et al. Long noncoding RNAs in cancer: from discovery to therapeutic targets. Adv Clin Chem. 2020;95:105–47.
Coan M, Haefliger S, Ounzain S, Johnson R. Targeting and engineering long non-coding RNAs for cancer therapy. Nat Rev Genet. 2024;25:578–95.
Sasaki N, Toyoda M, Yoshimura H, Matsuda Y, Arai T, Takubo K, et al. H19 long non-coding RNA contributes to sphere formation and invasion through regulation of CD24 and integrin expression in pancreatic cancer cells. Oncotarget. 2018;9:34719–34.
Wong CH, Li CH, He Q, Chan SL, Tong JH, To KF, et al. Ectopic HOTTIP expression induces noncanonical transactivation pathways to promote growth and invasiveness in pancreatic ductal adenocarcinoma. Cancer Lett. 2020;477:1–9.
Kim K, Jutooru I, Chadalapaka G, Johnson G, Frank J, Burghardt R, et al. HOTAIR is a negative prognostic factor and exhibits pro-oncogenic activity in pancreatic cancer. Oncogene. 2013;32:1616–25.
Nath P, Bhuyan K, Bhattacharyya DK, Barah P. ETENLNC: An end to end lncRNA identification and analysis framework to facilitate construction of known and novel lncRNA regulatory networks. Comput Biol Chem. 2024;112:108140.
Schmitz SU, Grote P, Herrmann BG. Mechanisms of long noncoding RNA function in development and disease. Cell Mol Life Sci. 2016;73:2491–509.
Postepska-Igielska A, Giwojna A, Gasri-Plotnitsky L, Schmitt N, Dold A, Ginsberg D, et al. LncRNA Khps1 regulates expression of the proto-oncogene SPHK1 via triplex-mediated changes in chromatin structure. Mol Cell. 2015;60:626–36.
Zealy RW, Fomin M, Davila S, Makowsky D, Thigpen H, McDowell CH, et al. Long noncoding RNA complementarity and target transcripts abundance. Biochim Biophys Acta Gene Regul Mech. 2018;1861:224–34.
Yamazaki T, Souquere S, Chujo T, Kobelke S, Chong YS, Fox AH, et al. Functional domains of NEAT1 architectural lncRNA induce paraspeckle assembly through phase separation. Mol Cell. 2018;70:1038–53.e7.
Bridges MC, Daulagala AC, Kourtidis A. LNCcation: lncRNA localization and function. J Cell Biol. 2021;220:e202009045.
Chen LL. Linking long noncoding RNA localization and function. Trends Biochem Sci. 2016;41:761–72.
Ferre F, Colantoni A, Helmer-Citterich M. Revealing protein-lncRNA interaction. Brief Bioinform. 2016;17:106–16.
Yamaguchi A, Takanashi K. FUS interacts with nuclear matrix-associated protein SAFB1 as well as Matrin3 to regulate splicing and ligand-mediated transcription. Sci Rep. 2016;6:35195.
Funding
This work was supported by National Nature Science Foundation of China (82370561, 82070567).
Author information
Authors and Affiliations
Contributions
GW and YL conceived the research and designed the experiments; RW and GL illustrated the figures; LC, ZH, YJ and HL performed the experiments and analyzed the original data; LC, GW and YL analyzed and interpreted the data and assisted with the adjustments of directions; LC, HY and YQ wrote the manuscript; GL, LC and TS revised the manuscript; all authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All methods were performed in accordance with the relevant guidelines and regulations. The animal study was approved by the Animal Experiment Ethics Committee of The First Affiliated Hospital of Harbin Medical University (Approval No. IRB-AF/SC-04/02.0), Additionally, the use of human PDAC tissues was approved by The First Affiliated Hospital of Harbin Medical University (Approval No. 2023086).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cheng, L., Yu, H., Qin, Y. et al. The lncRNA ST18-AS1 suppresses pancreatic cancer progression by enhancing ST18 mRNA stability through anchoring FUS in the cytoplasm. Oncogene 44, 2850–2863 (2025). https://doi.org/10.1038/s41388-025-03455-4
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-025-03455-4