Abstract
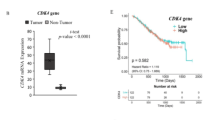
The cell membrane, as the boundary of the cellular system, plays an extremely important role in the regulation of cellular metabolism, substance transport, information exchange and cellular immunity. Complement component 1q subcomponent binding protein (C1QBP) is a ubiquitously expressed cellular protein, and although multiple studies have suggested that C1QBP may play an important role in cancer, the functions and mechanisms of C1QBP in the progression of many tumors remain unknown. We performed bioinformatics analysis and found that C1QBP was significantly overexpressed in a variety of tumor tissues and that high C1QBP expression correlates strongly with poor prognosis in tumor patients. Further validation in cholangiocarcinoma(CCA) revealed that C1QBP is the most upregulated membrane protein in CCA, where it is involved in energy metabolism, DNA repair, and tolerance to platinum-based chemotherapeutic agents. It also correlates strongly with CCA proliferation and poor prognosis, while silencing of C1QBP significantly inhibits CCA growth in mice. Mechanistic studies further demonstrated that c-MYC can upregulate C1QBP expression at the transcriptional level, subsequently influencing PAICS/FAK expression and promoting CCA growth. Interestingly, C1QBP also regulates c-MYC expression by increasing FAK phosphorylation, establishing a positive feedback loop that drives tumor progression. Additionally, we developed a novel siRNA delivery system, HA gel-siC1QBP, by encapsulating siC1QBP within hyaluronic acid-dopamine hydrogel-coated liposomes. In vivo experiments confirmed its ability to provide prolonged and stable C1QBP inhibition along with enhanced antitumor efficacy. In conclusion, our study suggests that C1QBP may serve as a valuable biomarker for tumour prognosis and that silencing C1QBP using HA gel-siC1QBP —either alone or combined with targeted/ immunotherapies —represents a promising therapeutic strategy against tumors.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The datasets analysed during the current study are available in the [GEO] repository.
References
Almén MS, Nordström KJ, Fredriksson R, Schiöth HB. Mapping the human membrane proteome: a majority of the human membrane proteins can be classified according to function and evolutionary origin. BMC Biol. 2009;7:50.
Gschwind A, Fischer OM, Ullrich A. The discovery of receptor tyrosine kinases: targets for cancer therapy. Nat Rev Cancer. 2004;4:361–70.
Gentles AJ, Newman AM, Liu CL, Bratman SV, Feng W, Kim D, et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat Med. 2015;21:938–45.
Kampen KR. Membrane proteins: the key players of a cancer cell. J Membr Biol. 2011;242:69–74.
Chen YB, Jiang CT, Zhang GQ, Wang JS, Pang D. Increased expression of hyaluronic acid binding protein 1 is correlated with poor prognosis in patients with breast cancer. J Surg Oncol. 2009;100:382–6.
Amamoto R, Yagi M, Song Y, Oda Y, Tsuneyoshi M, Naito S, et al. Mitochondrial p32/C1QBP is highly expressed in prostate cancer and is associated with shorter prostate-specific antigen relapse time after radical prostatectomy. Cancer Sci. 2011;102:639–47.
Yu H, Liu Q, Xin T, Xing L, Dong G, Jiang Q, et al. Elevated expression of hyaluronic acid binding protein 1 (HABP1)/P32/C1QBP is a novel indicator for lymph node and peritoneal metastasis of epithelial ovarian cancer patients. Tumour Biol. 2013;34:3981–7.
Gao H, Yao Q, Lan X, Li S, Wu J, Zeng G, et al. Elevated HABP1 protein expression correlates with progression and poor survival in patients with gastric cancer. Onco Targets Ther. 2016;9:6711–8.
Sinha S, Singh SK, Jangde N, Ray R, Rai V. p32 promotes melanoma progression and metastasis by targeting EMT markers, Akt/PKB pathway, and tumor microenvironment. Cell Death Dis. 2021;12:1012.
Egusquiza-Alvarez CA, Moreno-Londoño AP, Alvarado-Ortiz E, Ramos-Godínez MDP, Sarabia-Sánchez MA, Castañeda-Patlán MC, et al. Inhibition of Multifunctional Protein p32/C1QBP Promotes Cytostatic Effects in Colon Cancer Cells by Altering Mitogenic Signaling Pathways and Promoting Mitochondrial Damage. Int J Mol Sci. 2024;25:2712.
Fogal V, Zhang L, Krajewski S, Ruoslahti E. Mitochondrial/cell-surface protein p32/gC1qR as a molecular target in tumor cells and tumor stroma. Cancer Res. 2008;68:7210–8.
Zhai X, Liu K, Fang H, Zhang Q, Gao X, Liu F, et al. Mitochondrial C1qbp promotes differentiation of effector CD8+ T cells via metabolic-epigenetic reprogramming. Sci Adv. 2021;7:eabk0490.
Sarcognato S, Sacchi D, Fassan M, Fabris L, Cadamuro M, Zanus G, et al. Cholangiocarcinoma. Pathologica. 2021;113:158–69.
Banales JM, Marin JJG, Lamarca A, Rodrigues PM, Khan SA, Roberts LR, et al. Cholangiocarcinoma 2020: the next horizon in mechanisms and management. Nat Rev Gastroenterol Hepatol. 2020;17:557–88.
Dong LQ, Shi Y, Ma LJ, Yang LX, Wang XY, Zhang S, et al. Spatial and temporal clonal evolution of intrahepatic cholangiocarcinoma. J Hepatol. 2018;69:89–98.
Valle J, Wasan H, Palmer DH, Cunningham D, Anthoney A, Maraveyas A, et al. Cisplat in plus gemcitabine versus gemcitabine for biliary tract cancer. N Engl J Med. 2010;362:1273–81.
Goswami MT, Chen G, Chakravarthi BV, Pathi SS, Anand SK, Carskadon SL, et al. Role and regulation of coordinately expressed de novo purine biosynthetic enzymes PPAT and PAICS in lung cancer. Oncotarget. 2015;6:23445–61.
Qu ZY, Cui GY, Shi PJ, Wang HQ. Potential suppressive functions of microRNA-504 in cervical cancer cells malignant process were achieved by targeting PAICS and regulating EMT. Arch Gynecol Obstet. 2020;302:173–82.
Meng M, Chen Y, Jia J, Li L, Yang S. Knockdown of PAICS inhibits malignant proliferation of human breast cancer cell lines. Biol Res. 2018;51:24.
Huang N, Xu C, Deng L, Li X, Bian Z, Zhang Y, et al. PAICS contributes to gastric carcinogenesis and participates in DNA damage response by interacting with histone deacetylase 1/2. Cell Death Dis. 2020;11:507.
Keller KE, Doctor ZM, Dwyer ZW, Lee YS. SAICAR induces protein kinase activity of PKM2 that is necessary for sustained proliferative signaling of cancer cells. Mol Cell. 2014;53:700–9.
Du B, Zhang Z, Di W, Xu W, Yang L, Zhang S, et al. PAICS is related to glioma grade and can promote glioma growth and migration. J Cell Mol Med. 2021;25:7720–33.
Zhang J, Gao Q, Zhou Y, Dier U, Hempel N, Hochwald SN. Focal adhesion kinase-promoted tumor glucose metabolism is associated with a shift of mitochondrial respiration to glycolysis. Oncogene. 2016;35:1926–42.
Tapial Martínez P, López Navajas P, Lietha D. FAK Structure and Regulation by Membrane Interactions and Force in Focal Adhesions. Biomolecules. 2020;10:179.
Lang L, Tao J, Yang C, Li W. Tumor suppressive role of microRNA-4731-5p in breast cancer through reduction of PAICS-induced FAK phosphorylation. Cell Death Discov. 2022;8:154.
Diaz, Osterman CJ, Ozmadenci D, Kleinschmidt EG, Taylor KN, Barrie AM, et al. FAK activity sustains intrinsic and acquired ovarian cancer resistance to platinum chemotherapy. Elife. 2019;8:e47327.
Lin CY, Lee CH, Chuang YH, Lee JY, Chiu YY, Wu Lee YH, et al. Membrane protein-regulated networks across human cancers. Nat Commun. 2019;10:3131.
Eißmann M, Schwamb B, Melzer IM, Moser J, Siele D, Köhl U, et al. A functional yeast survival screen of tumor-derived cDNA libraries designed to identify anti-apoptotic mammalian oncogenes. PLoS ONE. 2013;8:e64873.
Schaller MD, Frisch SM. PND-1186 FAK inhibitor selectively promotes tumor cell apoptosis in three-dimensional environments. Cancer Biol Ther. 2010;9:791–3.
Kleinschmidt EG, Schlaepfer DD. Focal adhesion kinase signaling in unexpected places. Curr Opin Cell Biol. 2017;45:24–30.
Lei Y, Li X, Qin D, Zhang Y, Wang Y. gC1qR: A New Target for Cancer Immunotherapy. Front Immunol. 2023;14:1095943.
Herrera-Quiterio GA, Encarnación-Guevara S. The transmembrane proteins (TMEM) and their role in cell proliferation, migration, invasion, and epithelial-mesenchymal transition in cancer. Front Oncol. 2023;13:1244740.
Jager M, Blokzijl F, Kuijk E, Bertl J, Vougioukalaki M, Janssen R, et al. Deficiency of nucleotide excision repair is associated with mutational signature observed in cancer. Genome Res. 2019;29:1067–77.
Huang Y, Wang X, Niu X, Wang X, Jiang R, Xu T, et al. EZH2 suppresses the nucleotide excision repair in nasopharyngeal carcinoma by silencing XPA gene. Mol Carcinog. 2017;56:447–63.
Huo A, Xiong X. PAICS as a potential target for cancer therapy linking purine biosynthesis to cancer progression. Life Sci. 2023;331:122070.
Levy A, Alhazzani K, Dondapati P, Alaseem A, Cheema K, Thallapureddy K, et al. Focal Adhesion kinase in ovarian cancer: a potential therapeutic target for platinum and taxane-resistant tumors. Curr Cancer Drug Targets. 2019;19:179–88.
Tang X, Wu Y, Yang J, Zhu W. Regulating COX10-AS1 / miR-142-5p / PAICS axis inhibits the proliferation of non-small cell lung cancer. Bioengineered. 2021;12:4643–53.
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr, et al. Cancer genome landscapes. Science. 2013;339:1546–58. https://doi.org/10.1126/science.123.
Kaposi-Novak P, Libbrecht L, Woo HG, Lee YH, Sears NC, Coulouarn C, et al. Central role of c-Myc during malignant conversion in human hepatocarcinogenesis. Cancer Res. 2009;69:2775–82.
Yang H, Liu T, Wang J, Li TW, Fan W, Peng H, et al. Deregulated methionine adenosyltransferase α1, c-Myc, and Maf proteins together promote cholangiocarcinoma growth in mice and humans. Hepatology. 2016;64:439–55.
Brodeur GM, Seeger RC, Schwab M, Varmus HE, Bishop JM. Amplification of N-myc in untreated human neuroblastomas correlates with advanced disease stage. Science. 1984;224:1121–4.
Luo G, Li B, Duan C, Cheng Y, Xiao B, Yao F, et al. c-Myc promotes cholangiocarcinoma cells to overcome contact inhibition via the mTOR pathway. Oncol Rep. 2017;38:2498–506.
Acknowledgements
This study was supported by Natural Science Foundation of Hubei Province (Grants: 2023AFB1020(X.C.)). This study was supported by Natural Science Foundation of Hunan Province (Grant: 2023JJ60453 (B.T.)) and Hunan Provincial Health Commission Scientific Research Program (Grant :D202302085994 (B.T.)). This study was supported by National Natural Science Foundation of China (Grants: 82302971 (J.Z.)) and Hubei Provincial Natural Science Foundation of China(Grants: 2023AFB190 (J.Z.)). This study was supported by Natural Science Foundation of Hubei Province (Grants: 2024AFB667 (C.Z.)). This study was supported by National Natural Science Foundation of China (Grants: 81772800(P.L.) and 82072945(P.L.)).
Author information
Authors and Affiliations
Contributions
XC: Methodology, Investigation. MW: Methodology, Investigation. WL:Methodology, Visualization. BT: Methodology, Visualization. JZ:Visualization, Conceptualization. CZ: Visualization, Conceptualization. PL: Project administration, Conceptualization, Investigation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study was conducted in accordance with the principles of the Declaration of Helsinki principles. It was approved by the Animal Use and Care Committees at the Second Xiangya hospital, Central South University (approval number 20241070). All subjects have written informed consent.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cheng, X., Wang, M., Li, W. et al. C1QBP forms a positive feedback loop with the PAICS/FAK/C-MYC axis to promote cancer cell proliferation. Oncogene 44, 4363–4376 (2025). https://doi.org/10.1038/s41388-025-03568-w
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-025-03568-w