Abstract
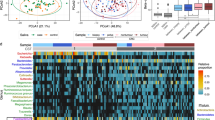
Circular RNAs (circRNAs) perform critical functions in cancer biology, commonly serving as microRNA (miRNA) sponges to modulate gene expression. Nevertheless, their participation in gut microbiota-driven colorectal cancer (CRC) has yet to be substantially investigated. Fusobacterium nucleatum (F. nucleatum), a well-recognized oncogenic bacterium in the human gut, has been implicated in CRC development, but the underlying mechanisms are not fully defined. In this study, we identified a novel circRNA, circPTBP3, which is the most significantly upregulated circRNA upon F. nucleatum infection, and is significantly upregulated in CRC tissues. CircPTBP3 is preferentially transcribed over its host gene PTBP3 in response to F. nucleatum through activation of the transcription factor ETS1. Functional assays demonstrated that circPTBP3 enhances CRC cell proliferation and tumor growth in vitro and in vivo. Mechanistically, circPTBP3 acts as a molecular sponge for miR-760, thereby relieving its suppression of the downstream target gene PUM1. In clinical CRC specimens, circPTBP3 expression showed a positive correlation with F. nucleatum abundance, PUM1 expression, larger tumor sizes, advanced TNM stages, and a negative correlation with miR-760 levels. These findings establish for the first time that circPTBP3 functions as a pivotal mediator of F. nucleatum ‘s oncogenicity, and reveal a novel F. nucleatum–circPTBP3–miR-760–PUM1 regulatory axis that promotes CRC progression. CircPTBP3 may serve as a potential biomarker and therapeutic target in F. nucleatum–associated colorectal carcinogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Brennan CA, Clay SL, Lavoie SL, Bae S, Lang JK, Fonseca-Pereira D, et al. Fusobacterium nucleatum drives a pro-inflammatory intestinal microenvironment through metabolite receptor- dependent modulation of IL-17 expression. Gut Microbes. 2021;13:1987780.
Wu J, Li Q, Fu X. Fusobacterium nucleatum contributes to the carcinogenesis of colorectal cancer by inducing inflammation and suppressing host immunity. Transl Oncol. 2019;12:846–51.
Yang Y, Weng W, Peng J, Hong L, Yang L, Toiyama Y, et al. Fusobacterium nucleatum increases proliferation of colorectal cancer cells and tumor development in mice by activating toll-like receptor 4 signaling to nuclear factor-κB, and up-regulating expression of microRNA-21. Gastroenterology. 2017;152:851–66.e24.
Yu T, Guo F, Yu Y, Sun T, Ma D, Han J, et al. Fusobacterium nucleatum promotes chemoresistance to colorectal cancer by modulating autophagy. Cell. 2017;170:548–63.e16.
Sanger HL, Klotz G, Riesner D, Gross HJ, Kleinschmidt AK. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci USA. 1976;73:3852–6.
Capel B, Swain A, Nicolis S, Hacker A, Walter M, Koopman P, et al. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell. 1993;73:1019–30.
Cocquerelle C, Mascrez B, Hétuin D, Bailleul B. Mis-splicing yields circular RNA molecules. FASEB J. 1993;7:155–60.
Zhang X, Xu Y, Qian Z, Zheng W, Wu Q, Chen Y, et al. circRNA_104075 stimulates YAP- dependent tumorigenesis through the regulation of HNF4a and may serve as a diagnostic marker in hepatocellular carcinoma. Cell Death Dis. 2018;9:1091.
Chen L, Nan A, Zhang N, Jia Y, Li X, Ling Y, et al. Circular RNA 100146 functions as an oncogene through direct binding to miR-361-3p and miR-615-5p in non-small cell lung cancer. Mol Cancer. 2019;18:13.
Liang HF, Zhang XZ, Liu BG, Jia GT, Li WL. Circular RNA circ-ABCB10 promotes breast cancer proliferation and progression through sponging miR-1271. Am J Cancer Res. 2017;7:1566–76.
Zhong Z, Huang M, Lv M, He Y, Duan C, Zhang L, et al. Circular RNA MYLK as a competing endogenous RNA promotes bladder cancer progression through modulating VEGFA/VEGFR2 signaling pathway. Cancer Lett. 2017;403:305–17.
Zhu M, Xu Y, Chen Y, Yan F. Circular BANP, an upregulated circular RNA that modulates cell proliferation in colorectal cancer. Biomed Pharmacother. 2017;88:138–44.
Zeng K, Chen X, Xu M, Liu X, Hu X, Xu T, et al. CircHIPK3 promotes colorectal cancer growth and metastasis by sponging miR-7. Cell Death Dis. 2018;9:417.
Yong W, Zhuoqi X, Baocheng W, Dongsheng Z, Chuan Z, Yueming S. Hsa_circ_0071589 promotes carcinogenesis via the miR-600/EZH2 axis in colorectal cancer. Biomed Pharmacother. 2018;102:1188–94.
Li X, Wang J, Zhang C, Lin C, Zhang J, Zhang W, et al. Circular RNA circITGA7 inhibits colorectal cancer growth and metastasis by modulating the Ras pathway and upregulating transcription of its host gene ITGA7. J Pathol. 2018;246:166–79.
Yao B, Zhang Q, Yang Z, An F, Nie H, Wang H, et al. CircEZH2/miR-133b/IGF2BP2 aggravates colorectal cancer progression via enhancing the stability of m(6)A-modified CREB1 mRNA. Mol Cancer. 2022;21:140.
Long F, Li L, Xie C, Ma M, Wu Z, Lu Z, et al. Intergenic CircRNA Circ_0007379 inhibits colorectal cancer progression by modulating miR-320a biogenesis in a KSRP-dependent manner. Int J Biol Sci. 2023;19:3781–803.
Du A, Li S, Zhou Y, Disoma C, Liao Y, Zhang Y, et al. M6A-mediated upregulation of circMDK promotes tumorigenesis and acts as a nanotherapeutic target in hepatocellular carcinoma. Mol Cancer. 2022;21:109.
Lee YS, Dutta A. MicroRNAs in cancer. Annu Rev Pathol. 2009;4:199–227.
Jay C, Nemunaitis J, Chen P, Fulgham P, Tong AW. miRNA profiling for diagnosis and prognosis of human cancer. DNA Cell Biol. 2007;26:293–300.
Zhang N, Hu X, Du Y, Du J. The role of miRNAs in colorectal cancer progression and chemoradiotherapy. Biomed Pharmacother. 2021;134:111099.
Chen Y, Fu LL, Wen X, Liu B, Huang J, Wang JH, et al. Oncogenic and tumor suppressive roles of microRNAs in apoptosis and autophagy. Apoptosis. 2014;19:1177–89.
Su MT, Kumata S, Endo S, Okada Y, Takai T. LILRB4 promotes tumor metastasis by regulating MDSCs and inhibiting miR-1 family miRNAs. Oncoimmunology. 2022;11:2060907.
Dunn KW, Kamocka MM, McDonald JH. A practical guide to evaluating colocalization in biological microscopy. Am J Physiol Cell Physiol. 2011;300:C723–42.
Wang N, Fang JY. Fusobacterium nucleatum, a key pathogenic factor and microbial biomarker for colorectal cancer. Trends Microbiol. 2023;31:159–72.
Yamane T, Kanamori Y, Sawayama H, Yano H, Nita A, Ohta Y, et al. Iron accelerates Fusobacterium nucleatum-induced CCL8 expression in macrophages and is associated with colorectal cancer progression. JCI Insight. 2022;7:e156802.
Meng S, Zhou H, Feng Z, Xu Z, Tang Y, Li P, et al. CircRNA: functions and properties of a novel potential biomarker for cancer. Mol Cancer. 2017;16:94.
Shen TD. Diet and gut microbiota in health and disease. Nestle Nutr Inst Workshop Ser. 2017;88:117–26.
Gentile CL, Weir TL. The gut microbiota at the intersection of diet and human health. Science. 2018;362:776–80.
Sun MF, Shen YQ. Dysbiosis of gut microbiota and microbial metabolites in Parkinson’s Disease. Ageing Res Rev. 2018;45:53–61.
Portincasa P, Bonfrate L, Vacca M, De Angelis M, Farella I, Lanza E, et al. Gut microbiota and short chain fatty acids: implications in glucose homeostasis. Int J Mol Sci. 2022;23:1105.
Xiao W, Su J, Gao X, Yang H, Weng R, Ni W, et al. The microbiota-gut-brain axis participates in chronic cerebral hypoperfusion by disrupting the metabolism of short-chain fatty acids. Microbiome. 2022;10:62.
Henao-Mejia J, Elinav E, Jin C, Hao L, Mehal WZ, Strowig T, et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature. 2012;482:179–85.
Miro-Blanch J, Yanes O. Epigenetic regulation at the interplay between gut microbiota and host metabolism. Front Genet. 2019;10:638.
Ansari I, Raddatz G, Gutekunst J, Ridnik M, Cohen D, Abu-Remaileh M, et al. The microbiota programs DNA methylation to control intestinal homeostasis and inflammation. Nat Microbiol. 2020;5:610–9.
Aleksandrova K, Romero-Mosquera B, Hernandez V. Gut microbiome and epigenetics: emerging links with inflammatory bowel diseases and prospects for management and prevention. Nutrients. 2017;9:962.
Gury-BenAri M, Thaiss CA, Serafini N, Winter DR, Giladi A, Lara-Astiaso D, et al. The spectrum and regulatory landscape of intestinal innate lymphoid cells are shaped by the microbiome. Cell. 2016;166:1231–46.e13.
Hong J, Guo F, Lu SY, Shen C, Ma D, Zhang X, et al. F. nucleatum targets lncRNA ENO1-IT1 to promote glycolysis and oncogenesis in colorectal cancer. Gut. 2021;70:2123–37.
Hui B, Zhou C, Xu Y, Wang R, Dong Y, Zhou Y, et al. Exosomes secreted by Fusobacterium nucleatum-infected colon cancer cells transmit resistance to oxaliplatin and 5-FU by delivering hsa_circ_0004085. J Nanobiotechnol. 2024;22:62.
Wang Z, Wei P, Wei D, Cao S, Liu H, Chen L, et al. Effect of up-regulation of circMATR3 on the proliferation, metastasis, progression and survival of hypopharyngeal carcinoma. J Cell Mol Med. 2020;24:4687–97.
Luo Z, Mao X, Cui W. Circular RNA expression and circPTPRM promotes proliferation and migration in hepatocellular carcinoma. Med Oncol. 2019;36:86.
Udayasuryan B, Zhou Z, Ahmad RN, Sobol P, Deng C, Nguyen TTD, et al. Fusobacterium nucleatum infection modulates the transcriptome and epigenome of HCT116 colorectal cancer cells in an oxygen-dependent manner. Commun Biol. 2024;7:551.
Qin J, Zhen S, Wang J, Lv W, Zhao Y, Duan Y, et al. Function of hsa_circ_0006646 as a competing endogenous RNA to promote progression in gastric cancer by regulating the miR-665-HMGB1 axis. J Gastrointest Oncol. 2023;14:1259–78.
Li Q, Geng S, Yuan H, Li Y, Zhang S, Pu L, et al. Circular RNA expression profiles in extracellular vesicles from the plasma of patients with pancreatic ductal adenocarcinoma. FEBS Open Bio. 2019;9:2052–62.
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495:384–8.
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–8.
Yan C, Zhang W, Shi X, Zheng J, Jin X, Huo J. MiR-760 suppresses non-small cell lung cancer proliferation and metastasis by targeting ROS1. Environ Sci Pollut Res Int. 2018;25:18385-91.
Li L, Yu P, Zhang P, Wu H, Chen Q, Li S, et al. Upregulation of hsa_circ_0007874 suppresses the progression of ovarian cancer by regulating the miR-760/SOCS3 pathway. Cancer Med. 2020;9:2491–9.
Ge L, Wang Y, Duan QH, Liu SS, Liu GJ. MicroRNA-760 acts as a tumor suppressor in gastric cancer development via inhibiting G-protein-coupled receptor kinase interacting protein-1 transcription. World J Gastroenterol. 2019;25:6619–33.
Cao L, Liu Y, Wang D, Huang L, Li F, Liu J, et al. MiR-760 suppresses human colorectal cancer growth by targeting BATF3/AP-1/cyclinD1 signaling. J Exp Clin Cancer Res. 2018;37:83.
Huang S, Long Y, Gao Y, Lin W, Wang L, Jiang J, et al. Combined inhibition of MET and VEGF enhances therapeutic efficacy of EGFR TKIs in EGFR-mutant non-small cell lung cancer with concomitant aberrant MET activation. Exp Hematol Oncol. 2024;13:97.
Dong L, Jiang CC, Thorne RF, Croft A, Yang F, Liu H, et al. Ets-1 mediates upregulation of Mcl-1 downstream of XBP-1 in human melanoma cells upon ER stress. Oncogene. 2011;30:3716–26.
Kim GC, Kwon HK, Lee CG, Verma R, Rudra D, Kim T, et al. Upregulation of ETS1 expression by NFATc2 and NFKB1/RELA promotes breast cancer cell invasiveness. Oncogenesis. 2018;7:91.
Kang W, Jia Z, Tang D, Zhang Z, Gao H, He K, et al. Fusobacterium nucleatum facilitates apoptosis, ROS generation, and inflammatory cytokine production by activating AKT/MAPK and NF-κB signaling pathways in human gingival fibroblasts. Oxid Med Cell Longev. 2019;2019:1681972.
Span PN, Manders P, Heuvel JJ, Thomas CM, Bosch RR, Beex LV, et al. Expression of the transcription factor Ets-1 is an independent prognostic marker for relapse-free survival in breast cancer. Oncogene. 2002;21:8506–9.
Yu Y, Zhang YC, Zhang WZ, Shen LS, Hertzog P, Wilson TJ, et al. Ets1 as a marker of malignant potential in gastric carcinoma. World J Gastroenterol. 2003;9:2154–9.
Zhou X, Zhou R, Zhou H, Li Q, Hong J, Meng R, et al. ETS-1 induces endothelial-like differentiation and promotes metastasis in non-small cell lung cancer. Cell Physiol Biochem. 2018;45:1827–39.
Nakayama T, Ito M, Ohtsuru A, Naito S, Sekine I. Expression of the ets-1 proto-oncogene in human colorectal carcinoma. Mod Pathol. 2001;14:415–22.
Oikawa M, Abe M, Kurosawa H, Hida W, Shirato K, Sato Y. Hypoxia induces transcription factor ETS-1 via the activity of hypoxia-inducible factor-1. Biochem Biophys Res Commun. 2001;289:39–43.
Wang S, Wan L, Zhang X, Fang H, Zhang M, Li F, et al. ETS-1 in tumor immunology: implications for novel anti-cancer strategies. Front Immunol. 2025;16:1526368.
Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12:381–8.
Santer L, Bär C, Thum T. Circular RNAs: a novel class of functional RNA molecules with a therapeutic perspective. Mol Ther. 2019;27:1350–63.
Li X, Yang L, Chen LL. The biogenesis, functions, and challenges of circular RNAs. Mol Cell. 2018;71:428–42.
Liu X, Zhang Y, Zhou S, Dain L, Mei L, Zhu G. Circular RNA: an emerging frontier in RNA therapeutic targets, RNA therapeutics, and mRNA vaccines. J Control Rel. 2022;348:84–94.
Acknowledgements
We thank the Department of Gastroenterology, Shanghai General Hospital and Shanghai Institute of Digestive Disease for providing support on research conditions in this study.
Funding
This work was financially supported by grants from the National Natural Science Foundation of China (Nos.82203500, Nos.82303215).
Author information
Authors and Affiliations
Contributions
Conception and design: CML, YXC. Development of methodology: CML, QQL, LNF. Acquisition of data (provided animals, acquired and managed patients, provided facilities, etc.): CML, QQL, THZ. Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): CML, QQL. Writing, review, and/or revision of the manuscript: QQL, HCS, CML, LNF. Administrative, technical, or material support (i.e., reporting or organizing data, constructing databases): CML, YXC. Study supervision: LNF, THZ, YXC.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, C., Liu, Q., Shen, H. et al. Fusobacterium nucleatum drives colorectal cancer progression through the circPTBP3/miR-760/PUM1 axis. Oncogene (2026). https://doi.org/10.1038/s41388-026-03746-4
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41388-026-03746-4