Abstract
Introduction
Early-life exposure to antibiotics (ABX) has been linked to increases in asthma severity and prevalence in both children and laboratory animals. We explored the immunologic mechanisms behind this association using a mouse model of house dust mite (HDM)-induced asthma and early-life ABX exposure.
Methods
Mice were exposed to three short courses of ABX following weaning and experimental asthma was thereafter induced. Airway cell counts and differentials; serum immunoglobulin E (IgE); pulmonary function; lung histopathology; pulmonary regulatory T cells (Tregs); and the fecal microbiome were characterized following ABX exposure and induction of experimental asthma.
Results
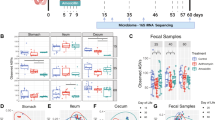
Asthma severity was increased in mice exposed to ABX, including: airway eosinophilia, airway hyper-reactivity, serum HDM-specific IgE, and lung histopathology. ABX treatment led to sharp reduction in fecal microbiome diversity, including the loss of pro-regulatory organisms such as Lachnospira. Pulmonary Tregs were reduced with ABX treatment, and this reduction was directly proportional to diminished microbiome diversity.
Conclusion
Intermittent exposure to ABX early in life worsened the severity of experimental asthma and reduced pulmonary Tregs; the latter change correlated with decreased microbiome diversity. These data may suggest targets for immunologic or probiotic therapy to counteract the harmful effects of childhood ABX.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Fanta, C. H. Asthma. N. Engl. J. Med. 360, 1002–1014 (2009).
Lötvall, J. et al. Asthma endotypes: a new approach to classification of disease entities within the asthma syndrome. J. Allergy Clin. Immunol. 127, 355–360 (2011).
Adami, A. J. & Bracken, S. J. Breathing better through bugs: asthma and the microbiome. Yale J. Biol. Med. 89, 309–324 (2016).
Sears, M. R. Trends in the prevalence of asthma. Chest 145, 219–225 (2014).
Eder, W., Ege, M. J. & von Mutius, E. The asthma epidemic. N. Engl. J. Med. 355, 2226–2235 (2006).
National Center for Health Statistics. Health, United States, 2014: With Special Feature on Adults Aged 55–64 [Internet]. (Centers for Disease Control, Hyattsville, 2015). http://www.cdc.gov/nchs/hus.htm
Yamamoto-Hanada, K., Yang, L., Narita, M., Saito, H. & Ohya, Y. Influence of antibiotic use in early childhood on asthma and allergic diseases at age 5. Ann. Allergy Asthma Immunol. 119, 54–58 (2017).
Walker, W. A. The importance of appropriate initial bacterial colonization of the intestine in newborn, child, and adult health. Pediatr. Res. 82, 387–395 (2017).
Noverr, M. C., Noggle, R. M., Toews, G. B. & Huffnagle, G. B. Role of antibiotics and fungal microbiota in driving pulmonary allergic responses. Infect. Immun. 72, 4996–5003 (2004).
Russell, S. L. et al. Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma. EMBO Rep. 13, 440–447 (2012).
Hill, D. A. et al. Commensal bacteria-derived signals regulate basophil hematopoiesis and allergic inflammation. Nat. Med. 18, 538–546 (2012).
Russell, S. L. et al. Perinatal antibiotic treatment affects murine microbiota, immune responses and allergic asthma. Gut Microbes 4, 158–164 (2013).
Zaiss, M. M. et al. The intestinal microbiota contributes to the ability of helminths to modulate allergic inflammation. Immunity 43, 998–1010 (2015).
Little, P. et al. Pragmatic randomised controlled trial of two prescribing strategies for childhood acute otitis media. BMJ 322, 336–342 (2001).
Bracken, S. J. et al. Long-term exposure to house dust mite leads to the suppression of allergic airway disease despite persistent lung inflammation. Int Arch. Allergy Immunol. 166, 243–258 (2015).
Nobel, Y. R. et al. Metabolic and metagenomic outcomes from early-life pulsed antibiotic treatment. Nat. Commun. 6, 7486 (2015).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol 79, 5112–5120 (2013).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Nelson, M. C., Morrison, H. G., Benjamino, J., Grim, S. L. & Graf, J. Analysis, optimization and verification of Illumina-generated 16S rRNA gene amplicon surveys. PLoS ONE 9, e94249 (2014).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 6, 610–618 (2012).
Simpson, E. H. Measurement of diversity. Nature 163, 688 (1949).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Carson, W. F. IV et al. Accumulation of regulatory T cells in local draining lymph nodes of the lung correlates with spontaneous resolution of chronic asthma in a murine model. Int Arch. Allergy Immunol. 145, 231–243 (2008).
Taylor, A., Verhagen, J., Akdis, C. A. & Akdis, M. T regulatory cells in allergy and health: a question of allergen specificity and balance. Int Arch. Allergy Immunol. 135, 73–82 (2004).
Robinson, D. S. Regulatory T cells and asthma. Clin. Exp. Allergy 39, 1314–1323 (2009).
Natarajan, P. et al. Regulatory B cells from hilar lymph nodes of tolerant mice in a murine model of allergic airway disease are CD5+, express TGF-β, and co-localize with CD4+Foxp3+T cells. Mucosal Immunol. 5, 691–701 (2012).
Abrahamsson, T. R. et al. Low gut microbiota diversity in early infancy precedes asthma at school age. Clin. Exp. Allergy 44, 842–850 (2014).
Dannemiller, K. C. et al. Next-generation DNA sequencing reveals that low fungal diversity in house dust is associated with childhood asthma development. Indoor Air 24, 236–247 (2014).
Arrieta, M.-C. et al. Early infancy microbial and metabolic alterations affect risk of childhood asthma. Sci. Transl. Med. 7, 307ra152 (2015).
Ball, T. M. et al. Siblings, day-care attendance, and the risk of asthma and wheezing during childhood. N. Engl. J. Med. 343, 538–543 (2000).
Thompson, A. L., Monteagudo-Mera, A., Cadenas, M. B., Lampl, M. L. & Azcarate-Peril, M. A. Milk- and solid-feeding practices and daycare attendance are associated with differences in bacterial diversity, predominant communities, and metabolic and immune function of the infant gut microbiome. Front. Cell. Infect. Microbiol. 5, 3 (2015).
Atarashi, K. et al. Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500, 232–236 (2013).
Koenig, J. E. et al. Succession of microbial consortia in the developing infant gut microbiome. Proc. Natl. Acad. Sci. USA 108(Suppl. 1), 4578–4585 (2011).
Elazab, N. et al. Probiotic administration in early life, atopy, and asthma: a meta-analysis of clinical trials. Pediatrics 132, e666–e676 (2013).
Blaser, M. J. Antibiotic use and its consequences for the normal microbiome. Science 352, 544–545 (2016).
Russell, S. L. et al. Perinatal antibiotic-induced shifts in gut microbiota have differential effects on inflammatory lung diseases. J. Allergy Clin. Immunol. 135, 100–109 (2015).
Labro, M. T. & Abdelghaffar, H. Immunomodulation by macrolide antibiotics. J. Chemother. (Florence, Italy) 13, 3–8 (2001).
Acknowledgements
We thank Jacqui Benjamino, Susan Janton, and Michael Nelson of the Graf Lab (UConn Storrs) for feedback and assistance. We also thank David Benson (UConn Storrs) for use of his laboratory space for experimentation. This work was supported by National Institutes of Health grants R01-AI43573 (R.S.T. and C.M.S.), F30-HL122018 (S.J.B.), and F30 HL-126324 (A.J.A.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Category of Study: Translational
Rights and permissions
About this article
Cite this article
Adami, A.J., Bracken, S.J., Guernsey, L.A. et al. Early-life antibiotics attenuate regulatory T cell generation and increase the severity of murine house dust mite-induced asthma. Pediatr Res 84, 426–434 (2018). https://doi.org/10.1038/s41390-018-0031-y
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41390-018-0031-y
This article is cited by
-
Microbiome in heritage: how maternal microbiome transmission impacts next generation health
Microbiome (2025)
-
Maternal emulsifier consumption alters the offspring early-life microbiota and goblet cell function leading to long-lasting diseases susceptibility
Nature Communications (2025)
-
Neonatal and early infancy antibiotic exposure is associated with childhood atopic dermatitis, wheeze and asthma
European Journal of Pediatrics (2024)
-
Influence of the early-life gut microbiota on the immune responses to an inhaled allergen
Mucosal Immunology (2022)
-
Dysbiosis of the gut and lung microbiome has a role in asthma
Seminars in Immunopathology (2020)