Abstract
Background
Special AT-rich sequence-binding protein 2 is essential for the development of cerebral cortex and key molecular node for the establishment of proper neural circuitry and function. Mutations in the SATB2 gene lead to SATB2-associated syndrome, which is characterized by abnormal development of skeleton and central nervous systems.
Methods
We generated Satb2 knockout mouse model through CRISPR-Cas9 technology and performed RNA-seq and ChIP-seq of embryonic cerebral cortex. We conducted RT-qPCR, western blot, immunofluorescence staining, luciferase reporter assay and behavioral analysis for experimental verification.
Results
We identified 1363 downstream effector genes of Satb2 and correlation analysis of Satb2-targeted genes and neurological disease genes showed that Satb2 contribute to cognitive and mental disorders from the early developmental stage. We found that Satb2 directly regulate the expression of Ntng1, Cdh13, Kitl, genes important for axon guidance, synaptic formation, neuron migration, and Satb2 directly activates the expression of Mef2c. We also showed that Satb2 heterozygous knockout mice showed impaired spatial learning and memory.
Conclusions
Taken together, our study supportsroles of Satb2 in the regulation of axonogenesis and synaptic formation at the early developmental stage and provides new insights into the complicated regulatory mechanism of Satb2 and new evidence to elucidate the pathogen of SATB2-associated syndrome.
Impact
-
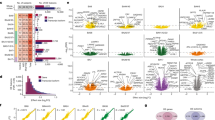
1363 downstream effector genes of Satb2 were classified into 5 clusters with different temporal expression patterns.
-
We identified Plxnd1, Ntng1, Efnb2, Ephb1, Plxna2, Epha3, Plxna4, Unc5c, and Flrt2 as axon guidance molecules to regulate axonogenesis.
-
168 targeted genes of Satb2 were found to regulate synaptic formation in the early development of the cerebral cortex.
-
Transcription factor Mef2c is positively regulated by Satb2, and 28 Mef2c-targeted genes can be directly regulated by Satb2.
-
In the Morris water maze test, Satb2+/− mice showed impaired spatial learning and memory, further strengthening that Satb2 can regulate synaptic functions.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
Data availability
RNAseq and ChIP-seq data that support the findings of this study have been deposited to GEO database with the accession number GSE201562.
References
Britanova, O., Akopov, S., Lukyanov, S., Gruss, P. & Tarabykin, V. Novel transcription factor Satb2 interacts with matrix attachment region DNA elements in a tissue-specific manner and demonstrates cell-type-dependent expression in the developing mouse CNS. Eur. J. Neurosci. 21, 658–668 (2005).
Dobreva, G. et al. SATB2 is a multifunctional determinant of craniofacial patterning and osteoblast differentiation. Cell 125, 971–986 (2006).
Alcamo, E. A. et al. Satb2 regulates callosal projection neuron identity in the developing cerebral cortex. Neuron 57, 364–377 (2008).
Britanova, O. et al. Satb2 is a postmitotic determinant for upper-layer neuron specification in the neocortex. Neuron 57, 378–392 (2008).
Li Y, et al. Satb2 ablation impairs hippocampus-based long-term spatial memory and short-term working memory and immediate early genes (IEGs)-mediated hippocampal synaptic plasticity. Mol. Neurobiol. https://doi.org/10.1007/s12035-017-0531-5 (2017).
Zarate, Y. A. & Fish, J. L. SATB2-associated syndrome: mechanisms, phenotype, and practical recommendations. Am. J. Med Genet. A 173, 327–337 (2017).
Lewis, H. et al. Epilepsy and electroencephalographic abnormalities in SATB2-associated syndrome. Pediatr. Neurol. 112, 94–100 (2020).
Leone, D. P. et al. Satb2 regulates the differentiation of both callosal and subcerebral projection neurons in the developing cerebral cortex. Cereb. Cortex 25, 3406–3419 (2015).
Hall, A. & Lalli, G. Rho and Ras GTPases in axon growth, guidance, and branching. Cold Spring Harb. Perspect. Biol. 2, a001818 (2010).
Tessier-Lavigne, M. & Goodman, C. S. The molecular biology of axon guidance. Science 274, 1123–1133 (1996).
Kamiguchi, H. The mechanism of axon growth: what we have learned from the cell adhesion molecule L1. Mol. Neurobiol. 28, 219–228 (2003).
Baranek, C. et al. Protooncogene Ski cooperates with the chromatin-remodeling factor Satb2 in specifying callosal neurons. Proc. Natl Acad. Sci. USA 109, 3546–3551 (2012).
Srivatsa, S. et al. Unc5C and DCC act downstream of Ctip2 and Satb2 and contribute to corpus callosum formation. Nat. Commun. 5, 3708 (2014).
He, C. -H. et al. Satb2 regulates EphA7 to control soma spacing and self-avoidance of cortical pyramidal neurons. Cereb. Cortex 32, 2321–2331 (2021).
Defelipe, J. The evolution of the brain, the human nature of cortical circuits, and intellectual creativity. Front. Neuroanat. 5, 29 (2011).
Takeuchi, T., Duszkiewicz, A. J. & Morris, R. G. M. The synaptic plasticity and memory hypothesis: encoding, storage and persistence. Philos. Trans. R. Soc. Lond. B Biol. Sci. 369, 20130288 (2014).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
McKenna, W. L. et al. Mutual regulation between Satb2 and Fezf2 promotes subcerebral projection neuron identity in the developing cerebral cortex. Proc. Natl Acad. Sci. USA 112, 11702–11707 (2015).
Cera, I. et al. Genes encoding SATB2-interacting proteins in adult cerebral cortex contribute to human cognitive ability. PLoS Genet. 15, e1007890 (2019).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Yu, G., Wang, L.-G. & He, Q.-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation (N. Y) 2, 100141 (2021).
Luo, W. & Brouwer, C. Pathview: an R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics 29, 1830–1831 (2013).
Hu, H. et al. AnimalTFDB 3.0: a comprehensive resource for annotation and prediction of animal transcription factors. Nucleic Acids Res. 47, D33–D38 (2019).
Weyn-Vanhentenryck, S. M. et al. Precise temporal regulation of alternative splicing during neural development. Nat. Commun. 9, 2189 (2018).
Kumar, L. & E Futschik, M. Mfuzz: a software package for soft clustering of microarray data. Bioinformation 2, 5–7 (2007).
Jin, K., Mao, X. O., Sun, Y., Xie, L. & Greenberg, D. A. Stem cell factor stimulates neurogenesis in vitro and in vivo. J. Clin. Invest. 110, 311–319 (2002).
Erlandsson, A., Larsson, J. & Forsberg-Nilsson, K. Stem cell factor is a chemoattractant and a survival factor for CNS stem cells. Exp. Cell Res. 301, 201–210 (2004).
Leoyklang, P. et al. Heterozygous nonsense mutation SATB2 associated with cleft palate, osteoporosis, and cognitive defects. Hum. Mutat. 28, 732–738 (2007).
Yin, Y., Miner, J. H. & Sanes, J. R. Laminets: laminin- and netrin-related genes expressed in distinct neuronal subsets. Mol. Cell Neurosci. 19, 344–358 (2002).
Pirooznia, M. et al. SynaptomeDB: an ontology-based knowledgebase for synaptic genes. Bioinformatics 28, 897–899 (2012).
Südhof, T. C. Towards an understanding of synapse formation. Neuron 100, 276–293 (2018).
Philippova, M., Ivanov, D., Tkachuk, V., Erne, P. & Resink, T. J. Polarisation of T-cadherin to the leading edge of migrating vascular cells in vitro: a function in vascular cell motility? Histochem. Cell Biol. 120, 353–360 (2003).
Paradis, S. et al. An RNAi-based approach identifies molecules required for glutamatergic and GABAergic synapse development. Neuron 53, 217–232 (2007).
Rivero, O. et al. Cadherin-13, a risk gene for ADHD and comorbid disorders, impacts GABAergic function in hippocampus and cognition. Transl. Psychiatry 5, e655 (2015).
Drgonova, J. et al. Cadherin 13: human cis-regulation and selectively-altered addiction phenotypes and cerebral cortical dopamine in knockout mice. Mol. Med. 22, 537–547 (2016).
Harrington, A. J. et al. MEF2C regulates cortical inhibitory and excitatory synapses and behaviors relevant to neurodevelopmental disorders. Elife 5, e20059 (2016).
Allaway, K. C. et al. Genetic and epigenetic coordination of cortical interneuron development. Nature 597, 693–697 (2021).
Kuwako, K.-I., Nishimoto, Y., Kawase, S., Okano, H. J. & Okano, H. Cadherin-7 regulates mossy fiber connectivity in the cerebellum. Cell Rep. 9, 311–323 (2014).
Föcking, M. et al. Proteomic analysis of the postsynaptic density implicates synaptic function and energy pathways in bipolar disorder. Transl. Psychiatry 6, e959 (2016).
Beeson, K. A., Beeson, R., Westbrook, G. L. & Schnell, E. α2δ-2 protein controls structure and function at the cerebellar climbing fiber synapse. J. Neurosci. 40, 2403–2415 (2020).
Di Gregorio, S. E., Volkening, K., Strong, M. J. & Duennwald, M. L. Inclusion formation and toxicity of the ALS protein RGNEF and its association with the microtubule network. Int. J. Mol. Sci. 21, E5597 (2020).
Reshetnikov, V. V. et al. Genes associated with cognitive performance in the Morris water maze: an RNA-seq study. Sci. Rep. 10, 22078 (2020).
Russell, S. A. & Bashaw, G. J. Axon guidance pathways and the control of gene expression. Dev. Dyn. 247, 571–580 (2018).
Sakurai, A. et al. Semaphorin 3E initiates antiangiogenic signaling through plexin D1 by regulating Arf6 and R-Ras. Mol. Cell Biol. 30, 3086–3098 (2010).
Hatanaka, Y. et al. Semaphorin 6A-plexin A2/A4 interactions with radial glia regulate migration termination of superficial layer cortical neurons. iScience 21, 359–374 (2019).
Limoni, G., Murthy, S., Jabaudon, D., Dayer, A. & Niquille, M. PlexinA4-Semaphorin3A-mediated crosstalk between main cortical interneuron classes is required for superficial interneuron lamination. Cell Rep. 34, 108644 (2021).
Altuame, F. D. et al. PLXNA2 as a candidate gene in patients with intellectual disability. Am. J. Med. Genet. A 185, 3859–3865 (2021).
Pijuan, J. et al. PLXNA2 and LRRC40 as candidate genes in autism spectrum disorder. Autism Res. 14, 1088–1100 (2021).
Noren, N. K. & Pasquale, E. B. Eph receptor-ephrin bidirectional signals that target Ras and Rho proteins. Cell Signal 16, 655–666 (2004).
Javier-Torrent, M. et al. Presenilin/γ-secretase-dependent EphA3 processing mediates axon elongation through non-muscle myosin IIA. Elife 8, e43646 (2019).
Salatino-Oliveira, A. et al. Catechol-O-methyltransferase valine158methionine polymorphism moderates methylphenidate effects on oppositional symptoms in boys with attention-deficit/hyperactivity disorder. Biol. Psychiatry 70, 216–221 (2011).
Salatino-Oliveira, A. et al. Cadherin-13 gene is associated with hyperactive/impulsive symptoms in attention/deficit hyperactivity disorder. Am. J. Med. Genet. B Neuropsychiatr. Genet. 168B, 162–169 (2015).
Rajkovich, K. E. et al. Experience-dependent and differential regulation of local and long-range excitatory neocortical circuits by postsynaptic Mef2c. Neuron 93, 48–56 (2017).
Assali, A., Harrington, A. J. & Cowan, C. W. Emerging roles for MEF2 in brain development and mental disorders. Curr. Opin. Neurobiol. 59, 49–58 (2019).
Zhang, Q., Huang, Y., Zhang, L., Ding, Y.-Q. & Song, N.-N. Loss of Satb2 in the cortex and hippocampus leads to abnormal behaviors in mice. Front. Mol. Neurosci. 12, 33 (2019).
Acknowledgements
We are very grateful to Dr. Zilong Qiu and Dr. Man Xiong for their comments and suggestions during the study. We are also grateful to the animal co-facility center of Children’s Hospital of Fudan University and our genetic laboratory teams who contributed to this study.
Funding
This study is funded by the National Natural Science Foundation of China (81471483, 81974237).
Author information
Authors and Affiliations
Contributions
H.W. designed the study and revised the manuscript; Q.G. performed experiment, collected and interpreted the data, drafted the initial manuscript; Y.W. analyzed the RNA-seq and ChIP-seq data; Y.Q., X.C. and X.L. involved experiment, X.D. and H.C. involved data analysis, Q.W., Y.J., S.Y., J.Z., and S.S. involved animal experiment; B.W. and W.Z. revised the manuscript; and all authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guo, Q., Wang, Y., Wang, Q. et al. In the developing cerebral cortex: axonogenesis, synapse formation, and synaptic plasticity are regulated by SATB2 target genes. Pediatr Res 93, 1519–1527 (2023). https://doi.org/10.1038/s41390-022-02260-z
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41390-022-02260-z
This article is cited by
-
Advances in research on SATB2 and its role in tumor development
Cell & Bioscience (2025)
-
Sex differences in brain protein expression and disease
Nature Medicine (2023)