Abstract
Introduction
Pharmacogenomics (PGx) testing aims to identify the most appropriate drug and dose for individual patients based on their genetic profiles. In Jordan, patients with genetic disorders often use multiple medications, some of which have clinical guidelines recommending PGx testing.
Aim
This study aimed to screen the frequency of clinically relevant PGx biomarkers among a sample of Jordanian patients with genetic disorders.
Methods
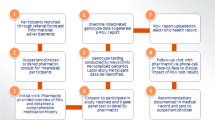
A total of 76 patients (average age 13 ± 14 years; 71% under 13 years old) attending INNOVIA Biobank in Amman between January 2023 and January 2024 participated. Buccal swabs were collected, and DNA was extracted for whole-genome sequencing using Illumina technology. Variant calling and annotation were performed using DRAGEN, Geneyx, and ANNOVAR tools. A PGx panel based on PharmCAT v2.8.3 and Clinical Pharmacogenetics Implementation Consortium v1.30.0, covering 20 pharmacogenes, was applied.
Result
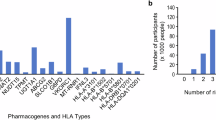
In Phase I enzymes, CYP2D6*10 (13.2%) and CYP2C19*1/*17 (18.4%) were most common, while CYP2C9 and CYP3A4 variants were less frequent. In Phase II enzymes, UGT1A180 + 2B appeared in 7.9% and multiple DPYD variants found in heterozygous forms 925%). Among toxicity-related markers, G6PD and HLA-B*57:01 were detected in 3.9 and 2.6%, respectively. Transporter gene variants in SLCO1B1 (15%) and ABCB1 (21.1%) were relatively frequent. For pharmacodynamic genes, VKORC1 −1639G > A (52.6%) and CYP4F2 V433M (40.8%) were most prevalent. Accordingly, over half of the patients had genetic variants affecting warfarin response, with additional impacts seen on antidepressants (45%), clopidogrel (35%), and anticancers (30%). Conclusions, this study demonstrates the presence of key PGx biomarkers among Jordanian patients with genetic diseases and supports the integration of PGx testing to optimize the use of drugs like antidepressants, clopidogrel, and warfarin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Data availability
Data are available with the corresponding author upon request. Conflicts of Interest: The authors declare no conflicts of interest.
References
Roden DM, Wilke RA, Kroemer HK, Stein CM. Pharmacogenomics: the genetics of variable drug responses. Circulation. 2011;123:1661–70.
Pinto N, Dolan ME. Clinically relevant genetic variations in drug metabolizing enzymes. Curr Drug Metab. 2011;12:487–97.
Whirl-Carrillo M, Huddart R, Gong L, Sangkuhl K, Thorn CF, Whaley R, et al. An evidence-based framework for evaluating pharmacogenomics knowledge for personalized medicine. Clin Pharmacol Ther. 2021;110:563–72.
Ortega VE, Meyers DA. Pharmacogenetics: implications of race and ethnicity on defining genetic profiles for personalized medicine. J Allergy Clin Immunol. 2014;133:16–26.
Jarrar YB, Balasmeh AA, Jarrar W. Sequence analysis of the N-acetyltransferase 2 gene (NAT2) among Jordanian volunteers. Libyan J Med. 2018;13:1408381.
Johnson JA, Caudle KE, Gong L, Whirl-Carrillo M, Stein CM, Scott SA, et al. Clinical pharmacogenetics implementation consortium (CPIC) guideline for pharmacogenetics-guided warfarin dosing: 2017 update. Clin Pharmacol Ther. 2017;102:397–404.
Al-Mahayri ZN, Patrinos GP, Wattanapokayakit S, Iemwimangsa N, Fukunaga K, Mushiroda T, et al. Variation in 100 relevant pharmacogenes among emiratis with insights from understudied populations. Sci Rep. 2020;10:21310.
Abdel-Latif R, Badji R, Mohammed S, Al-Muftah W, Mbarek H, Darwish D, et al. QPGx-CARES: qatar pharmacogenetics clinical applications and research enhancement strategies. Clin Transl Sci. 2024;17:e13800.
Goljan E, Abouelhoda M, ElKalioby MM, Jabaan A, Alghithi N, Meyer BF, et al. Identification of pharmacogenetic variants from large scale next generation sequencing data in the Saudi population. PLoS One. 2022;17:e0263137.
Alhawari H, Jarrar Y, AlKhatib MA, Alhawari H, Momani M, Zayed A, et al. The Association of 3-hydroxy-3-methylglutaryl-CoA reductase, apolipoprotein e, and solute carrier organic anion genetic variants with atorvastatin response among jordanian patients with type 2 diabetes. Life (Basel). 2020;10:232.
Hakooz N, Jarrar YB, Zihlif M, Imraish A, Hamed S, Arafat T. Effects of the genetic variants of organic cation transporters 1 and 3 on the pharmacokinetics of metformin in Jordanians. Drug Metab Pers Ther. 2017;32:157–62.
Alhawari H, Jarrar Y, Zihlif M, Wahbeh A, Alshelleh S, Ojjoh K, et al. Variability in response to valsartan and its relationship with AGT M235T genotype and other nongenetic parameters among a sample of hypertensive individuals in jordan: a prospective pilot study. Health Sci Rep. 2025;8:e70611.
Hamamy H, Al-Hait S, Alwan A, Ajlouni K. Jordan: communities and community genetics. Community Genet. 2007;10:52–60.
El-Shanti H. The impact of genetic diseases on jordanians: strategies towards prevention. J Biomed Biotechnol. 2001;1:45–47.
Ghanim M, Mosleh R, Hamdan A, Amer J, Alqub M, Jarrar Y, et al. Assessment of perceptions and predictors towards consanguinity: a cross-sectional study from palestine. J Multidiscip Healthc. 2023;16:3443–53.
Hulley SB, Cummings SR, Browner WS, Grady DG, Newman TB. Designing Clinical Research. 4th ed. Philadelphia: Lippincott Williams & Wilkins; 2013.
Schneider VA, Graves-Lindsay T, Howe K, Bouk N, Chen HC, Kitts PA, et al. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res. 2017;27:849–64.
den Dunnen JT, Dalgleish R, Maglott DR, Hart RK, Greenblatt MS, McGowan-Jordan J, et al. HGVS recommendations for the description of sequence variants: 2016 update. Hum Mutat. 2016;37:564–9.
Clinical Pharmacogenetics Implementation Consortium (CPIC). CPIC website. Stanford (CA): CPIC. Accessed date: 14 January 2025. https://cpicpgx.org/.
Abed E, Jarrar Y, Alhawari H, Abdullah S, Zihlif M. How the cytochrome 7a1 (CYP7A1) and ATP-binding cassette G8 (ABCG8) genetic variants affect atorvastatin response among type 2 diabetic patients attending the university of jordan hospital. Int J Clin Pharmacol Ther. 2021;59:99–108.
Abdullah S, Jarrar Y, Alhawari H, Abed E, Zihlif M. The Influence of endothelial nitric oxide synthase (eNOS) genetic polymorphisms on cholesterol blood levels among type 2 diabetic patients on atorvastatin therapy. Endocr Metab Immune Disord Drug Targets. 2021;21:352–9.
Ryan SL, Peden JF, Kingsbury Z, Schwab CJ, James T, Polonen P, et al. Whole genome sequencing provides comprehensive genetic testing in childhood B-cell acute lymphoblastic leukaemia. Leukemia. 2023;37:518–28.
Austin-Zimmerman I, Wronska M, Wang B, Irizar H, Thygesen JH, Bhat A, et al. The influence of CYP2D6 and CYP2C19 genetic variation on diabetes mellitus risk in people taking antidepressants and antipsychotics. Genes (Basel). 2021;12:1758.
Gong L, Whirl-Carrillo M, Klein TE. PharmGKB, an integrated resource of pharmacogenomic knowledge. Curr Protoc. 2021;1:e226.
Bousman CA, Stevenson JM, Ramsey LB, Sangkuhl K, Hicks JK, Strawn JR, et al. Clinical pharmacogenetics implementation consortium (CPIC) guideline for CYP2D6, CYP2C19, CYP2B6, SLC6A4, and HTR2A genotypes and serotonin reuptake inhibitor antidepressants. Clin Pharmacol Ther. 2023;114:51–68.
Bertilsson L, Dahl ML, Dalén P, Al-Shurbaji A. Molecular genetics of CYP2D6: clinical relevance with focus on psychotropic drugs. Br J Clin Pharmacol. 2002;53:111–22.
Cuisset T, Loosveld M, Morange PE, Quilici J, Moro PJ, Saut N, et al. CYP2C19*2 and *17 alleles have a significant impact on platelet response and bleeding risk in patients treated with prasugrel after acute coronary syndrome. JACC Cardiovasc Interv. 2012;5:1280–7.
Langmia IM, Just KS, Yamoune S, Brockmöller J, Masimirembwa C, Stingl JC. CYP2B6 functional variability in drug metabolism and exposure across populations-implication for drug safety, dosing, and individualized therapy. Front Genet. 2021;12:692234.
Zanger UM, Klein K. Pharmacogenetics of cytochrome P450 2B6 (CYP2B6): advances on polymorphisms, mechanisms, and clinical relevance. Front Genet. 2013;4:24.
Li Y, Jackson KA, Slon B, Hardy JR, Franco M, William L, et al. CYP2B6*6 allele and age substantially reduce steady-state ketamine clearance in chronic pain patients: impact on adverse effects. Br J Clin Pharmacol. 2015;80:276–84.
Alrabayah M, Massad IM, Imraish A, Jarrar Y, Massad F, Guzu H, et al. Effects of CYP2B6 genetic variants on the propofol dose and response among jordanian arabic patients undergoing general anesthesia. Curr Drug Metab. 2023;23:1156–61.
Marcuello E, Altés A, Menoyo A, Del Rio E, Gómez-Pardo M, Baiget M. UGT1A1 gene variations and irinotecan treatment in patients with metastatic colorectal cancer. Br J Cancer. 2004;91:678–82.
Hulshof EC, Deenen MJ, Nijenhuis M, Soree B, de Boer-Veger NJ, Buunk AM, et al. Dutch pharmacogenetics working group (DPWG) guideline for the gene-drug interaction between UGT1A1 and irinotecan. Eur J Hum Genet. 2023;31:982–7.
de Moraes FCA, de Almeida Barbosa AB, Sano VKT, Kelly FA, Burbano RMR. Pharmacogenetics of DPYD and treatment-related mortality on fluoropyrimidine chemotherapy for cancer patients: a meta-analysis and trial sequential analysis. BMC Cancer. 2024;24:1210.
Al-Khateeb M, Awidi A, Al-Hadidi K, Battah A. Low incidence of the DPD IVS14 + 1G>a polymorphism in jordanian breast and colorectal cancer patients. Asian Pac J Cancer Prev. 2017;18:1651–4.
Amstutz U, Henricks LM, Offer SM, Barbarino J, Schellens JHM, Swen JJ, et al. Clinical pharmacogenetics implementation consortium (CPIC) guideline for dihydropyrimidine dehydrogenase genotype and fluoropyrimidine dosing: 2017 update. Clin Pharmacol Ther. 2018;103:210–6.
Awab GR, Aaram F, Jamornthanyawat N, Suwannasin K, Pagornrat W, Watson JA, et al. Protective effect of mediterranean-type glucose-6-phosphate dehydrogenase deficiency against Plasmodium vivax malaria. Elife. 2021;10:e62448.
Martin MA, Hoffman JM, Freimuth RR, Klein TE, Dong BJ, Pirmohamed M, et al. Clinical pharmacogenetics implementation consortium. clinical pharmacogenetics implementation consortium guidelines for HLA-B genotype and abacavir dosing: 2014 update. Clin Pharmacol Ther. 2014;95:499–500.
Cooper-DeHoff RM, Niemi M, Ramsey LB, Luzum JA, Tarkiainen EK, Straka RJ, et al. The clinical pharmacogenetics implementation consortium guideline for SLCO1B1, ABCG2, and CYP2C9 genotypes and statin-associated musculoskeletal symptoms. Clin Pharmacol Ther. 2022;111:1007–21.
McDonald MG, Rieder MJ, Nakano M, Hsia CK, Rettie AE. CYP4F2 is a vitamin K1 oxidase: an explanation for altered warfarin dose in carriers of the V433M variant. Mol Pharmacol. 2009;75:1337–46.
Jarrar Y, Alkhalili M, Alhawari H, Abaalkhail SJ, Alkhalili S, Alhawari H, et al. The frequency of cytochrome 4F2 rs2108622 genetic variant and its effects on the lipid profile and complications of type II diabetes among a sample of patients in Jordan: A pilot study. Prostaglandins Other Lipid Mediat. 2023;165:106715.
Jarrar Y, Hamadneh L, Iqtait D, Sadieh R, Al‑Bawab AQ, Zihlif M. The cytochrome 4F2 rs2108622 genetic variant among unrelated Arab jordanian volunteers. Jordan Med J. 2017;51:69–76.
Johnson JA, Gong L, Whirl-Carrillo M, Gage BF, Scott SA, Stein CM, et al. Clinical pharmacogenetics implementation consortium. clinical pharmacogenetics implementation consortium guidelines for CYP2C9 and VKORC1 genotypes and warfarin dosing. Clin Pharmacol Ther. 2011;90:625–9.
Mizzi C, Dalabira E, Kumuthini J, Dzimiri N, Balogh I, Başak N, et al. A European spectrum of pharmacogenomic biomarkers: implications for clinical pharmacogenomics. PLoS One. 2016;11:e0162866.
Atasilp C, Biswas M, Jinda P, Nuntharadthanaphong N, Rachanakul J, Hongkaew Y, et al. Association of UGT1A1*6, UGT1A1*28, or ABCC2 c.3972C>T genetic polymorphisms with irinotecan-induced toxicity in Asian cancer patients: meta-analysis. Clin Transl Sci. 2022;15:1613–33.
Bertholim-Nasciben L, Scliar MO, Debortoli G, Thiruvahindrapuram B, Scherer SW, Duarte YAO, et al. Characterization of pharmacogenomic variants in a Brazilian admixed cohort of elderly individuals based on whole-genome sequencing data. Front Pharmacol. 2023;14:1178715.
Jarrar Y, Hakooz N, Ajamieh MA, Shawagfeh K, Zaidan R, Al Shareef O. Evaluation of pharmacogenomic information in drug labeling: a case study from Jordan. Per Med. 2025;22:1–7.
Acknowledgements
The authors would like to thank INNOVIA Biobank for supporting this research.
Funding
This research did not receive any funding.
Author information
Authors and Affiliations
Contributions
Conceptualization, YJ, NH, AB, and MA; methodology, DA, MH, SAA, MAG, and RA; formal analysis, YJ, DA, MH, and SAA; writing—original draft preparation, YJ DA; writing—review and editing, NH, AB, and MA. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jarrar, Y., Hakooz, N., Hashem, D.A. et al. Genome-wide screening of pharmacogenomic biomarkers in jordanian patients with genetic disorders. Pharmacogenomics J 25, 28 (2025). https://doi.org/10.1038/s41397-025-00387-w
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41397-025-00387-w