Abstract
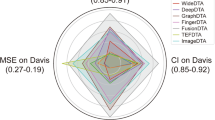
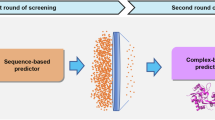
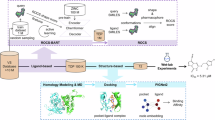
N-methyl-D-aspartate receptors (NMDARs) are glutamate-gated ion channels essential for synaptic transmission and plasticity in the central nervous system. GluN1/GluN3A, an unconventional NMDAR subtype functioning as an excitatory glycine receptor, has been implicated in mood regulation, with high expression in brain regions governing emotional and motivational states. However, therapeutic exploration has been significantly hindered by a lack of potent and selective modulators, limited structural data and the intrinsic complexity of ion channels. Here, we introduce a compound virtual screening pipeline that combines artificial intelligence and physical models, integrating two sequence-based deep learning prediction models (TEFDTA and ESMLigSite) with a molecular docking approach. This approach was employed to identify potential inhibitors against GluN1/GluN3A by screening a commercial database containing 18 million compounds. The strategy resulted in an impressive hit rate of 50% for discovering inhibitors, with the most promising compound exhibiting strong inhibitory activity (IC50 = 1.26 ± 0.23 μM) and remarkable target specificity (>23-fold selectivity over the GluN1/GluN2A receptor). These findings highlight the effectiveness of AI-assisted strategies in addressing challenges related to unconventional ion channels and pave the way for new therapeutic exploration.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Paoletti P, Bellone C, Zhou Q. NMDA receptor subunit diversity: impact on receptor properties, synaptic plasticity and disease. Nat Rev Neurosci. 2013;14:383–400.
Hansen KB, Yi F, Perszyk RE, Furukawa H, Wollmuth LP, Gibb AJ, et al. Structure, function, and allosteric modulation of NMDA receptors. J Gen Physiol. 2018;150:1081–105.
Hansen KB, Wollmuth LP, Bowie D, Furukawa H, Menniti FS, Sobolevsky AI, et al. Structure, function, and pharmacology of glutamate receptor ion channels. Pharmacol Rev. 2021;73:1469–658.
Das S, Sasaki YF, Rothe T, Premkumar LS, Takasu M, Crandall JE, et al. Increased NMDA current and spine density in mice lacking the NMDA receptor subunit NR3A. Nature. 1998;393:377–81.
Perez-Otano I, Schulteis CT, Contractor A, Lipton SA, Trimmer JS, Sucher NJ, et al. Assembly with the NR1 subunit is required for surface expression of NR3A-containing NMDA receptors. J Neurosci. 2001;21:1228–37.
Chatterton JE, Awobuluyi M, Premkumar LS, Takahashi H, Talantova M, Shin Y, et al. Excitatory glycine receptors containing the NR3 family of NMDA receptor subunits. Nature. 2002;415:793–8.
Pérez-Otaño I, Larsen RS, Wesseling JF. Emerging roles of GluN3-containing NMDA receptors in the CNS. Nat Rev Neurosci. 2016;17:623–35.
Sasaki YF, Rothe T, Premkumar LS, Das S, Cui J, Talantova MV, et al. Characterization and comparison of the NR3A subunit of the NMDA receptor in recombinant systems and primary cortical neurons. J Neurophysiol. 2002;87:2052–63.
Murillo A, Navarro AI, Puelles E, Zhang Y, Petros TJ, Pérez-Otaño I. Temporal dynamics and neuronal specificity of Grin3a expression in the mouse forebrain. Cereb Cortex. 2021;31:1914–26.
Marco S, Giralt A, Petrovic MM, Pouladi MA, Martínez-Turrillas R, Martínez-Hernández J, et al. Suppressing aberrant GluN3A expression rescues NMDA receptor dysfunction, synapse loss and motor and cognitive decline in Huntington’s disease models. Nat Med. 2013;19:1030.
Yuan T, Mameli M, O’Connor EC, Dey PN, Verpelli C, Sala C, et al. Expression of cocaine-evoked synaptic plasticity by GluN3A-containing NMDA receptors. Neuron. 2013;80:1025–38.
Glantz LA, Lewis DA. Decreased dendritic spine density on prefrontal cortical pyramidal neurons in schizophrenia. Arch Gen Psychiatry. 2000;57:65–73.
Pfisterer U, Petukhov V, Demharter S, Meichsner J, Thompson JJ, Batiuk MY, et al. Identification of epilepsy-associated neuronal subtypes and gene expression underlying epileptogenesis. Nat Commun. 2020;11:5038.
Bossi S, Pizzamiglio L, Paoletti P. Excitatory GluN1/GluN3A glycine receptors (eGlyRs) in brain signaling. Trends Neurosci. 2023;46:667–81.
Otsu Y, Darcq E, Pietrajtis K, Mátyás F, Schwartz E, Bessaih T, et al. Control of aversion by glycine-gated GluN1/GluN3A NMDA receptors in the adult medial habenula. Science. 2019;366:250–4.
Bossi S, Dhanasobhon D, Ellis-Davies GC, Frontera J, Van Velze MdB, Lourenço J, et al. GluN3A excitatory glycine receptors control adult cortical and amygdalar circuits. Neuron. 2022;110:2438–54.e8.
Yao Y, Mayer ML. Characterization of a soluble ligand binding domain of the NMDA receptor regulatory subunit NR3A. J Neurosci. 2006;26:4559–66.
Zhu Z, Yi F, Epplin MP, Liu D, Summer SL, Mizu R, et al. Negative allosteric modulation of GluN1/GluN3 NMDA receptors. Neuropharmacology. 2020;176:108117.
Kvist T, Greenwood JR, Hansen KB, Traynelis SF, Bräuner-Osborne H. Structure-based discovery of antagonists for GluN3-containing N-methyl-D-aspartate receptors. Neuropharmacology. 2013;75:324–36.
Zeng Y, Zheng Y, Zhang T, Ye F, Zhan L, Kou Z, et al. Identification of a subtype-selective allosteric inhibitor of GluN1/GluN3 NMDA receptors. Front Pharmacol. 2022;13:888308.
Rouzbeh N, Rau AR, Benton AJ, Yi F, Anderson CM, Johns MR, et al. Allosteric modulation of GluN1/GluN3 NMDA receptors by GluN1-selective competitive antagonists. J Gen Physiol. 2023;155:e202313340.
Li Z, Ren P, Yang H, Zheng J, Bai F. TEFDTA: a transformer encoder and fingerprint representation combined prediction method for bonded and non-bonded drug–target affinities. Bioinformatics. 2024;40:btad778.
Chen L, Tan X, Wang D, Zhong F, Liu X, Yang T, et al. TransformerCPI: improving compound–protein interaction prediction by sequence-based deep learning with self-attention mechanism and label reversal experiments. Bioinformatics. 2020;36:4406–14.
Zhao Q, Duan G, Yang M, Cheng Z, Li Y, Wang J. AttentionDTA: Drug–target binding affinity prediction by sequence-based deep learning with attention mechanism. IEEE/ACM Trans Comput Biol Bioinform. 2022;20:852–63.
Nguyen TM, Nguyen T, Le TM, Tran T. Gefa: early fusion approach in drug-target affinity prediction. IEEE/ACM Trans Comput Biol Bioinform. 2021;19:718–28.
Wang K, Zhou R, Li Y, Li M. DeepDTAF: a deep learning method to predict protein–ligand binding affinity. Brief Bioinform. 2021;22:bbab072.
Wang L, Wang S, Yang H, Li S, Wang X, Zhou Y, et al. Conformational space profiling enhances generic molecular representation for AI‐powered ligand‐based drug discovery. Adv Sci. 2024:2403998.
Wang R, Fang X, Lu Y, Yang CY, Wang S. The PDBbind database: methodologies and updates. J Med Chem. 2005;48:4111–9.
Burley SK, Berman HM, Bhikadiya C, Bi C, Chen L, Di Costanzo L, et al. RCSB Protein Data Bank: biological macromolecular structures enabling research and education in fundamental biology, biomedicine, biotechnology and energy. Nucleic Acids Res. 2019;47:D464–D74.
Lin Z, Akin H, Rao R, Hie B, Zhu Z, Lu W, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science. 2023;379:1123–30.
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition. Las Vegas (USA). 2016–2016. p 770–8.
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–9.
Knox C, Wilson M, Klinger CM, Franklin M, Oler E, Wilson A, et al. DrugBank 6.0: the DrugBank knowledgebase for 2024. Nucleic Acids Res. 2024;52:D1265–D75.
Madhavi Sastry G, Adzhigirey M, Day T, Annabhimoju R, Sherman W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des. 2013;27:221–34.
Greenwood JR, Calkins D, Sullivan AP, Shelley JC. Towards the comprehensive, rapid, and accurate prediction of the favorable tautomeric states of drug-like molecules in aqueous solution. J Comput Aided Mol Des. 2010;24:591–604.
Yang Y, Yao K, Repasky MP, Leswing K, Abel R, Shoichet BK, et al. Efficient exploration of chemical space with docking and deep learning. J Chem Theory Comput. 2021;17:7106–19.
Ogden KK, Traynelis SF. Contribution of the M1 transmembrane helix and pre-M1 region to positive allosteric modulation and gating of N-methyl-D-aspartate receptors. Mol Pharmacol. 2013;83:1045–56.
Chaput L, Martinez-Sanz J, Saettel N, Mouawad L. Benchmark of four popular virtual screening programs: construction of the active/decoy dataset remains a major determinant of measured performance. J Cheminform. 2016;8:1–17.
Sliwoski G, Kothiwale S, Meiler J, Lowe EW. Computational methods in drug discovery. Pharmacol Rev. 2014;66:334–95.
Wee KSL, Zhang Y, Khanna S, Low CM. Immunolocalization of NMDA receptor subunit NR3B in selected structures in the rat forebrain, cerebellum, and lumbar spinal cord. J Comp Neurol. 2008;509:118–35.
Nishi M, Hinds H, Lu HP, Kawata M, Hayashi Y. Motoneuron-specific expression of NR3B, a novel NMDA-type glutamate receptor subunit that works in a dominant-negative manner. J Neurosci. 2001;21:RC185.
Acknowledgements
This work was supported by Shanghai Science and Technology Development Funds (Grant IDs: 24JS2850200 and 24JS2850100), the Strategic Priority Research Program of the Chinese Academy of Sciences (Grant ID: XDB0830403), ShanghaiTech AI4S Initiative SHTAI4S202404, National Key R&D Program of China (Grant IDs: 2022YFC3400501 & 2022YFC3400500), start-up package from ShanghaiTech University, and Shanghai Frontiers Science Center for Biomacromolecules and Precision Medicine at ShanghaiTech University, and the National Science and Technology Innovation 2030 Major Program (Grant ID: 2021ZD0200900). The authors appreciate the technical support provided by the high-performance computing cluster of ShanghaiTech University.
Author information
Authors and Affiliations
Contributions
FB and ZBG designed the study. FB and SWL proposed the screening pipeline, SNW, CX and XYM collected corresponding data for the screening, ZQL conducted initial virtual screening, SWL, SNW and CX conducted fine screening. YZ, SF, and XQC designed and conducted wet-lab experiments. SWL and YZ wrote the manuscript. All authors reviewed the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Sw., Zeng, Y., Wu, Sn. et al. Discovery of selective GluN1/GluN3A NMDA receptor inhibitors using integrated AI and physics-based approaches. Acta Pharmacol Sin 47, 22–31 (2026). https://doi.org/10.1038/s41401-025-01607-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41401-025-01607-6
Keywords
This article is cited by
-
AI-driven breakthroughs in ion channel drug discovery: the future is now
Acta Pharmacologica Sinica (2026)
-
New advances in small molecule drugs targeting NMDA receptors
Acta Pharmacologica Sinica (2026)