Abstract
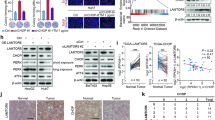
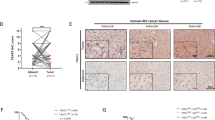
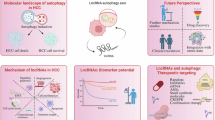
In the etiology of cancer, p62 is a well-known autophagic receptor and signaling adapter. High p62 expression is known to accelerate hepatocellular carcinoma (HCC) growth by activating various downstream signaling pathways. In this study, we investigated the activity of elevated p62 and its associated regulatory mechanisms during HCC progression. By conducting immunohistochemical staining on a human liver tissue microarray including 10 liver cancer tissues and 10 paracancerous tissues, we found that the expression levels of p62 and oncoprotein LAMTOR5 were markedly increased in HCC tissues compared with noncancerous tissues; LAMTOR5 was positively associated with p62 expression, and high LAMTOR5 or p62 expression predicted reduced overall and release-free survival. Transcriptomic analysis revealed that LAMTOR5 overexpression inhibited autophagy in HepG2 cells. We demonstrated that LAMTOR5 interacted with the LC3-interacting region domain of p62 and inhibited autophagy caused by the binding of p62 to LC3, thereby leading to the accumulation of p62 protein in HCC. Moreover, LAMTOR5 blocked p62 ubiquitination-mediated proteasome degradation, which increased the stability of p62. Functionally, p62 overexpression reversed LAMTOR5 deficiency-reduced hepatoma cell proliferation in vitro and in vivo. Lenvatinib, a multi-receptor tyrosine kinase inhibitor, significantly suppressed HCC growth in vitro and in vivo by downregulating LAMTOR5 and p62 expression. We conclude that LAMTOR5-mediated p62 stabilization is a novel HCC growth mechanism, targeting this axis as a promising therapeutic strategy.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Siegel RL, Kratzer TB, Giaquinto AN, Sung H, Jemal A. Cancer statistics, 2025. CA Cancer J Clin. 2025;75:10–45.
Chan SL, Sun HC, Xu Y, Zeng H, El-Serag HB, Lee JM, et al. The Lancet Commission on addressing the global hepatocellular carcinoma burden: comprehensive strategies from prevention to treatment. Lancet. 2025;406:731–78.
Zhao Y, Zhang YN, Wang KT, Chen L. Lenvatinib for hepatocellular carcinoma: From preclinical mechanisms to anti-cancer therapy. Biochim Biophys Acta Rev Cancer. 2020;1874:188391.
Al-Salama ZT, Syed YY, Scott LJ. Lenvatinib: a review in hepatocellular carcinoma. Drugs. 2019;79:665–74.
Zhang H, Zhang Y, Zhu X, Chen C, Zhang C, Xia Y, et al. DEAD box protein 5 inhibits liver tumorigenesis by stimulating autophagy via interaction with p62/SQSTM1. Hepatology. 2019;69:1046–63.
Zhang M, Liu S, Chua MS, Li H, Luo D, Wang S, et al. SOCS5 inhibition induces autophagy to impair metastasis in hepatocellular carcinoma cells via the PI3K/Akt/mTOR pathway. Cell Death Dis. 2019;10:612.
Zhou X, Chen D, Yan Y, Li Q, Xing W, Liu Y, et al. The nociceptin receptor promotes autophagy through NF-kB signaling and is transcriptionally regulated by E2F1 in HCC. Cell Death Discov. 2022;8:165.
Liu Z, Chen P, Gao H, Gu Y, Yang J, Peng H, et al. Ubiquitylation of autophagy receptor Optineurin by HACE1 activates selective autophagy for tumor suppression. Cancer Cell. 2014;26:106–20.
Deng D, Luo K, Liu H, Nie X, Xue L, Wang R, et al. p62 acts as an oncogene and is targeted by miR-124-3p in glioma. Cancer Cell Int. 2019;19:280.
Tao M, Liu T, You Q, Jiang Z. p62 as a therapeutic target for tumor. Eur J Med Chem. 2020;193:112231.
Moscat J, Diaz-Meco MT. p62: a versatile multitasker takes on cancer. Trends Biochem Sci. 2012;37:230–6.
Liu JL, Chen FF, Lung J, Lo CH, Lee FH, Lu YC, et al. Prognostic significance of p62/SQSTM1 subcellular localization and LC3B in oral squamous cell carcinoma. Br J Cancer. 2014;111:944–54.
Umemura A, He F, Taniguchi K, Nakagawa H, Yamachika S, Font-Burgada J, et al. p62, upregulated during preneoplasia, induces hepatocellular carcinogenesis by maintaining survival of stressed HCC-initiating cells. Cancer Cell. 2016;29:935–48.
Xu Y, Zhu C, Zhu C, Peng L, Ji D, Wu Q, et al. SQSTM1/p62 promotes the progression of gastric cancer through epithelial-mesenchymal transition. Heliyon. 2024;10:e24409.
Duran A, Linares JF, Galvez AS, Wikenheiser K, Flores JM, Diaz-Meco MT, et al. The signaling adaptor p62 is an important NF-kappaB mediator in tumorigenesis. Cancer Cell. 2008;13:343–54.
Xu D, Li X, Shao F, Lv G, Lv H, Lee JH, et al. The protein kinase activity of fructokinase A specifies the antioxidant responses of tumor cells by phosphorylating p62. Sci Adv. 2019;5:eaav4570.
Peng H, Yang J, Li G, You Q, Han W, Li T, et al. Ubiquitylation of p62/sequestosome1 activates its autophagy receptor function and controls selective autophagy upon ubiquitin stress. Cell Res. 2017;27:657–74.
Lee Y, Weihl CC. Regulation of SQSTM1/p62 via UBA domain ubiquitination and its role in disease. Autophagy. 2017;13:1615–6.
You Z, Jiang WX, Qin LY, Gong Z, Wan W, Li J, et al. Requirement for p62 acetylation in the aggregation of ubiquitylated proteins under nutrient stress. Nat Commun. 2019;10:5792.
Song P, Li S, Wu H, Gao R, Rao G, Wang D, et al. Parkin promotes proteasomal degradation of p62: implication of selective vulnerability of neuronal cells in the pathogenesis of Parkinson’s disease. Protein Cell. 2016;7:114–29.
Pankiv S, Clausen TH, Lamark T, Brech A, Bruun JA, Outzen H, et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J Biol Chem. 2007;282:24131–45.
Tang C, Lai Y, Li L, Situ MY, Li S, Cheng B, et al. SERPINH1 modulates apoptosis by inhibiting P62 ubiquitination degradation to promote bone metastasis of prostate cancer. iScience. 2024;27:110427.
Bar-Peled L, Schweitzer LD, Zoncu R, Sabatini DM. Ragulator is a GEF for the rag GTPases that signal amino acid levels to mTORC1. Cell. 2012;150:1196–208.
Li Y, Wang Z, Shi H, Li H, Li L, Fang R, et al. HBXIP and LSD1 scaffolded by lncRNA hotair mediate transcriptional activation by c-Myc. Cancer Res. 2016;76:293–304.
Liu B, Wang T, Wang H, Zhang L, Xu F, Fang R, et al. Oncoprotein HBXIP enhances HOXB13 acetylation and co-activates HOXB13 to confer tamoxifen resistance in breast cancer. J Hematol Oncol. 2018;11:26.
Fang R, Xu F, Shi H, Wu Y, Cao C, Li H, et al. LAMTOR5 raises abnormal initiation of O-glycosylation in breast cancer metastasis via modulating GALNT1 activity. Oncogene. 2019;39:2290–2230.
Wu Y, Wang X, Xu F, Zhang L, Wang T, Fu X, et al. The regulation of acetylation and stability of HMGA2 via the HBXIP-activated Akt-PCAF pathway in promotion of esophageal squamous cell carcinoma growth. Nucleic Acids Res. 2020;48:4858–76.
Xu FF, Sun HM, Fang RP, Zhang L, Shi H, Wang X, et al. The modulation of PD-L1 induced by the oncogenic HBXIP for breast cancer growth. Acta Pharmacol Sin. 2022;43:429–45.
Zhang L, Zhou X, Liu B, Shi X, Li X, Xu F, et al. HBXIP blocks myosin-IIA assembly by phosphorylating and interacting with NMHC-IIA in breast cancer metastasis. Acta Pharm Sin B. 2023;13:1053–70.
Fu XL, Guo SM, Ma JQ, Ma FY, Wang X, Tang YX, et al. HBXIP induces PARP1 via WTAP-mediated m(6)A modification and CEBPA-activated transcription in cisplatin resistance to hepatoma. Acta Pharmacol Sin. 2024;45:2405–19.
Marusawa H, Matsuzawa S, Welsh K, Zou H, Armstrong R, Tamm I, et al. HBXIP functions as a cofactor of survivin in apoptosis suppression. EMBO J. 2003;22:2729–40.
Garcia-Saez I, Lacroix FB, Blot D, Gabel F, Skoufias DA. Structural characterization of HBXIP: the protein that interacts with the anti-apoptotic protein survivin and the oncogenic viral protein HBx. J Mol Biol. 2011;405:331–40.
Zhang W, Lu Z, Kong G, Gao Y, Wang T, Wang Q, et al. Hepatitis B virus X protein accelerates hepatocarcinogenesis with partner survivin through modulating miR-520b and HBXIP. Mol Cancer. 2014;13:1–14.
Xu F, You X, Liu F, Shen X, Yao Y, Ye L, et al. The oncoprotein HBXIP up-regulates Skp2 via activating transcription factor E2F1 to promote proliferation of breast cancer cells. Cancer Lett. 2013;333:124–32.
Shi H, Fang R, Li Y, Li L, Zhang W, Wang H, et al. The oncoprotein HBXIP suppresses gluconeogenesis through modulating PCK1 to enhance the growth of hepatoma cells. Cancer Lett. 2016;382:147–56.
Wang Y, Fang R, Cui M, Zhang W, Bai X, Wang H, et al. The oncoprotein HBXIP up-regulates YAP through activation of transcription factor c-Myb to promote growth of liver cancer. Cancer Lett. 2017;385:234–42.
Jiang Y, Wang D, Ren H, Shi Y, Gao Y. Oncogenic HBXIP enhances ZEB1 through Sp1 to accelerate breast cancer growth. Thorac Cancer. 2018;9:1664–70.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60.
Zhang Y, Mun SR, Linares JF, Towers CG, Thorburn A, Diaz-Meco MT, et al. Mechanistic insight into the regulation of SQSTM1/p62. Autophagy. 2019;15:735–7.
Zhang D, Zhang Y, Pan J, Cao J, Sun X, Li X, et al. Degradation of NLRP3 by p62-dependent-autophagy improves cognitive function in Alzheimer’s disease by maintaining the phagocytic function of microglia. CNS Neurosci Ther. 2023;29:2826–42.
Kurusu R, Fujimoto Y, Morishita H, Noshiro D, Takada S, Yamano K, et al. Integrated proteomics identifies p62-dependent selective autophagy of the supramolecular vault complex. Dev Cell. 2023;58:1189–1205 e1111.
Moscat J, Karin M, Diaz-Meco MT. p62 in cancer: signaling adaptor beyond autophagy. Cell. 2016;167:606–9.
Wang Y, Cui M, Cai X, Sun B, Liu F, Zhang X, et al. The oncoprotein HBXIP up-regulates SCG3 through modulating E2F1 and miR-509-3p in hepatoma cells. Cancer Lett. 2014;352:169–78.
Jiang TX, Zou JB, Zhu QQ, Liu CH, Wang GF, Du TT, et al. SIP/CacyBP promotes autophagy by regulating levels of BRUCE/Apollon, which stimulates LC3-I degradation. Proc Natl Acad Sci USA. 2019;116:13404–13.
Matsumoto G, Wada K, Okuno M, Kurosawa M, Nukina N. Serine 403 phosphorylation of p62/SQSTM1 regulates selective autophagic clearance of ubiquitinated proteins. Mol Cell. 2011;44:279–89.
Li L, Fang R, Liu B, Shi H, Wang Y, Zhang W, et al. Deacetylation of tumor-suppressor MST1 in Hippo pathway induces its degradation through HBXIP-elevated HDAC6 in promotion of breast cancer growth. Oncogene. 2016;35:4048–57.
Zhang L, Li XM, Shi XH, Ye K, Fu XL, Wang X, et al. Sorafenib triggers ferroptosis via inhibition of HBXIP/SCD axis in hepatocellular carcinoma. Acta Pharmacol Sin. 2023;44:622–34.
Zhao Y, Li H, Zhang Y, Li L, Fang R, Li Y, et al. Oncoprotein HBXIP modulates abnormal lipid metabolism and growth of breast cancer cells by activating the LXRs/SREBP-1c/FAS signaling cascade. Cancer Res. 2016;76:4696–707.
Chen M, Dong X, Deng H, Ye F, Zhao Y, Cheng J, et al. Targeting TRPV1-mediated autophagy attenuates nitrogen mustard-induced dermal toxicity. Signal Transduct Target Ther. 2021;6:29.
Wu Y, Zhang Y, Qin X, Geng H, Zuo D, Zhao Q. PI3K/AKT/mTOR pathway-related long non-coding RNAs: roles and mechanisms in hepatocellular carcinoma. Pharmacol Res. 2020;160:105195.
Wang X, Li QQ, Tang YX, Li Y, Zhang L, Xu FF, et al. Oncoprotein LAMTOR5-mediated CHOP silence via DNA hypermethylation and miR-182/miR-769 in promotion of liver cancer growth. Acta Pharmacol Sin. 2024;45:2625–45.
Fang R, Xu F, Shi H, Wu Y, Cao C, Li H, et al. LAMTOR5 raises abnormal initiation of O-glycosylation in breast cancer metastasis via modulating GALNT1 activity. Oncogene. 2020;39:2290–304.
Zhou J, Li Y, Li D, Liu Z, Zhang J. Oncoprotein LAMTOR5 activates GLUT1 via upregulating NF-kappaB in liver cancer. Open Med. 2019;14:264–70.
Mondal G, Gonzalez H, Marsh T, Leidal AM, Vlahakis A, Phadatare PR, et al. Autophagy-targeted NBR1-p62/SQSTM1 complexes promote breast cancer metastasis by sequestering ITCH. Nat Cell Biol. 2025;27:1098–113.
Huang X, Zhang J, Yao J, Mi N, Yang A. Phase separation of p62: roles and regulations in autophagy. Trends Cell Biol. 2025;35:854–65.
Jiang G, Liang X, Huang Y, Lan Z, Zhang Z, Su Z, et al. p62 promotes proliferation, apoptosisresistance and invasion of prostate cancer cells through the Keap1/Nrf2/ARE axis. Oncol Rep. 2020;43:1547–57.
Yang P, Gao S, Shen J, Liu T, Lu K, Han X, et al. TRIM21-mediated ubiquitination of SQSTM1/p62 abolishes its Ser403 phosphorylation and enhances palmitic acid cytotoxicity. Autophagy. 2025;21:178–90.
Shan Q, Yin L, Zhan Q, Yu J, Pan S, Zhuo J, et al. The p-MYH9/USP22/HIF-1α axis promotes lenvatinib resistance and cancer stemness in hepatocellular carcinoma. Signal Transduct Target Ther. 2024;9:249.
Acknowledgements
We thank Zhuo-wei Hu (Chinese Academy of Medical Sciences, Peking Union Medical College) for Flag-tagged p62 plasmids and Xiao-bo Qiu (Beijing Normal University) for GFP-RFP-LC3 plasmids. This work was supported by the National Natural Scientific Foundation of China [82202950, 82072943, and 82072929].
Author information
Authors and Affiliations
Contributions
FFX, ZTZ, WYZ, and LHY designed the research; FFX, HMS, YL, KY, ZYL, and XLF conducted experiments; FFX, HMS, and YL analyzed data and wrote the main paper; ZTZ, WYZ, and LHY revised the paper. All authors read and provided approval for the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, Ff., Sun, Hm., Liu, Y. et al. LAMTOR5 promotes hepatoma growth in mice by disrupting LC3-p62-mediated autophagy and preventing p62 proteasome degradation. Acta Pharmacol Sin 47, 1042–1057 (2026). https://doi.org/10.1038/s41401-025-01687-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41401-025-01687-4