Abstract
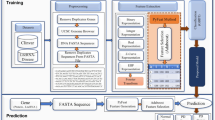
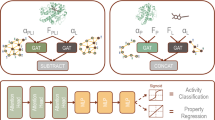
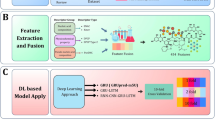
Accurately quantifying protein-DNA interactions (PDIs) is critical for understanding biological processes and facilitating drug design. However, the inherent flexibility of nucleic acids limits the availability of experimentally determined structures of PDI complexes, posing a significant challenge for training reliable scoring functions (SFs). To address this, we developed PDIScore, a novel deep learning-based SF for PDI prediction. PDIScore utilizes a comprehensive graph representation to capture nucleotide flexibility, employs a scalable GraphGPS architecture with BigBird linear global attention to handle large interaction interfaces, and leverages Mixture Density Networks (MDNs) to model residue-nucleotide distance distributions. PDIScore was trained on a self-collected dataset of ~7000 protein-nucleic acid complex structures and validated on three rigorous test sets for evaluating its screening, docking, and ranking capabilities. The results illustrated that PDIScore significantly outperformed existing methods: it achieved the best screening power on the screening set (e.g., EF1% = 14.13, AUROC = 0.82 using AlphaFold3 structures), the highest docking success rate on the docking set (48.94% top1), and superior ranking capability on the ranking set (PCC = 0.50). Case studies demonstrated PDIScore’s ability to elucidate biological mechanisms (e.g., adenovirus transcription, SOCS1 regulation) and its interpretability at the nucleotide level for identifying key interaction sites. PDIScore represents a robust, generalizable tool with significant potential for advancing PDI-related research and therapeutic design.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets are available at https://doi.org/10.5281/zenodo.13764615. The code is available at https://github.com/roger-yh-zhao/pdiscore.
References
Charoensawan V, Wilson D, Teichmann SA. Genomic repertoires of DNA-binding transcription factors across the tree of life. Nucleic Acids Res. 2010;38:7364–77.
Markodimitraki CM, Rang FJ, Rooijers K, de Vries SS, Chialastri A, de Luca KL, et al. Simultaneous quantification of protein-DNA interactions and transcriptomes in single cells with scDam&T-seq. Nat Protoc. 2020;15:1922–53.
Yu L, Liu P. Cytosolic DNA sensing by cGAS: regulation, function, and human diseases. Signal Transduct Target Ther. 2021;6:170.
Wells A, Heckerman D, Torkamani A, Yin L, Sebat J, Ren B, et al. Ranking of non-coding pathogenic variants and putative essential regions of the human genome. Nat Commun. 2019;10:5241.
Jiao W, Atwal G, Polak P, Karlic R, Cuppen E, Subtypes PT, et al. A deep learning system accurately classifies primary and metastatic cancers using passenger mutation patterns. Nat Commun. 2020;11:728.
Guo AD, Yan KN, Hu H, Zhai L, Hu TF, Su H, et al. Spatiotemporal and global profiling of DNA-protein interactions enables discovery of low-affinity transcription factors. Nat Chem. 2023;15:803–14.
Chan LL, Pineda M, Heeres JT, Hergenrother PJ, Cunningham BT. A general method for discovering inhibitors of protein-DNA interactions using photonic crystal biosensors. ACS Chem Biol. 2008;3:437–48.
Barabasi AL, Gulbahce N, Loscalzo J. Network medicine: a network-based approach to human disease. Nat Rev Genet. 2011;12:56–68.
Nikolov DB, Chen H, Halay ED, Usheva AA, Hisatake K, Lee DK, et al. Crystal structure of a TFIIB-TBP-TATA-element ternary complex. Nature. 1995;377:119–28.
Hashimoto H, Olanrewaju YO, Zheng Y, Wilson GG, Zhang X, Cheng X. Wilms tumor protein recognizes 5-carboxylcytosine within a specific DNA sequence. Genes Dev. 2014;28:2304–13.
Kays AR, Schepartz A. Virtually unidirectional binding of TBP to the AdMLP TATA box within the quaternary complex with TFIIA and TFIIB. Chem Biol. 2000;7:601–10.
Mostecki J, Showalter BM, Rothman PB. Early growth response-1 regulates lipopolysaccharide-induced suppressor of cytokine signaling-1 transcription. J Biol Chem. 2005;280:2596–605.
Sobah ML, Liongue C, Ward AC. SOCS proteins in immunity, inflammatory diseases, and immune-related cancer. Front Med. 2021;8:727987.
Sefah K, Shangguan D, Xiong X, O’Donoghue MB, Tan W. Development of DNA aptamers using Cell-SELEX. Nat Protoc. 2010;5:1169–85.
Huynh L, Chen A. Design of a protein-targeted DNA aptamer using atomistic simulation. J Biomol Struct Dyn. 2023;41:672–80.
Wang X, Wang Y, Cao A, Luo Q, Chen D, Zhao W, et al. Development of cyclopeptide inhibitors of cGAS targeting protein-DNA interaction and phase separation. Nat Commun. 2023;14:6132.
Kimoto M, Yamashige R, Matsunaga K, Yokoyama S, Hirao I. Generation of high-affinity DNA aptamers using an expanded genetic alphabet. Nat Biotechnol. 2013;31:453–7.
Hellman LM, Fried MG. Electrophoretic mobility shift assay (EMSA) for detecting protein-nucleic acid interactions. Nat Protoc. 2007;2:1849–61.
Hadzi S, Lah J. Analysis of protein-DNA interactions using isothermal titration calorimetry: successes and failures. Methods Mol Biol. 2022;2516:239–57.
Majka J, Speck C. Analysis of protein-DNA interactions using surface plasmon resonance. Adv Biochem Eng Biotechnol. 2007;104:13–36.
Townshend RJL, Eismann S, Watkins AM, Rangan R, Karelina M, Das R, et al. Geometric deep learning of RNA structure. Science. 2021;373:1047–51.
Roche R, Moussad B, Shuvo MH, Tarafder S, Bhattacharya D. EquiPNAS: improved protein-nucleic acid binding site prediction using protein-language-model-informed equivariant deep graph neural networks. Nucleic Acids Res. 2024;52:e27.
Delgado J, Radusky LG, Cianferoni D, Serrano L. FoldX 5.0: working with RNA, small molecules and a new graphical interface. Bioinformatics. 2019;35:4168–9.
Schymkowitz J, Borg J, Stricher F, Nys R, Rousseau F, Serrano L. The FoldX web server: an online force field. Nucleic Acids Res. 2005;33:W382–8.
Wang E, Sun H, Wang J, Wang Z, Liu H, Zhang JZH, et al. End-point binding free energy calculation with MM/PBSA and MM/GBSA: strategies and applications in drug design. Chem Rev. 2019;119:9478–508.
Chaudhury S, Lyskov S, Gray JJ. PyRosetta: a script-based interface for implementing molecular modeling algorithms using Rosetta. Bioinformatics. 2010;26:689–91.
Yang W, Deng L. PreDBA: a heterogeneous ensemble approach for predicting protein-DNA binding affinity. Sci Rep. 2020;10:1278.
Yang S, Gong W, Zhou T, Sun X, Chen L, Zhou W, et al. emPDBA: protein-DNA binding affinity prediction by combining features from binding partners and interface learned with ensemble regression model. Brief Bioinform. 2023;24:bbad192.
Zhang X, Mei LC, Gao YY, Hao GF, Song BA. Web tools support predicting protein-nucleic acid complexes stability with affinity changes. WIREs RNA. 2023;14:e1781.
Yan Y, Zhang D, Zhou P, Li B, Huang SY. HDOCK: a web server for protein-protein and protein-DNA/RNA docking based on a hybrid strategy. Nucleic Acids Res. 2017;45:W365–73.
Rodriguez-Lumbreras LA, Jimenez-Garcia B, Gimenez-Santamarina S, Fernandez-Recio J. pyDockDNA: a new web server for energy-based protein-DNA docking and scoring. Front Mol Biosci. 2022;9:988996.
van Zundert GCP, Rodrigues J, Trellet M, Schmitz C, Kastritis PL, Karaca E, et al. The HADDOCK2.2 web server: user-friendly integrative modeling of biomolecular complexes. J Mol Biol. 2016;428:720–5.
Honorato RV, Koukos PI, Jimenez-Garcia B, Tsaregorodtsev A, Verlato M, Giachetti A, et al. Structural biology in the clouds: the WeNMR-EOSC ecosystem. Front Mol Biosci. 2021;8:729513.
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–9.
Abramson J, Adler J, Dunger J, Evans R, Green T, Pritzel A, et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature. 2024;630:493–500.
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, et al. The Protein Data Bank. Nucleic Acids Res. 2000;28:235–42.
Afek A, Shi H, Rangadurai A, Sahay H, Senitzki A, Xhani S, et al. DNA mismatches reveal conformational penalties in protein-DNA recognition. Nature. 2020;587:291–6.
van Dijk M, Bonvin AM. A protein-DNA docking benchmark. Nucleic Acids Res. 2008;36:e88.
Larkin C, Datta S, Harley MJ, Anderson BJ, Ebie A, Hargreaves V, et al. Inter- and intramolecular determinants of the specificity of single-stranded DNA binding and cleavage by the F factor relaxase. Structure. 2005;13:1533–44.
Li J, Lee JC. Modulation of allosteric behavior through adjustment of the differential stability of the two interacting domains in E. coli cAMP receptor protein. Biophys Chem. 2011;159:210–6.
Yu S, Maillard RA, Gribenko AV, Lee JC. The N-terminal capping propensities of the D-helix modulate the allosteric activation of the Escherichia coli cAMP receptor protein. J Biol Chem. 2012;287:39402–11.
Seldeen KL, Deegan BJ, Bhat V, Mikles DC, McDonald CB, Farooq A. Energetic coupling along an allosteric communication channel drives the binding of Jun-Fos heterodimeric transcription factor to DNA. FEBS J. 2011;278:2090–104.
Rampášek L, Galkin M, Dwivedi VP, Luu AT, Wolf G, Beaini D. Recipe for a general, powerful, scalable graph transformer. NeurIPS. 2022;35:14501–15.
Zaheer M, Guruganesh G, Dubey KA, Ainslie J, Alberti C, Ontanon S, et al. Big bird: transformers for longer sequences. NeurIPS. 2020;33:17283–97.
Shen C, Zhang X, Hsieh CY, Deng Y, Wang D, Xu L, et al. A generalized protein-ligand scoring framework with balanced scoring, docking, ranking and screening powers. Chem Sci. 2023;14:8129–46.
Shen C, Zhang X, Deng Y, Gao J, Wang D, Xu L, et al. Boosting protein-ligand binding pose prediction and virtual screening based on residue-atom distance likelihood potential and graph transformer. J Med Chem. 2022;65:10691–706.
Wang R, Fang X, Lu Y, Wang S. The PDBbind database: collection of binding affinities for protein-ligand complexes with known three-dimensional structures. J Med Chem. 2004;47:2977–80.
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, et al. UCSF Chimera-a visualization system for exploratory research and analysis. J Comput Chem. 2004;25:1605–12.
Leaver-Fay A, Tyka M, Lewis SM, Lange OF, Thompson J, Jacak R, et al. ROSETTA3: an object-oriented software suite for the simulation and design of macromolecules. Methods Enzymol. 2011;487:545–74.
Lu XJ, Olson WK. 3DNA: a software package for the analysis, rebuilding and visualization of three-dimensional nucleic acid structures. Nucleic Acids Res. 2003;31:5108–21.
Steinegger M, Soding J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat Biotechnol. 2017;35:1026–8.
Michaud-Agrawal N, Denning EJ, Woolf TB, Beckstein O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J Comput Chem. 2011;32:2319–27.
Wang M, Xu L, Zheng D, Gan Q, Gai Y, Ye Z, et al. Deep graph library: towards efficient and scalable deep learning on graphs. Preprint at. https://doi.org/10.48550/arXiv.1909.01315 (2019).
Hu W, Liu B, Gomes J, Zitnik M, Liang P, Pande V, et al. In presented in part at the International Conference on Learning Representations (ICLR), (ICLR, 2020).
Bauer MR, Ibrahim TM, Vogel SM, Boeckler FM. Evaluation and optimization of virtual screening workflows with DEKOIS 2.0-a public library of challenging docking benchmark sets. J Chem Inf Model. 2013;53:1447–62.
Concino M, Goldman RA, Caruthers MH, Weinmann R. Point mutations of the adenovirus major late promoter with different transcriptional efficiencies in vitro. J Biol Chem. 1983;258:8493–6.
Ilangumaran S, Gui Y, Shukla A, Ramanathan S. SOCS1 expression in cancer cells: potential roles in promoting antitumor immunity. Front Immunol. 2024;15:1362224.
Acknowledgements
This work was financially supported by National Natural Science Foundation of China (22220102001, 22307112, 92370130, 22303081) and Postdoctoral Fellowship Program of CPSF (GZB20230648).
Author information
Authors and Affiliations
Contributions
TJH and YK designed the research study. YHZ developed the method and wrote the code. YHZ, YW, CS, DJJ, SKG, HFZ, and ZYY performed the analysis. YHZ and TJH wrote the paper. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, Yh., Wang, Y., Shen, C. et al. Graph-based deep learning approach for high-throughput protein-DNA interaction scoring. Acta Pharmacol Sin 47, 977–989 (2026). https://doi.org/10.1038/s41401-025-01688-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41401-025-01688-3