Abstract
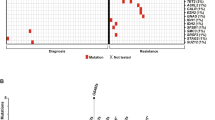
Contemporary risk models in chronic myelomonocytic leukemia (CMML) focus on the prognostic relevance of individual rather than concurrent mutations. In the current study of 605 Mayo Clinic patients with CMML, we applied machine-learning algorithms in order to examine the influence of cooperative mutational interactions on blast transformation (BT). A hierarchical clustering algorithm was developed and tailored for patient stratification using survival outcomes and co-occurrence of genomic alterations. Five molecular clusters were identified with 3-year blast BT rates ranging from 0% to 100% (AUC at 3 years 0.78). A subsequent Cox regression analysis confirmed independent detrimental impact of specific mutations or their combinations including NPM1 (HR 26.7; p < 0.01), “NRAS + SETBP1” (HR 12.6; p < 0.01), “ASXL1 + BCOR” (HR 8.4; p < 0.01), “ASXL1 + RUNX1” (HR 2.2, p < 0.01), JAK2 (HR 2.1; p < 0.01), and “ASXL1 + TET2” (HR 1.7; p = 0.02) while “PHF6+wild-type ASXL1” (HR 5.61e−10; p < 0.01) had a favorable impact. Furthermore, compared to NPM1 wild-type cases, NPM1-mutated patients were less likely to have co-occurring mutations involving ASXL1 (0% vs. 43%, p < 0.01), RUNX1 (0% vs. 17%, p = 0.02), and SRSF2 (7% vs. 39%, p < 0.01) and were more likely DNMT3A (71% vs. 7%, p < 0.01). The prognostic relevance of “NRAS + SETBP1”, “ASXL1 + RUNX1”, NPM1 and BCOR was validated in an external cohort from Italy (N = 501). Taken together, these observations highlight i) the possibility of prognostic interaction of mutations in CMML that should be considered in the development of future risk models and ii) the distinct genotypic and prognostic characteristics of NPM1-mutated CMML.

Similar content being viewed by others
Data availability
By email request to the corresponding author.
References
Khoury JD, Solary E, Abla O, Akkari Y, Alaggio R, Apperley JF, et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/Dendritic Neoplasms. Leukemia. 2022;36:1703–1719.
Arber DA, Orazi A, Hasserjian RP, Borowitz MJ, Calvo KR, Kvasnicka HM, et al. International Consensus Classification of Myeloid Neoplasms and Acute Leukemias: integrating morphologic, clinical, and genomic data. Blood. 2022;140:1200–1228.
Patnaik MM, Tefferi A. Chronic myelomonocytic leukemia: 2024 update on diagnosis, risk stratification and management. Am J Hematol. 2024;99:1142–1165.
Tefferi A, Fathima S, Abdelmagid M, Alsugair A, Aperna F, Rezasoltani M, et al. BLAST: a globally applicable and molecularly versatile survival model for chronic myelomonocytic leukemia. Blood. 2025;146:874-886.
Patnaik MM, Wassie EA, Lasho TL, Hanson CA, Ketterling R, Tefferi A. Blast transformation in chronic myelomonocytic leukemia: Risk factors, genetic features, survival, and treatment outcome. Am J Hematol. 2015;90:411–416.
Elena C, Gallì A, Such E, Meggendorfer M, Germing U, Rizzo E, et al. Integrating clinical features and genetic lesions in the risk assessment of patients with chronic myelomonocytic leukemia. Blood. 2016;128:1408–1417.
Patnaik MM, Padron E, LaBorde RR, Lasho TL, Finke CM, Hanson CA, et al. Mayo prognostic model for WHO-defined chronic myelomonocytic leukemia: ASXL1 and spliceosome component mutations and outcomes. Leukemia. 2013;27:1504–1510.
Patnaik MM, Itzykson R, Lasho T, Kosmider O, Finke C, Hanson C, et al. ASXL1 and SETBP1 mutations and their prognostic contribution in chronic myelomonocytic leukemia: a two-center study of 466 patients. Leukemia. 2014;28:2206–2212.
He R, Devine DJ, Tu ZJ, Mai M, Chen D, Nguyen PL, et al. Hybridization capture-based next generation sequencing reliably detects FLT3 mutations and classifies FLT3-internal tandem duplication allelic ratio in acute myeloid leukemia: a comparative study to standard fragment analysis. Mod Pathol. 2020;33:334–343.
Castaño-Díez S, López-Guerra M, Zugasti I, Calvo X, Schulz FI, Avendaño A, et al. AML typical mutations (CEBPA, FLT3, NPM1) identify a high-risk chronic myelomonocytic leukemia independent of CPSS molecular. Blood Adv. 2025;9:39–53.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Mehta N, He R, Viswanatha DS. Internal standardization of the interpretation and reporting of sequence variants in hematologic neoplasms. Mol Diagn Ther. 2021;25:517–526.
McGowan-Jordan J, Hastings R, Moore S. ISCN 2020: an international system for human cytogenomic nomenclature (2020) Cytogenet. Genome Res. 2020;161:225–226.
Kröger N, Zabelina T, Guardiola P, Runde V, Sierra J, Van Biezen A, et al. Allogeneic stem cell transplantation of adult chronic myelomonocytic leukaemia. A report on behalf of the Chronic Leukaemia Working Party of the European Group for Blood and Marrow Transplantation (EBMT). Br J Haematol. 2002;118:67–73.
Gagelmann N, Badbaran A, Beelen DW, Salit RB, Stölzel F, Rautenberg C, et al. A prognostic score including mutation profile and clinical features for patients with CMML undergoing stem cell transplantation. Blood Adv. 2021;5:1760–1769.
Pophali P, Matin A, Mangaonkar AA, Carr R, Binder M, Al-Kali A, et al. Prognostic impact and timing considerations for allogeneic hematopoietic stem cell transplantation in chronic myelomonocytic leukemia. Blood Cancer J. 2020;10:121.
Carratt SA, Braun TP, Coblentz C, Schonrock Z, Callahan R, Curtiss BM, et al. Mutant SETBP1 enhances NRAS-driven MAPK pathway activation to promote aggressive leukemia. Leukemia. 2021;35:3594–3599.
Bera R, Chiu MC, Huang YJ, Lin TH, Kuo MC, Shih LY. RUNX1 mutations promote leukemogenesis of myeloid malignancies in ASXL1-mutated leukemia. J Hematol Oncol. 2019;12:104.
Montalban-Bravo G, Rodriguez-Sevilla JJ, Swanson DM, Kanagal-Shamanna R, Hammond D, Chien K, et al. Influence of co-mutational patterns in disease phenotype and clinical outcomes of chronic myelomonocytic leukemia. Leukemia. 2024;38:1178–1181.
Patnaik MM, Lasho TL, Vijayvargiya P, Finke CM, Hanson CA, Ketterling RP, et al. Prognostic interaction between ASXL1 and TET2 mutations in chronic myelomonocytic leukemia. Blood Cancer J. 2016;6:e385.
Zhao W, Zhang C, Li Y, Li Y, Liu Y, Sun X, et al. The prognostic value of the interaction between ASXL1 and TET2 gene mutations in patients with chronic myelomonocytic leukemia: a meta-analysis. Hematology. 2022;27:367–378.
You X, Liu F, Binder M, Vedder A, Lasho T, Wen Z, et al. Asxl1 loss cooperates with oncogenic Nras in mice to reprogram the immune microenvironment and drive leukemic transformation. Blood. 2022;139:1066–1079.
Noushin Farnoud KM, Guglielmelli P, et al. Identification of ultra high-risk genotypes for progression to blast phase disease redefines molecular-based stratification of myelofibrosis. European Hematology Association, EHA Library, 2025;4160895:PS1820.
Yoshimi A, Lin K-T, Wiseman DH, Rahman MA, Pastore A, Wang B, et al. Coordinated alterations in RNA splicing and epigenetic regulation drive leukaemogenesis. Nature. 2019;574:273–277.
Peng J, Zuo Z, Fu B, Oki Y, Tang G, Goswami M, et al. Chronic myelomonocytic leukemia with nucleophosmin (NPM1) mutation. Eur J Haematol. 2016;96:65–71.
Author information
Authors and Affiliations
Contributions
SF, MY, PF, AA, CC, MN, MGDP, LL, AC, NF, RR, MG and AT were involved in study design, and gathering of data; LR, SF and AT conducted the data analysis; AAM, AP, MMP, LL, MG and NG participated in patient care; SF, LR and AT wrote the paper. All authors approved the final script.
Corresponding author
Ethics declarations
Competing interests
SF: Nothing to disclose; LR: Nothing to disclose; MY: Nothing to disclose; PF: Nothing to disclose; AA: Nothing to disclose; CC: Nothing to disclose; MN: Nothing to disclose; AAM: Research funding from Novartis, BMS, Solu Therapeutics and Sanofi; AP: Nothing to disclose; LL: Nothing to disclose; AC: Nothing to disclose; GM: Nothing to disclose; NF: Nothing to disclose; RR: Honoraria Incyte Corp.; MGDP: Nothing to disclose; NG: Advisory board to DISC Medicine and Agios; MMP: Research funding from Kura Oncology, Stemline therapeutics, Epigenetix, Solutherapeutics, Polaris and has served on the advisory board for AstraZeneca and SOBI pharmaceuticals, AT: Nothing to disclose; AT and NG are members of the editorial board for BCJ.
Ethics approval
This study was approved by the Institutional Review Board of the Mayo Clinic (IRB protocol number: 12-003574). All methods were performed in accordance with the Declaration of Helsinki, the relevant guidelines, and regulations.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fathima, S., Rokach, L., Yousuf, M. et al. CMML2AML: machine-learning discovery of co-mutations and specific single mutations predictive of blast transformation in chronic myelomonocytic leukemia. Blood Cancer J. (2026). https://doi.org/10.1038/s41408-026-01491-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41408-026-01491-1