Abstract
Background
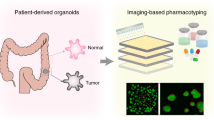
Colorectal cancer (CRC) is a substantial global health concern due to its limited treatment options, especially for oxaliplatin (L-OHP) regimen resistance. This study used organoid-based screening methodologies to evaluate drug responses in CRC while validating the approach with patient-derived CRC organoids and investigating potential biomarkers.
Methods
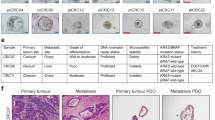
Patient-derived organoids were created from CRC surgical specimens, and drug screening were performed. Selected organoids with high and low L-OHP sensitivity underwent next-generation sequencing (NGS), and in vivo experiments using xenotransplantation were used to validate in vitro results. Moreover, the clinical application of homologous recombination deficiency (HRD) as a biomarker was investigated.
Results
Organoid drug screening revealed differences in L-OHP sensitivity among 34 patient-derived CRC organoids, and NGS deemed HRD as a potential biomarker. In vivo experiments validated the correlation between HRD status and L-OHP sensitivity, and clinical data suggested the potential of HRD as a biomarker for recurrence-free survival in patients treated with L-OHP. Additionally, HRD exhibited potential as a biomarker for other platinum agents and poly (ADP-ribose) polymerase inhibitors in CRC.
Conclusions
The study underscores HRD as a potential biomarker for predicting L-OHP sensitivity, expanding its application to other drugs in CRC. Organoid screening is reliable, providing insights into the intricate association between genetic features and treatment responses.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data used for the manuscript will be available from the corresponding author upon reasonable request.
References
Globocan Global Cancer Observatory, 2018, Cancer Today. Available from: https://gco.iarc.fr [Accessed 14.04.2024].
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Hong YS, Kim SY, Lee JS, Nam BH, Kim KP, Kim JE, et al. Oxaliplatin-based adjuvant chemotherapy for rectal cancer after preoperative chemoradiotherapy (ADORE): long-term results of a randomized controlled trial. J Clin Oncol. 2019;37:3111–23.
Suwa Y, Watanabe J, Suwa H, Ozawa M, Momiyama M, Ishibe A, et al. Exploratory randomized phase II trial for optimizing treatment dosage and duration of adjuvant S-1 plus oxaliplatin in patients with stage III colon cancer: YCOG1402 (SOAP trial). Ann Gastroenterol Surg. 2023;7:922–31.
Kelland L. The resurgence of platinum-based cancer chemotherapy. Nat Rev Cancer. 2007;7:573–84.
Zhou J, Zhou Y, Yin B, Hao W, Zhao L, Ju W, et al. 5-Fluorouracil and oxaliplatin modify the expression profiles of microRNAs in human colon cancer cells in vitro. Oncol Rep. 2010;23:121–8.
Chen ZH, Qi JJ, Wu QN, Lu JH, Liu ZX, Wang Y, et al. Eukaryotic initiation factor 4A2 promotes experimental metastasis and oxaliplatin resistance in colorectal cancer. J Exp Clin Cancer Res. 2019;38:196.
Cercek A, Dos Santos Fernandes G, Roxburgh CS, Ganesh K, Ng S, Sanchez-Vega F, et al. Mismatch repair-deficient rectal cancer and resistance to neoadjuvant chemotherapy. Clin Cancer Res. 2020;26:3271–9.
Ciccia A, Elledge SJ. The DNA damage response: making it safe to play with knives. Mol Cell. 2010;40:179–204.
Paulet L, Trecourt A, Leary A, Peron J, Descotes F, Devouassoux-Shisheboran M, et al. Cracking the homologous recombination deficiency code: how to identify responders to PARP inhibitors. Eur J Cancer. 2022;166:87–99.
Lord CJ, Ashworth A. BRCAness revisited. Nat Rev Cancer. 2016;16:110–20.
Kaufman B, Shapira-Frommer R, Schmutzler RK, Audeh MW, Friedlander M, Balmaña J, et al. Olaparib monotherapy in patients with advanced cancer and a germline BRCA1/2 mutation. J Clin Oncol. 2015;33:244–50.
Tutt A, Tovey H, Cheang MCU, Kernaghan S, Kilburn L, Gazinska P, et al. Carboplatin in BRCA1/2-mutated and triple-negative breast cancer BRCAness subgroups: the TNT Trial. Nat Med. 2018;24:628–37.
Moore K, Colombo N, Scambia G, Kim BG, Oaknin A, Friedlander M, et al. Maintenance olaparib in patients with newly diagnosed advanced ovarian cancer. N Engl J Med. 2018;379:2495–505.
Ray-Coquard I, Pautier P, Pignata S, Pérol D, González-Martín A, Berger R, et al. Olaparib plus bevacizumab as first-line maintenance in ovarian cancer. N Engl J Med. 2019;381:2416–28.
Oh M, McBride A, Yun S, Bhattacharjee S, Slack M, Martin JR, et al. BRCA1 and BRCA2 gene mutations and colorectal cancer risk: systematic review and meta-analysis. J Natl Cancer Inst. 2018;110:1178–89.
Soyano AE, Baldeo C, Kasi PM. BRCA mutation and its association with colorectal cancer. Clin Colorectal Cancer. 2018;17:e647–50.
Ogiri M, Seishima R, Nakamura K, Aimono E, Matsui S, Shigeta K, et al. Real-world application of next-generation sequencing-based test for surgically resectable colorectal cancer in clinical practice. Future Oncol. 2022;18:2701–11.
Clevers H. Modeling development and disease with organoids. Cell. 2016;165:1586–97.
Vlachogiannis G, Hedayat S, Vatsiou A, Jamin Y, Fernández-Mateos J, Khan K, et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science. 2018;359:920–6.
Ando K, Hu Q, Kasagi Y, Oki E, Mori M. Recent developments in cancer research: expectations for a new remedy. Ann Gastroenterol Surg. 2021;5:419–26.
Driehuis E, Kretzschmar K, Clevers H. Establishment of patient-derived cancer organoids for drug-screening applications. Nat Protoc. 2020;15:3380–409.
Fujii M, Matano M, Toshimitsu K, Takano A, Mikami Y, Nishikori S, et al. Human intestinal organoids maintain self-renewal capacity and cellular diversity in niche-inspired culture condition. Cell Stem Cell. 2018;23:787–93.e6.
Kawasaki K, Toshimitsu K, Matano M, Fujita M, Fujii M, Togasaki K, et al. An organoid biobank of neuroendocrine neoplasms enables genotype-phenotype mapping. Cell. 2020;183:1420–35.e21.
Fujii M, Shimokawa M, Date S, Takano A, Matano M, Nanki K, et al. A colorectal tumor organoid library demonstrates progressive loss of niche factor requirements during tumorigenesis. Cell Stem Cell. 2016;18:827–38.
Nakamura K, Aimono E, Oba J, Hayashi H, Tanishima S, Hayashida T, et al. Estimating copy number using next-generation sequencing to determine ERBB2 amplification status. Med Oncol. 2021;38:36.
Tsumura K, Arai E, Tian Y, Shibuya A, Nishihara H, Yotani T, et al. Establishment of permutation for cancer risk estimation in the urothelium based on genome-wide DNA methylation analysis. Carcinogenesis. 2019;40:1308–19.
Hilton JL, Geisler JP, Rathe JA, Hattermann-Zogg MA, DeYoung B, Buller RE. Inactivation of BRCA1 and BRCA2 in ovarian cancer. J Natl Cancer Inst. 2002;94:1396–406.
Loeffler CML, El Nahhas OSM, Muti HS, Carrero ZI, Seibel T, van Treeck M, et al. Prediction of homologous recombination deficiency from routine histology with attention-based multiple instance learning in nine different tumor types. BMC Biol. 2024;22:225.
Birkbak NJ, Wang ZC, Kim JY, Eklund AC, Li Q, Tian R, et al. Telomeric allelic imbalance indicates defective DNA repair and sensitivity to DNA-damaging agents. Cancer Discov. 2012;2:366–75.
Popova T, Manié E, Rieunier G, Caux-Moncoutier V, Tirapo C, Dubois T, et al. Ploidy and large-scale genomic instability consistently identify basal-like breast carcinomas with BRCA1/2 inactivation. Cancer Res. 2012;72:5454–62.
Pennington KP, Walsh T, Harrell MI, Lee MK, Pennil CC, Rendi MH, et al. Germline and somatic mutations in homologous recombination genes predict platinum response and survival in ovarian, fallopian tube, and peritoneal carcinomas. Clin Cancer Res. 2014;20:764–75.
Wen H, Feng Z, Ma Y, Liu R, Ou Q, Guo Q, et al. Homologous recombination deficiency in diverse cancer types and its correlation with platinum chemotherapy efficiency in ovarian cancer. BMC Cancer. 2022;22:550.
Zhao EY, Shen Y, Pleasance E, Kasaian K, Leelakumari S, Jones M, et al. Homologous recombination deficiency and platinum-based therapy outcomes in advanced breast cancer. Clin Cancer Res. 2017;23:7521–30.
Do Canto LM, Larsen SJ, Catin Kupper BE, Begnami M, Scapulatempo-Neto C, Petersen AH, et al. Increased levels of genomic instability and mutations in homologous recombination genes in locally advanced rectal carcinomas. Front Oncol. 2019;9:395.
Tsang ES, Csizmok V, Williamson LM, Pleasance E, Topham JT, Karasinska JM, et al. Homologous recombination deficiency signatures in gastrointestinal and thoracic cancers correlate with platinum therapy duration. NPJ Precis Oncol. 2023;7:31.
Sundar R, Miranda S, Rodrigues DN, Chénard-Poirier M, Dolling D, Clarke M, et al. Ataxia telangiectasia mutated protein loss and benefit from oxaliplatin-based chemotherapy in colorectal cancer. Clin Colorectal Cancer. 2018;17:280–4.
O’Connor MJ. Targeting the DNA damage response in cancer. Mol Cell. 2015;60:547–60.
Mansoori B, Mohammadi A, Davudian S, Shirjang S, Baradaran B. The different mechanisms of cancer drug resistance: a brief review. Adv Pharm Bull. 2017;7:339–48.
Ciardiello F, Ciardiello D, Martini G, Napolitano S, Tabernero J, Cervantes A. Clinical management of metastatic colorectal cancer in the era of precision medicine. CA Cancer J Clin. 2022;72:372–401.
Ghiringhelli F, Richard C, Chevrier S, Végran F, Boidot R. Efficiency of olaparib in colorectal cancer patients with an alteration of the homologous repair protein. World J Gastroenterol. 2016;22:10680–6.
Arena S, Corti G, Durinikova E, Montone M, Reilly NM, Russo M, et al. A subset of colorectal cancers with cross-sensitivity to olaparib and oxaliplatin. Clin Cancer Res. 2020;26:1372–84.
Funding
This work was supported by JST SPRING (Grant Number JPMJSP2123) and Sysmex Corporation (Hyogo, Japan).
Author information
Authors and Affiliations
Contributions
Yujin K, RS, KH and HI carried out experiments. Yujin K, HK and KN collected the samples. Yujin K, ES, KS, KO, TT, HN, HS and Yuko K analysed the data. Yujin K and RS conceived experiments and wrote the paper. All authors had final approval of the submitted and published versions.
Corresponding author
Ethics declarations
Competing interests
Yuko K received grants from TAIHO PHARMACEUTICAL CO., LTD.; CHUGAI PHARMACEUTICAL CO., LTD.; Yakult Honsha Co. Ltd.; DAIICHI SANKYO COMPANY; Merck Serono Co., Ltd; AsahiKASEI Co., Ltd; EA Pharma Co., Ltd.; Otsuka Pharmaceutical Co., Ltd.; Takeda Pharmaceutical Co., Ltd.; Otsuka Pharmaceutical Factory Inc.; SHIONOGI & CO., LTD.; KAKEN PHARMACEUTICAL CO., LTD.; Kowa Pharmaceutical Co., Ltd.; Astellas Pharma Inc.; MEDICON INC., Dainippon Sumitomo Pharma Co., Ltd.; Taisho Toyama Pharmaceutical Co., Ltd.; Kyouwa Hakkou Kirin Co., Ltd.; Pfizer Japan Inc.; ONO PHARMACEUTICAL CO., LTD.; NIHON PHARMACEUTICAL CO., LTD.; Japan Blood Products Organization; Medtronic Japan Co., Ltd.; Sanofi K.K.; Eisai Co., Ltd; TSUMURA & CO.; KCI Licensing, Inc.; ABBOTT JAPAN CO., LTD; and FUJIFILM Toyama Chemical Co., Ltd., outside the submitted work.
Ethics approval and consent to participate
The protocol of this study was approved by The Ethics Committee of Keio University, and all methods were performed in accordance with the Helsinki Declaration of 1996 (approval number: 20150148). The Ethics Committee at the Laboratory Animal Care and Use Committee at Keio University School of Medicine approved all experimental procedures, and the study was performed in accordance with the Care and Use of Laboratory Animals (NIH). Informed consent was obtained from all individual participants included in the study.
Consent for publication
All authors critically reviewed and approved the final manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kato, Y., Seishima, R., Hattori, K. et al. Significance of homologous recombinant deficiency as a biomarker for drug sensitivity in colorectal cancer. Br J Cancer 132, 533–542 (2025). https://doi.org/10.1038/s41416-025-02950-5
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41416-025-02950-5