Abstract
Background
To develop and test a relation knowledge distillation three-dimensional residual network (RKD-R3D) model for predicting breast cancer molecular subtypes using ultrasound (US) videos to aid clinical personalized management.
Methods
This multicentre study retrospectively included 882 breast cancer patients (2375 US videos and 9499 images) between January 2017 and December 2021, which was divided into training, validation, and internal test cohorts. Additionally, 86 patients was collected between May 2023 and November 2023 as the external test cohort. St. Gallen molecular subtypes (luminal A, luminal B, HER2-positive, and triple-negative) were confirmed via postoperative immunohistochemistry. The RKD-R3D based on US videos was developed and validated to predict four-classification molecular subtypes of breast cancer. The predictive performance of RKD-R3D was compared with RKD-R2D, traditional R3D, and preoperative core needle biopsy (CNB). The area under the receiver operating characteristic curve (AUC), sensitivity, specificity, accuracy, balanced accuracy, precision, recall, and F1-score were analyzed.
Results
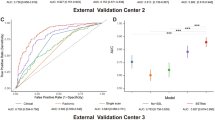
RKD-R3D (AUC: 0.88, 0.95) outperformed RKD-R2D (AUC: 0.72, 0.85) and traditional R3D (AUC: 0.65, 0.79) in predicting four-classification breast cancer molecular subtypes in the internal and external test cohorts. RKD-R3D outperformed CNB (Accuracy: 0.87 vs. 0.79) in the external test cohort, achieved good performance in predicting triple negative from non-triple negative breast cancers (AUC: 0.98), and obtained satisfactory prediction performance for both T1 and non-T1 lesions (AUC: 0.96, 0.90).
Conclusions
RKD-R3D when used with US videos becomes a potential supplementary tool to non-invasively assess breast cancer molecular subtypes.
This is a preview of subscription content, access via your institution
Access options






Similar content being viewed by others
Data availability
Some or all data, models, or code generated or used during the study are available from the corresponding author by request.
Code availability
Some or all data, models, or code generated or used during the study are available from the corresponding author by request.
References
Zardavas D, Irrthum A, Swanton C, Piccart M. Clinical management of breast cancer heterogeneity. Nat Rev Clin Oncol. 2015;12:381–94.
Goldhirsch A, Wood WC, Coates AS, Gelber RD, Thürlimann B, Senn HJ. Strategies for subtypes-dealing with the diversity of breast cancer: highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol. 2011;22:1736–47.
Zhou BY, Wang LF, Yin HH, Wu TF, Ren TT, Peng C, et al. Decoding the molecular subtypes of breast cancer seen on multimodal ultrasound images using an assembled convolutional neural network model: a prospective and multicentre study. eBioMedicine. 2021;74:103684.
Yeo SK, Guan JL. Breast cancer: multiple subtypes within a tumor?. Trends Cancer. 2017;3:753–60.
Wang M, He X, Chang Y, Sun G, Thabane L. A sensitivity and specificity comparison of fine needle aspiration cytology and core needle biopsy in evaluation of suspicious breast lesions: a systematic review and meta-analysis. Breast. 2017;31:157–66.
Zhu JY, He HL, Jiang XC, Bao HW, Chen F. Multimodal ultrasound features of breast cancers: correlation with molecular subtypes. BMC Med Imaging. 2023;23:57.
Zhang L, Li J, Xiao Y, Cui H, Du G, Wang Y, et al. Identifying ultrasound and clinical features of breast cancer molecular subtypes by ensemble decision. Sci Rep. 2015;5:11085.
Yang S, Gao X, Liu L, Shu R, Yan J, Zhang G, et al. Performance and reading time of automated breast US with or without computer-aided detection. Radiology. 2019;292:540–9.
Li X, Zhang S, Zhang Q, Wei X, Pan Y, Zhao J, et al. Diagnosis of thyroid cancer using deep convolutional neural network models applied to sonographic images: a retrospective, multicohort, diagnostic study. Lancet Oncol. 2019;20:193–201.
Trepanier C, Huang A, Liu M, Ha R. Emerging uses of artificial intelligence in breast and axillary ultrasound. Clin Imaging. 2023;100:64–68.
Cuocolo R, Caruso M, Perillo T, Ugga L, Petretta M. Machine Learning in oncology: a clinical appraisal. Cancer Lett. 2020;481:55–62.
Jiang M, Zhang D, Tang SC, Luo XM, Chuan ZR, Lv WZ, et al. Deep learning with convolutional neural network in the assessment of breast cancer molecular subtypes based on US images: a multicenter retrospective study. Eur Radiol. 2021;31:3673–82.
Zhang X, Li H, Wang C, Cheng W, Zhu Y, Li D, et al. Evaluating the accuracy of breast cancer and molecular subtype diagnosis by ultrasound image deep learning model. Front Oncol. 2021;11:623506.
Zhao G, Kong D, Xu X, Hu S, Li Z, Tian J. Deep learning-based classification of breast lesions using dynamic ultrasound video. Eur J Radiol. 2023;165:110885.
Qiu S, Zhuang S, Li B, Wang J, Zhuang Z. Prospective assessment of breast lesions AI classification model based on ultrasound dynamic videos and ACR BI-RADS characteristics. Front Oncol. 2023;13:1274557.
Meng H, Lin Z, Yang F, Xu Y, Cui L. Knowledge distillation in medical data mining: a survey. 5th international conference on crowd science and engineering. ACM Digital Library. 2021:175–82.
Xie X, Niu J, Liu X, Chen Z, Tang S, Yu S. A survey on incorporating domain knowledge into deep learning for medical image analysis. Med Image Anal. 2021;69:101985.
Canny J. A computational approach to edge detection. IEEE Trans Pattern Anal Mach Intell. 1986;8:679–98.
Kataoka H, Wakamiya T, Hara K, Satoh Y. Would mega-scale datasets further enhance spatiotemporal 3D CNNs? arXiv: 2004.04968 [Preprint] 2020. Available from https://arxiv.org/abs/2004.04968.
Park W, Kim D, Lu Y, Cho M. Relational knowledge distillation. 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR). IEEE. 2019: 3962–71.
Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D. Grad-cam: visual explanations from deep networks via gradient-based localization. 2017 IEEE International Conference on Computer Vision (ICCV). IEEE. 2017:618–26.
Tian Y, Krishnan D, Isola P. Contrastive representation distillation. arXiv: 1910.10699 [Preprint] 2019. Available from: https://arxiv.org/abs/1910.10699.
Passalis N, Tefas A. Learning deep representations with probabilistic knowledge transfer. European Conference on Computer Vision. Cham: Springer International Publishing. 2018:283–99.
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). IEEE. 2016: 770–8.
Luo WQ, Huang QX, Huang XW, Hu HT, Zeng FQ, Wang W. Predicting breast cancer in breast imaging reporting and data system (BI-RADS) ultrasound category 4 or 5 lesions: a nomogram combining radiomics and BI-RADS. Sci Rep. 2019;9:11921.
Zhang Y, Chen JH, Lin Y, Chan S, Zhou J, Chow D, et al. Prediction of breast cancer molecular subtypes on DCE-MRI using convolutional neural network with transfer learning between two centers. Eur Radiol. 2021;31:2559–67.
Mao N, Zhang H, Dai Y, Li Q, Lin F, Gao J, et al. Attention-based deep learning for breast lesions classification on contrast enhanced spectral mammography: a multicentre study. Br J Cancer. 2023;128:793–804.
de Margerie-Mellon C, Chassagnon G. Artificial intelligence: a critical review of applications for lung nodule and lung cancer. Diagn Interv Imaging. 2023;104:11–17.
Raja H, Akram MU, Shaukat A, Khan SA, Alghamdi N, Khawaja SG, et al. Extraction of retinal layers through convolution neural network (CNN) in an OCT image for glaucoma diagnosis. J Digital Imaging. 2020;33:1428–42.
Musthafa MM, RM T, VK V, Guluwadi S. Enhanced skin cancer diagnosis using optimized CNN architecture and checkpoints for automated dermatological lesion classification. BMC Med Imaging. 2024;24:201.
Xu C, Coen-Pirani P, Jiang X. Empirical study of overfitting in deep learning for predicting breast cancer metastasis. Cancers. 2023;15:1969.
Zhang T, Tan T, Han L, Appelman L, Veltman J, Wessels R, et al. Predicting breast cancer types on and beyond molecular level in a multi-modal fashion. NPJ Breast Cancer. 2023;9:16.
Wang F, Casalino LP, Khullar D. Deep learning in medicine-promise, progress, and challenges. JAMA Intern Med. 2019;179:293–4.
Tan PH, Ellis I, Allison K, Brogi E, Fox SB, Lakhani S, et al. The 2019 World Health Organization classification of tumours of the breast. Histopathology. 2020;77:181–5.
Pölcher M, Braun M, Tischitz M, Hamann M, Szeterlak N, Kriegmair A, et al. Concordance of the molecular subtype classification between core needle biopsy and surgical specimen in primary breast cancer. Arch Gynecol Obstet. 2021;304:783–90.
Acknowledgements
We thank the Harbin Medical University Cancer Hospital for the data support. We thank all those who helped us during the writing of this research.
Funding
Key Research and Development Project of Vanguard and Leading Goose in Zhejiang Province grant Nos. 2024C03069 (SLT) Medical Science and Technology Project of Zhejiang Province grant Nos. 2025KY564 (WYN) National Natural Science Foundation of China grant Nos. 82371984 (TJW).
Author information
Authors and Affiliations
Contributions
YNW: Conceptualization, Data curation, Formal analysis, Investigation, Writing - original draft, Supervision, Writing - review & editing. LZ: Data curation, Investigation, Writing- original draft. JZ: Data curation, Resources. YQP: Data curation, Software, Validation, Visualization; XYL: Formal analysis. YTW: Data curation, Software, Validation, Visualization. STZ: Software, Validation, Visualization. CJH: Formal analysis. PD: Formal analysis. LL: Data curation, Resources. YW: Data curation, Resources; JWT: Funding acquisition, Supervision, Writing - review & editing. LTS: Methodology, Project administration, Supervision, Writing - review & editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study protocol was centrally approved by the institutional Clinical Research Ethics Committee of the Second Affiliated Hospital of Harbin Medical University (approval number: KY2016-127), which conforms to the ethical guidelines of the 1975 Declaration of Helsinki. As this was a retrospective cohort study and individually identifiable information was removed during retrospective collection, informed consent was waived.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, Y., Zhou, L., Zhao, J. et al. Relation knowledge distillation 3D-ResNet-based deep learning for breast cancer molecular subtypes prediction on ultrasound videos: a multicenter study. Br J Cancer 133, 1178–1188 (2025). https://doi.org/10.1038/s41416-025-03146-7
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41416-025-03146-7
This article is cited by
-
Optimizing YOLOv11 for automated classification of breast cancer in medical images
Scientific Reports (2025)