Abstract
Background
This study aims to unravel the underlying mechanisms of osteosarcoma (OS) metastasis by single-cell RNA-sequencing.
Methods
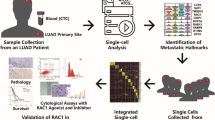
Ten pediatrics OS samples were analyzed using scRNA-seq and divided into three groups: primary tumor without lung metastasis (Pri_non_MT), primary tumor with lung metastasis (Pri_MT), and lung metastasis site (MT). Candidate genes associated with metastasis were identified by bioinformatics analysis and was confirmed at mRNA and protein levels in OS tissues. The function of candidate gene was identified in vitro and in vivo. Bulk RNA sequencing was used to explore downstream mechanism after candidate gene silenced.
Results
A total of 17 cell clusters were identified. ENO1 was selected as the candidate gene and was significantly expressed in Pri_MT and MT groups. In OS tissues, ENO1 significantly overexpressed in patients with lung metastasis compared to those without. Knocking down ENO1 resulted in a marked decrease in migration and invasion in vitro and a reduction in lung metastasis in vivo. Additionally, ENO1 suppression resulted in shifting the primary ATP production pathway from glycolysis to oxidative phosphorylation.
Conclusions
Our findings highlight ENO1 as a key regulator of glycolysis and metastasis in osteosarcoma, offering a novel therapeutic target for OS treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The analyzed data sets generated during the present study are available from the corresponding author on reasonable request.
References
Corre I, Verrecchia F, Crenn V, Redini F, Trichet V The Osteosarcoma Microenvironment: A Complex But Targetable Ecosystem. Cells. 2020;9:976.
Lindsey BA, Markel JE, Kleinerman ES. Osteosarcoma Overview. Rheumatol Ther. 2017;4:25–43.
Ahmed G, Zamzam M, Kamel A, Ahmed S, Salama A, Zaki I, et al. Effect of timing of pulmonary metastasis occurrence on the outcome of metastasectomy in osteosarcoma patients. J Pediatr Surg. 2019;54:775–9.
Yang C, Tian Y, Zhao F, Chen Z, Su P, Li Y, et al. Bone Microenvironment and Osteosarcoma Metastasis. Int J Mol Sci. 2020;21:6985.
Cui J, Dean D, Hornicek FJ, Chen Z, Duan Z. The role of extracelluar matrix in osteosarcoma progression and metastasis. J Exp Clin Cancer Res. 2020;39:178.
Long Z, Sun C, Tang M, Wang Y, Ma J, Yu J, et al. Single-cell multiomics analysis reveals regulatory programs in clear cell renal cell carcinoma. Cell Discov. 2022;8:68.
Kuchina A, Brettner LM, Paleologu L, Roco CM, Rosenberg AB, Carignano A, et al. Microbial single-cell RNA sequencing by split-pool barcoding. Science. 2021;371:eaba5257.
He S, Wang J, Huang Y, Kong F, Yang R, Zhan Y, et al. Intestinal fibrosis in aganglionic segment of Hirschsprung’s disease revealed by single-cell RNA sequencing. Clin Transl Med. 2023;13:e1193.
Zhou Y, Yang D, Yang Q, Lv X, Huang W, Zhou Z, et al. Single-cell RNA landscape of intratumoral heterogeneity and immunosuppressive microenvironment in advanced osteosarcoma. Nat Commun. 2020;11:6322.
Wang L, Huang X, You X, Yi T, Lu B, Liu J, et al. Nanoparticle enhanced combination therapy for stem-like progenitors defined by single-cell transcriptomics in chemotherapy-resistant osteosarcoma. Signal Transduct Target Ther. 2020;5:196.
Liu Y, Feng W, Dai Y, Bao M, Yuan Z, He M, et al. Single-Cell Transcriptomics Reveals the Complexity of the Tumor Microenvironment of Treatment-Naive Osteosarcoma. Front Oncol. 2021;11:709210.
Huang X, Wang L, Guo H, Zhang W, Shao Z. Single-cell transcriptomics reveals the regulative roles of cancer associated fibroblasts in tumor immune microenvironment of recurrent osteosarcoma. Theranostics. 2022;12:5877–87.
Guo J, Tang H, Huang P, Guo J, Shi Y, Yuan C, et al. Single-Cell Profiling of Tumor Microenvironment Heterogeneity in Osteosarcoma Identifies a Highly Invasive Subcluster for Predicting Prognosis. Front Oncol. 2022;12:732862.
Mereu E, Lafzi A, Moutinho C, Ziegenhain C, McCarthy DJ, Álvarez-Varela A, et al. Benchmarking single-cell RNA-sequencing protocols for cell atlas projects. Nat Biotechnol. 2020;38:747–55.
Hao Y, Hao S, Andersen-Nissen E, Mauck WM 3rd, Zheng S, Butler A, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184:3573–87.e29.
Wolock SL, Lopez R, Klein AM. Scrublet: Computational Identification of Cell Doublets in Single-Cell Transcriptomic Data. Cell Syst. 2019;8:281–91.e9.
Patel AP, Tirosh I, Trombetta JJ, Shalek AK, Gillespie SM, Wakimoto H, et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science. 2014;344:1396–401.
Gaujoux R, Seoighe C. A flexible R package for nonnegative matrix factorization. BMC Bioinforma. 2010;11:367.
Wu Y, Yang S, Ma J, Chen Z, Song G, Rao D, et al. Spatiotemporal Immune Landscape of Colorectal Cancer Liver Metastasis at Single-Cell Level. Cancer Discov. 2022;12:134–53.
Yin H, Gao T, Xie J, Huang Z, Zhang X, Yang F, et al. FUBP1 promotes colorectal cancer stemness and metastasis via DVL1-mediated activation of Wnt/β-catenin signaling. Mol Oncol. 2021;15:3490–512.
Niini T, Lahti L, Michelacci F, Ninomiya S, Hattinger CM, Guled M, et al. Array comparative genomic hybridization reveals frequent alterations of G1/S checkpoint genes in undifferentiated pleomorphic sarcoma of bone. Genes Chromosomes Cancer. 2011;50:291–306.
Liu W, Wang R, Zhang Y, Wang H, Huang Z, Jin T, et al. Whole-exome sequencing in osteosarcoma with distinct prognosis reveals disparate genetic heterogeneity. Cancer Genet. 2021;256-257:149–57.
Sayles LC, Breese MR, Koehne AL, Leung SG, Lee AG, Liu HY, et al. Genome-Informed Targeted Therapy for Osteosarcoma. Cancer Discov. 2019;9:46–63.
Atiye J, Wolf M, Kaur S, Monni O, Böhling T, Kivioja A, et al. Gene amplifications in osteosarcoma-CGH microarray analysis. Genes Chromosomes Cancer. 2005;42:158–63.
Chu Y, Dai E, Li Y, Han G, Pei G, Ingram DR, et al. Pan-cancer T cell atlas links a cellular stress response state to immunotherapy resistance. Nat Med. 2023;29:1550–62.
Engblom C, Pfirschke C, Pittet MJ. The role of myeloid cells in cancer therapies. Nat Rev Cancer. 2016;16:447–62.
Gentles AJ, Newman AM, Liu CL, Bratman SV, Feng W, Kim D, et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat Med. 2015;21:938–45.
Cui C, Chakraborty K, Tang XA, Zhou G, Schoenfelt KQ, Becker KM, et al. Neutrophil elastase selectively kills cancer cells and attenuates tumorigenesis. Cell. 2021;184:3163–77.e21.
Wang L, Liu Y, Dai Y, Tang X, Yin T, Wang C, et al. Single-cell RNA-seq analysis reveals BHLHE40-driven pro-tumour neutrophils with hyperactivated glycolysis in pancreatic tumour microenvironment. Gut. 2023;72:958–71.
Rahman I, Collado Sánchez A, Davies J, Rzeniewicz K, Abukscem S, Joachim J, et al. L-selectin regulates human neutrophil transendothelial migration. J Cell Sci. 2021;134:jcs250340.
Xue R, Zhang Q, Cao Q, Kong R, Xiang X, Liu H, et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature. 2022;612:141–7.
Ribatti D. Mast cells as therapeutic target in cancer. Eur J Pharm. 2016;778:152–7.
Marone G, Varricchi G, Loffredo S, Granata F. Mast cells and basophils in inflammatory and tumor angiogenesis and lymphangiogenesis. Eur J Pharm. 2016;778:146–51.
Cheng S, Li Z, Gao R, Xing B, Gao Y, Yang Y, et al. A pan-cancer single-cell transcriptional atlas of tumor infiltrating myeloid cells. Cell. 2021;184:792–809.e23.
Mellman I. Dendritic cells: master regulators of the immune response. Cancer Immunol Res. 2013;1:145–9.
Zhang Q, He Y, Luo N, Patel SJ, Han Y, Gao R, et al. Landscape and Dynamics of Single Immune Cells in Hepatocellular Carcinoma. Cell. 2019;179:829–45.e20.
Burr ML, Sparbier CE, Chan YC, Williamson JC, Woods K, Beavis PA, et al. CMTM6 maintains the expression of PD-L1 and regulates anti-tumour immunity. Nature. 2017;549:101–5.
Jessa S, Blanchet-Cohen A, Krug B, Vladoiu M, Coutelier M, Faury D, et al. Stalled developmental programs at the root of pediatric brain tumors. Nat Genet. 2019;51:1702–13.
Yuan Q, Yu H, Chen J, Song X, Sun L. Knockdown of pyruvate kinase type M2 suppresses tumor survival and invasion in osteosarcoma cells both in vitro and in vivo. Exp Cell Res. 2018;362:209–16.
Li Z, Geng M, Ye X, Ji Y, Li Y, Zhang X, et al. IRF7 inhibits the Warburg effect via transcriptional suppression of PKM2 in osteosarcoma. Int J Biol Sci. 2022;18:30–42.
Feng Z, Ou Y, Hao L. The roles of glycolysis in osteosarcoma. Front Pharm. 2022;13:950886.
Zhang J, Li H, Miao L, Ding J. Silencing of ENO1 inhibits the proliferation, migration and invasion of human breast cancer cells. J buon. 2020;25:696–701.
Li J, Hu ZQ, Yu SY, Mao L, Zhou ZJ, Wang PC, et al. CircRPN2 Inhibits Aerobic Glycolysis and Metastasis in Hepatocellular Carcinoma. Cancer Res. 2022;82:1055–69.
Li HJ, Ke FY, Lin CC, Lu MY, Kuo YH, Wang YP, et al. ENO1 Promotes Lung Cancer Metastasis via HGFR and WNT Signaling-Driven Epithelial-to-Mesenchymal Transition. Cancer Res. 2021;81:4094–109.
Wei ZQ, Ding S, Yang YC. TYROBP-positive endothelial cell-derived TWEAK as a promoter of osteosarcoma progression: insights from single-cell omics. Front Oncol. 2023;13:1200203.
Jiang Y, Ning Y, Cheng S, Huang Y, Deng M, Chen C. Single-cell aggrephagy-related patterns facilitate tumor microenvironment intercellular communication, influencing osteosarcoma progression and prognosis. Apoptosis. 2024;29:521–35.
Cheng D, Zhang Z, Mi Z, Tao W, Liu D, Fu J, et al. Deciphering the heterogeneity and immunosuppressive function of regulatory T cells in osteosarcoma using single-cell RNA transcriptome. Comput Biol Med. 2023;165:107417.
Bian X, Jiang H, Meng Y, Li YP, Fang J, Lu Z. Regulation of gene expression by glycolytic and gluconeogenic enzymes. Trends Cell Biol. 2022;32:786–99.
Chen K, Zhu C, Cai M, Fu D, Cheng B, Cai Z, et al. Integrative metabolome and transcriptome profiling reveals discordant glycolysis process between osteosarcoma and normal osteoblastic cells. J Cancer Res Clin Oncol. 2014;140:1715–21.
Huang CK, Sun Y, Lv L, Ping Y. ENO1 and Cancer. Mol Ther Oncolytics. 2022;24:288–98.
Luo M, Wu S, Ma Y, Liang H, Luo Y, Gu W, et al. Evaluating a Panel of Autoantibodies Against Tumor-Associated Antigens in Human Osteosarcoma. Front Genet. 2022;13:872253.
Nishizawa K, Nishiyama H, Matsui Y, Kobayashi T, Saito R, Kotani H, et al. Thioredoxin-interacting protein suppresses bladder carcinogenesis. Carcinogenesis. 2011;32:1459–66.
Kamitori K, Yamaguchi F, Dong Y, Hossain A, Katagi A, Noguchi C, et al. Both Ser361 phosphorylation and the C-arrestin domain of thioredoxin interacting protein are important for cell cycle blockade at the G1/S checkpoint. FEBS Open Bio. 2018;8:1804–19.
Deng J, Pan T, Liu Z, McCarthy C, Vicencio JM, Cao L, et al. The role of TXNIP in cancer: a fine balance between redox, metabolic, and immunological tumor control. Br J Cancer. 2023;129:1877–92.
Shen S, Yao T, Xu Y, Zhang D, Fan S, Ma J. CircECE1 activates energy metabolism in osteosarcoma by stabilizing c-Myc. Mol Cancer. 2020;19:151.
Qiao G, Wu A, Chen X, Tian Y, Lin X. Enolase 1, a Moonlighting Protein, as a Potential Target for Cancer Treatment. Int J Biol Sci. 2021;17:3981–92.
Vaupel P. Metabolic microenvironment of tumor cells: a key factor in malignant progression. Exp Oncol. 2010;32:125–7.
Reina-Campos M, Moscat J, Diaz-Meco M. Metabolism shapes the tumor microenvironment. Curr Opin Cell Biol. 2017;48:47–53.
Paul S, Ghosh S, Kumar S. Tumor glycolysis, an essential sweet tooth of tumor cells. Semin Cancer Biol. 2022;86:1216–30.
Acknowledgements
This work was supported by the Medical Research Data Center of Fudan University.
Funding
This study was supported by the National Natural Science Foundation of China (project approval No. 82272441). Science and Technology Commission of Shanghai Municipality (CN). Grant No. 22Y11912200.
Author information
Authors and Affiliations
Contributions
PH and SWH participated in the design of the study and performed the data analysis, PH, ZQ and YN carried out the in vitro and vivo studies. JFW participated in the data analysis. ZCL and YYM performed part of the experiments. RD, GC and BN conceived of the study, provided the funding, and participated in its design and coordination and helped to draft the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study protocol was approved by Children’s Hospital of Fudan University (2022-202). All experiments were performed incompliance with the relevant regulations, and all patients provided written informed consent. Besides, the animal studies were approved by the Animal Ethics Committee of Children’s Hospital of Fudan University (00036). The experiments followed the Guidelines for the Care and Use of Laboratory Animals issued by the Chinese Council on Animal Research.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, P., He, S., Qi, Z. et al. ENO1 promotes cancer metastasis via stimulating metabolism reprogramming in osteosarcoma. Br J Cancer 133, 1625–1639 (2025). https://doi.org/10.1038/s41416-025-03182-3
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41416-025-03182-3