Abstract
Background
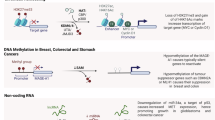
Despite significant advances in diagnosis and therapy, the prognosis of late-stage lung adenocarcinoma (LUAD) remains poor, underscoring the urgent need for effective biomarkers to enable early detection. Epigenetic alterations, particularly DNA methylation, is essential for controlling gene expression, and its abnormality plays critical roles in promoting carcinogenesis.
Methods
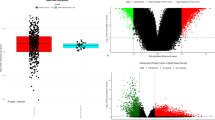
Reduced representation bisulfite sequencing (RRBS) technique was used to establish the DNA methylation profile in early-stage LUAD in comparison to normal tissues. The epigenetic abnormalities and expression, as well as the functions of ArfGAP with coiled-coil, ankyrin repeat and PH domains 3 (ACAP3) in LUAD carcinogenesis were further investigated.
Results
we investigated the DNA methylation dysregulation during tumorigenesis in the early-stage LUAD, and identified that Myc-mediated DNA hypermethylation and deacetylation as key mechanisms suppressing ACAP3 expression. ACAP3 significantly suppresses the proliferation of LUAD cells in vitro and in vivo. Mechanically, ACAP3 inhibits epidermal growth factor receptor (EGFR) signalling via impairing EGFR recycling and accelerating lysosome-mediated EGFR degradation in a GTPase-activating protein (GAP) activity-dependent manner.
Conclusion
Our finding reveals that ACAP3, suppressed by Myc-mediated epigenetic abnormality in early-stage LUAD, acts as a tumour suppressor by inhibiting EGFR signalling and cells proliferation, suggesting its potential as a diagnostic and therapeutic target.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The datasets generated in the current study are available from the corresponding authors on reasonable request.
References
Leiter A, Veluswamy RR, Wisnivesky JP. The global burden of lung cancer: current status and future trends. Nat Rev Clin Oncol. 2023;20:624–39.
Seguin L, Durandy M, Feral CC. Lung adenocarcinoma tumor origin: a guide for personalized medicine. Cancers. 2022;14:1759.
Thai AA, Solomon BJ, Sequist LV, Gainor JF, Heist RS. Lung cancer. Lancet. 2021;398:535–54.
Kim D, Lee YS, Kim DH, Bae SC. Lung cancer staging and associated genetic and epigenetic events. Mol Cells. 2020;43:1–9.
Zubair T, Bandyopadhyay D. Small molecule EGFR inhibitors as anti-cancer agents: discovery, mechanisms of action, and opportunities. Int J Mol Sci. 2023;24:2651.
Guo H, Zhou C, Zheng M, Zhang J, Wu H, He Q, et al. Insights into the role of derailed endocytic trafficking pathway in cancer: from the perspective of cancer hallmarks. Pharmacol Res. 2024;201:107084.
Zhang Y. Targeting Epidermal growth factor receptor for cancer treatment: abolishing both Kinase-dependent and kinase-independent functions of the receptor. Pharmacol Rev. 2023;75:1218–32.
Banushi B, Joseph SR, Lum B, Lee JJ, Simpson F. Endocytosis in cancer and cancer therapy. Nat Rev Cancer. 2023;23:450–73.
Duruisseaux M, Esteller M. Lung cancer epigenetics: from knowledge to applications. Semin Cancer Biol. 2018;51:116–28.
Lin RK, Hsu HS, Chang JW, Chen CY, Chen JT, Wang YC. Alteration of DNA methyltransferases contributes to 5’CpG methylation and poor prognosis in lung cancer. Lung Cancer. 2007;55:205–13.
Al-Yozbaki M, Jabre I, Syed NH, Wilson CM. Targeting DNA methyltransferases in non-small-cell lung cancer. Semin Cancer Biol. 2022;83:77–87.
Dong Z, Li J, Dai W, Yu D, Zhao Y, Liu S, et al. RRP15 deficiency induces ribosome stress to inhibit colorectal cancer proliferation and metastasis via LZTS2-mediated beta-catenin suppression. Cell Death Dis. 2023;14:89.
Hao Q, Bai Y, Guan R, Dong R, Bai W, Hamdy H, et al. VPS35/retromer-dependent MT1-MMP regulation confers melanoma metastasis. Sci China Life Sci. 2025;68:1996–2009.
Chen M, Zhou Y, Fu Z, Wu C. Transcription factor occupancy limits DNA methylation and determines ICAM1 expression in breast cancer. Acta Biochim Biophys Sin. 2025;57:818–33.
Zhang S, Liu J, Li F, Yang M, Wang J. EZH2 suppresses insulinoma development by epigenetically reducing KIF4A expression via H3K27me3 modification. Gene. 2022;822:146317.
Bai W, Yan C, Yang Y, Sang L, Hao Q, Yao X, et al. EGF/EGFR-YAP1/TEAD2 signaling upregulates STIM1 in vemurafenib resistant melanoma cells. FEBS J. 2024;291:4969–83.
Dong Z, Zhu C, Zhan Q, Jiang W. Cdk phosphorylation licenses Kif4A chromosome localization required for early mitotic progression. J Mol Cell Biol. 2018;10:358–70.
Zheng Z, Zhao Y, Yu H, Wang T, Li J, Xu L, et al. Suppressing MTERF3 inhibits proliferation of human hepatocellular carcinoma via ROS-mediated p38 MAPK activation. Commun Biol. 2024;7:18.
Liu SH, Shao FG, Wang YR, Zhang YR, Yu HJ, Zhang NX et al. COX6C expression driven by copy amplification of 8q22.2 regulates cell proliferation via mediation of mitosis by ROS-AMPK signaling in lung adenocarcinoma. Cell Death Dis. 2014;15:74.
Chen PH, Bendris N, Hsiao YJ, Reis CR, Mettlen M, Chen HY, et al. Crosstalk between CLCb/Dyn1-mediated adaptive clathrin-mediated endocytosis and epidermal growth factor receptor signaling increases metastasis. Dev Cell. 2017;40:278–88.
Yu JX, Feng HR, Sang QQ, Li FY, Chen MD, Yu BQ, et al. VPS35 promotes cell proliferation via EGFR recycling and enhances EGFR inhibitors response in gastric cancer. Ebiomedicine. 2023;89:104451.
Zhang R, Liu S, Gong B, Xie W, Zhao Y, Xu L, et al. Kif4A mediates resistance to neoadjuvant chemoradiotherapy in patients with advanced colorectal cancer via regulating DNA damage response. Acta Biochim Biophys Sin. 2022;54:940–51.
Jia Y, Li L, Li Y, Zhu X, Wang H, Xu B, et al. Iron overload mediates cytarabine resistance in AML by inhibiting the TP53 signaling pathway. Acta Biochim Biophys Sin. 2025;57:646–55.
Gopaldass N, Chen KE, Collins B, Mayer A. Assembly and fission of tubular carriers mediating protein sorting in endosomes. Nat Rev Mol Cell Biol. 2024;25:765–83.
Haglund K, Dikic I. The role of ubiquitylation in receptor endocytosis and endosomal sorting. J Cell Sci. 2012;125:265–75.
Perez Verdaguer M, Zhang T, Surve S, Paulo JA, Wallace C, Watkins SC, et al. Time-resolved proximity labeling of protein networks associated with ligand-activated EGFR. Cell Rep. 2022;39:110950.
Guo H, Wang J, Ren S, Zheng LF, Zhuang YX, Li DL, et al. Targeting EGFR-dependent tumors by disrupting an ARF6-mediated sorting system. Nat Commun. 2022;13:6004.
Sun D, Guo Y, Tang P, Li H, Chen L. Arf6 as a therapeutic target: structure, mechanism, and inhibitors. Acta Pharm Sin B. 2023;13:4089–104.
Miura Y, Hongu T, Yamauchi Y, Funakoshi Y, Katagiri N, Ohbayashi N, et al. ACAP3 regulates neurite outgrowth through its GAP activity specific to Arf6 in mouse hippocampal neurons. Biochem J. 2016;473:2591–602.
Dawson MA, Kouzarides T. Cancer epigenetics: from mechanism to therapy. Cell. 2012;150:12–27.
Wang Y, Wang J, Yang L, Qiu L, Hua Y, Wu S, et al. Epigenetic regulation of intestinal peptide transporter PEPT1 as a potential strategy for colorectal cancer sensitization. Cell Death Dis. 2021;12:532.
Smith ZD, Hetzel S, Meissner A. DNA methylation in mammalian development and disease. Nat Rev Genet. 2024;75:98–111.
Pommert L, Schafer ES, Malvar J, Gossai N, Florendo E, Pulakanti K, et al. Decitabine and vorinostat with FLAG chemotherapy in pediatric relapsed/refractory AML: report from the therapeutic advances in childhood leukemia and lymphoma (TACL) consortium. Am J Hematol. 2022;97:613–22.
Falchi L, Ma H, Klein S, Lue JK, Montanari F, Marchi E, et al. Combined oral 5-azacytidine and romidepsin are highly effective in patients with PTCL: a multicenter phase 2 study. Blood. 2021;137:2161–70.
Gaillard SL, Zahurak M, Sharma A, Durham JN, Reiss KA, Sartorius-Mergenthaler S, et al. A phase 1 trial of the oral DNA methyltransferase inhibitor CC-486 and the histone deacetylase inhibitor romidepsin in advanced solid tumors. Cancer. 2019;125:2837–45.
Zhan F, Zhang R, Qiu L, Ren Y. ACAP3 negatively regulated by HDAC2 inhibits the malignant development of papillary thyroid carcinoma cells. Int J Biochem Cell Biol. 2024;174:106635.
Jakobsen ST, Siersbaek R. Transcriptional regulation by MYC: an emerging new model. Oncogene. 2024;44:1–7.
Lourenco C, Resetca D, Redel C, Lin P, MacDonald AS, Ciaccio R, et al. MYC protein interactors in gene transcription and cancer. Nat Rev Cancer. 2021;21:579–91.
Dong Y, Tu R, Liu H, Qing G. Regulation of cancer cell metabolism: oncogenic MYC in the driver’s seat. Signal Transduct Target Ther. 2020;5:124.
Qiu X, Boufaied N, Hallal T, Feit A, de Polo A, Luoma AM, et al. MYC drives aggressive prostate cancer by disrupting transcriptional pause release at androgen receptor targets. Nat Commun. 2022;13:2559.
Feng YC, Liu XY, Teng L, Ji Q, Wu Y, Li JM, et al. c-Myc inactivation of p53 through the pan-cancer lncRNA MILIP drives cancer pathogenesis. Nat Commun. 2020;11:4980.
Zimmerli D, Brambillasca CS, Talens F, Bhin J, Linstra R, Romanens L, et al. MYC promotes immune-suppression in triple-negative breast cancer via inhibition of interferon signaling. Nat Commun. 2022;13:6579.
McFadden DG, Politi K, Bhutkar A, Chen FK, Song X, Pirun M, et al. Mutational landscape of EGFR-, MYC-, and Kras-driven genetically engineered mouse models of lung adenocarcinoma. Proc Natl Acad Sci USA. 2016;113:E6409–E6417.
Iwakawa R, Kohno T, Kato M, Shiraishi K, Tsuta K, Noguchi M, et al. MYC amplification as a prognostic marker of early-stage lung adenocarcinoma identified by whole genome copy number analysis. Clin Cancer Res. 2011;17:1481–9.
Seo AN, Yang JM, Kim H, Jheon S, Kim K, Lee CT, et al. Clinicopathologic and prognostic significance of c-MYC copy number gain in lung adenocarcinomas. Br J Cancer. 2014;110:2688–99.
Li C, Sun Y, Fang Z, Han X, Fang R, Zhang Y, et al. Comprehensive analysis of epidermal growth factor receptor gene status in lung adenocarcinoma. J Thorac Oncol. 2011;6:1016–21.
Okabe T, Okamoto I, Tamura K, Terashima M, Yoshida T, Satoh T, et al. Differential constitutive activation of the epidermal growth factor receptor in non-small cell lung cancer cells bearing EGFR gene mutation and amplification. Cancer Res. 2007;67:2046–53.
Devarakonda S, Morgensztern D, Govindan R. Genomic alterations in lung adenocarcinoma. Lancet Oncol. 2015;16:e342–351.
Roper N, Brown AL, Wei JS, Pack S, Trindade C, Kim C, et al. Clonal evolution and heterogeneity of osimertinib acquired resistance mechanisms in EGFR Mutant lung cancer. Cell Rep Med. 2020;1:100007.
Nukaga S, Yasuda H, Tsuchihara K, Hamamoto J, Masuzawa K, Kawada I, et al. Amplification of EGFR wild-type alleles in non-small cell lung cancer cells confers acquired resistance to mutation-selective EGFR tyrosine kinase inhibitors. Cancer Res. 2017;77:2078–89.
Blaquier JB, Ortiz-Cuaran S, Ricciuti B, Mezquita L, Cardona AF, Recondo G. Tackling osimertinib resistance in EGFR-mutant non-small cell lung cancer. Clin Cancer Res. 2023;29:3579–91.
Wu J, Gao Q, Xia Q, Wang Y, Zheng Z, He A, et al. Highly specific cytokine receptor-targeting chimeras for targeted membrane protein degradation and sensitization of Osimertinib in EGFR-mutated non-small-cell lung cancer. Adv Mater. 2025;37:e2504050.
Yao N, Wang CR, Liu MQ, Li YJ, Chen WM, Li ZQ, et al. Discovery of a novel EGFR ligand DPBA that degrades EGFR and suppresses EGFR-positive NSCLC growth. Signal Transduct Target Ther. 2020;5:214.
Miura Y, Kanaho Y. ACAP3, the GTPase-activating protein specific to the small GTPase Arf6, regulates neuronal migration in the developing cerebral cortex. Biochem Biophys Res Commun. 2017;493:1089–94.
Qi S, Su L, Li J, Zhang C, Ma Z, Liu G, et al. Arf6-driven endocytic recycling of CD147 determines HCC malignant phenotypes. J Exp Clin Cancer Res. 2019;38:471.
Morishige M, Hashimoto S, Ogawa E, Toda Y, Kotani H, Hirose M, et al. GEP100 links epidermal growth factor receptor signalling to Arf6 activation to induce breast cancer invasion. Nat Cell Biol. 2008;10:85–92.
Acknowledgements
We thank Haishan Huang and Yu Zhang for kindly supporting of materials.
Funding
This work was supported by National Natural Science Foundation of China (Grant No. 82402714), Zhejiang Provincial Natural Science Foundation of China (Grant No. ZCLMS25H2001), the Medical and Health Science and Technology Program of Zhejiang Province (Grant No. 2024KY1243), Key Clinical Specialty Construction Project of Zhejiang Province, Key Laboratory of Clinical Laboratory Diagnosis and Translational Research of Zhejiang Province (Grant No. 2022E10022).
Author information
Authors and Affiliations
Contributions
ZD: Methodology, Visualisation, Writing-Original draft preparation, Writing—Review & Editing, Supervision. WX: Methodology, Visualisation, Validation, Writing-Original draft preparation. NZ: Methodology, Investigation, Visualisation. YG: Methodology, Validation, Resources. YZ: Investigation, Resources. YL: Methodology, Investigation. CL: Validation, Conceptualisation, Writing—Review & Editing. FS: Resources, Funding acquisition, Writing—Review & Editing, Supervision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
Informed consent was obtained from all subjects involved in the study.
Ethics approval and consent to participate
All aspects of this study were approved by Institutional Research Ethics Committee of Wenzhou Medical University.
Informed consent
Informed consent was obtained from all subjects involved in the study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dong, Z., Xie, W., Zhang, N. et al. Myc-mediated epigenetic silencing of ACAP3 promotes lung adenocarcinoma proliferation via regulating EGFR dynamics. Br J Cancer 134, 860–873 (2026). https://doi.org/10.1038/s41416-025-03305-w
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41416-025-03305-w