Abstract
Background
TCF3::HLF-positive B-cell acute lymphoblastic leukemia (B-ALL) is a rare, highly aggressive subtype with historically poor outcomes. Despite its classification as a distinct entity, its clinical and molecular landscape remains poorly understood.
Methods
This study presents a single-center cohort of 34 TCF3::HLF-positive B-ALL patients, providing comprehensive clinical and molecular characterization by integrating clinical data, treatment responses, survival outcomes, whole-transcriptome sequencing (WTS), targeted sequencing, and flow cytometry.
Results
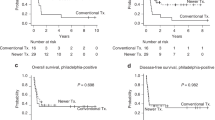
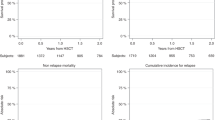
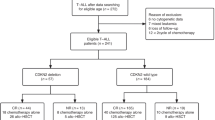
TCF3::HLF accounted for 1.59% of B-ALL cases. Three fusion isoforms were identified, with Isoform III likely arising from alternative splicing. No significant clinical or transcriptomic differences were observed between Isoform I and II. The 5-year overall survival (OS) was 35.2%. Allogeneic hematopoietic stem cell transplantation (allo-HSCT) significantly improved OS and event-free survival (p < 0.0001), while chimeric antigen receptor T-cell (CAR-T) therapy facilitated allo-HSCT but lacked durable efficacy. RAS pathway mutations were prevalent (85.7%), and CD33 expression was frequent (79.4%), suggesting potential therapeutic targets. WTS analysis revealed dysregulation of epithelial-mesenchymal transition, coagulation, and immune pathways.
Conclusions
TCF3::HLF-positive B-ALL represents an ultra-high-risk leukemia requiring allo-HSCT for long-term remission. CAR-T serves as a bridge to transplantation, while RAS and CD33-directed therapies warrant further investigation. These findings provide critical insights into disease biology and potential treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
For original data, please contact starliu@pku.edu.cn
References
Raimondi SC, Privitera E, Williams DL, Look AT, Behm F, Rivera GK, et al. New recurring chromosomal translocations in childhood acute lymphoblastic leukemia. Blood. 1991;77:2016–22.
Inaba T, Roberts WM, Shapiro LH, Jolly KW, Raimondi SC, Smith SD, et al. Fusion of the leucine zipper gene HLF to the E2A gene in human acute B-lineage leukemia. Science. 1992;257:531–4.
Hunger SP, Ohyashiki K, Toyama K, Cleary ML. Hlf, a novel hepatic bZIP protein, shows altered DNA-binding properties following fusion to E2A in t(17;19) acute lymphoblastic leukemia. Genes. 1992;6:1608–20.
Fischer U, Forster M, Rinaldi A, Risch T, Sungalee S, Warnatz HJ, et al. Genomics and drug profiling of fatal TCF3-HLF-positive acute lymphoblastic leukemia identifies recurrent mutation patterns and therapeutic options. Nat Genet. 2015;47:1020–9.
Tasian SK. TCF3::HLF acute lymphoblastic leukemia: still challenging to cure thirty years later. Haematologica. 2023;108:1713–4.
Alaggio R, Amador C, Anagnostopoulos I, Attygalle AD, Araujo IBO, Berti E, et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: lymphoid Neoplasms. Leukemia. 2022;36:1720–48.
Arber DA, Orazi A, Hasserjian RP, Borowitz MJ, Calvo KR, Kvasnicka HM, et al. International consensus classification of myeloid neoplasms and acute leukemias: integrating morphologic, clinical, and genomic data. Blood. 2022;140:1200–28.
Hirai M, Yagasaki H, Fujimura J, Inoue M, Shimozawa K, Okuma H, et al. Successful preemptive donor lymphocyte infusions from a haploidentical donor in a boy with E2A-HLF-positive ALL. Leuk Lymphoma. 2018;59:746–8.
Panagopoulos I, Micci F, Thorsen J, Haugom L, Tierens A, Ulvmoen A, et al. A novel TCF3-HLF fusion transcript in acute lymphoblastic leukemia with a t(17;19)(q22;p13). Cancer Genet. 2012;205:669–72.
Takeda R, Yokoyama K, Ogawa M, Kawamata T, Fukuyama T, Kondoh K, et al. The first case of elderly TCF3-HLF-positive B-cell acute lymphoblastic leukemia. Leuk Lymphoma. 2019;60:2821–4.
Ahmed MZ, Venkatadasari I, Dyer S, Wall K, Huxley E, Lovell R, et al. Clonal evolution in adult TCF3::HLF-positive acute lymphoblastic leukemia undergoing stem cell transplantation. Ann Hematol. 2022;101:2553–4.
Lejman M, Włodarczyk M, Zawitkowska J, Kowalczyk JR. Comprehensive chromosomal aberrations in a case of a patient with TCF3-HLF-positive BCP-ALL. BMC Med Genom. 2020;13:58.
Mouttet B, Vinti L, Ancliff P, Bodmer N, Brethon B, Cario G, et al. Durable remissions in TCF3-HLF positive acute lymphoblastic leukemia with blinatumomab and stem cell transplantation. Haematologica. 2019;104:e244–e247.
Zeckanovic A, Mouttet B, Vinti L, Ancliff P, Brethon B, Cario G, et al. Update on long-term outcomes of a cohort of patients with TCF3::HLF-positive acute lymphoblastic leukemia treated with blinatumomab and stem cell transplantation. Haematologica. 2025;110:1373–8.
Chen S, Li Y, Wang Z, Feng L, Jia Y, Mo X, et al. Improved outcomes in E2A::HLF positive B-cell acute lymphoblastic leukemia by chimeric antigen receptor T cell therapy and BCL-2 inhibitor. Chin Med J (Engl). 2023;136:1382–4.
Wang T, Wan X, Yang F, Shi W, Liu R, Ding L, et al. Successful treatment of TCF3-HLF-positive Childhood B-ALL with Chimeric Antigen Receptor T-Cell Therapy. Clin Lymphoma Myeloma Leuk. 2021;21:386–92.
Minson KA, Prasad P, Vear S, Borinstein S, Ho R, Domm J, et al. t(17;19) in children with acute lymphocytic leukemia: a report of 3 cases and a review of the literature. Case Rep Hematol. 2013;2013:563291.
Dahéron L, Brizard F, Millot F, Cividin M, Lacotte L, Guilhot F, et al. E2A/HLF fusion gene in an acute lymphoblastic leukemia patient with disseminated intravascular coagulation and a normal karyotype. Hematol J. 2002;3:153–6.
Glover JM, Loriaux M, Tyner JW, Druker BJ, Chang BH. In vitro sensitivity to dasatinib in lymphoblasts from a patient with t(17;19)(q22;p13) gene rearrangement pre-B acute lymphoblastic leukemia. Pediatr Blood Cancer. 2012;59:576–9.
Shimonodan H, Nagayama J, Nagatoshi Y, Hatanaka M, Takada A, Iguchi H, et al. Acute lymphocytic leukemia in adolescence with multiple osteolytic lesions and hypercalcemia mediated by lymphoblast-producing parathyroid hormone-related peptide: a case report and review of the literature. Pediatr Blood Cancer. 2005;45:333–9.
Zerkalenkova E, Menchits Y, Borkovskaia A, Sokolova S, Soldatkina O, Mikhailova E, et al. TCF3 gene rearrangements in pediatric B-cell acute lymphoblastic leukemia-A single center experience. Int J Lab Hematol. 2023;45:533–40.
Li MJ, Yu CH, Chou SW, Su YH, Liao KW, Chang HH, et al. TCF3-HLF-positive acute lymphoblastic leukemia resembling burkitt leukemia: cell morphologic and immunophenotypic findings. JCO Precis Oncol. 2022;6:e2200236.
Wu S, Lu J, Su D, Yang F, Zhang Y, Hu S. The advantage of chimeric antigen receptor T cell therapy in pediatric acute lymphoblastic leukemia with E2A-HLF fusion gene positivity: a case series. Transl Pediatr. 2021;10:686–91.
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127:2391–405.
Chen X, Wang F, Zhang Y, Wang M, Tian W, Teng W, et al. Retrospective analysis of 36 fusion genes in 2479 Chinese patients of de novo acute lymphoblastic leukemia. Leuk Res. 2018;72:99–104.
Chen X, Wang F, Zhang Y, Wang M, Tian W, Teng W, et al. Panoramic view of common fusion genes in a large cohort of Chinese de novo acute myeloid leukemia patients. Leuk Lymphoma. 2019;60:1071–8.
Chen X, Wang F, Zhang Y, Ma X, Cao P, Yuan L, et al. Fusion gene map of acute leukemia revealed by transcriptome sequencing of a consecutive cohort of 1000 cases in a single center. Blood Cancer J. 2021;11:112.
Chen X, Yuan L, Zhang Y, Wang F, Ma X, Fang J, et al. Advances towards genome-based acute myeloid leukemia classification: a comparative analysis of WHO-HAEM4R, WHO-HAEM5, and International Consensus Classification. Am J Hematol. 2024;99:824–35.
Marx V. Seeing data as t-SNE and UMAP do. Nat Methods. 2024;21:930–3.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7.
Zhang Y, Wang F, Chen X, Zhang Y, Wang M, Liu H, et al. CSF3R mutations are frequently associated with abnormalities of RUNX1, CBFB, CEBPA, and NPM1 genes in acute myeloid leukemia. Cancer. 2018;124:3329–38.
Zhang Y, Wang F, Chen X, Liu H, Wang X, Chen J, et al. Next-generation sequencing reveals the presence of DDX41 mutations in acute lymphoblastic leukemia and aplastic anemia. EJHaem. 2021;2:508–13.
McGowan-Jordan J, Hastings RJ, Moore S. ISCN 2020: an International System for Human Cytogenomic Nomenclature. Basel, Switzerland: Karger, 2020.
Zhang X, Lu XA, Yang J, Zhang G, Li J, Song L, et al. Efficacy and safety of anti-CD19 CAR T-cell therapy in 110 patients with B-cell acute lymphoblastic leukemia with high-risk features. Blood Adv. 2020;4:2325–38.
Beder T, Hansen BT, Hartmann AM, Zimmermann J, Amelunxen E, Wolgast N, et al. The gene expression classifier ALLCatchR identifies B-cell precursor ALL subtypes and underlying developmental trajectories across age. Hemasphere. 2023;7:e939.
Huang Y, Mouttet B, Warnatz HJ, Risch T, Rietmann F, Frommelt F, et al. The leukemogenic TCF3-HLF complex rewires enhancers driving cellular identity and self-renewal conferring EP300 vulnerability. Cancer Cell. 2019;36:630–44.
Priebe V, Drakul A, Galvan B, Aguadé-Gorgorió J, Mikkola HKA, Bornhauser B, et al. TCF3::HLF orchestrates an enhancer-promoter network with activation of MEF2C to promote immature HSC gene expression in leukemia. bioRxiv. 2024;11:625006.
de Boer J, Yeung J, Ellu J, Ramanujachar R, Bornhauser B, Solarska O, et al. The E2A-HLF oncogenic fusion protein acts through Lmo2 and Bcl-2 to immortalize hematopoietic progenitors. Leukemia. 2011;25:321–30.
Hirose K, Inukai T, Kikuchi J, Furukawa Y, Ikawa T, Kawamoto H, et al. Aberrant induction of LMO2 by the E2A-HLF chimeric transcription factor and its implication in leukemogenesis of B-precursor ALL with t(17;19). Blood. 2010;116:962–70.
Liu Y, Klein J, Bajpai R, Dong L, Tran Q, Kolekar P, et al. Etiology of oncogenic fusions in 5,190 childhood cancers and its clinical and therapeutic implication. Nat Commun. 2023;14:1739.
Roth DB V(D)J Recombination: Mechanism, Errors, and Fidelity. Microbiol Spectr. 2014;2:https://doi.org/10.1128/microbiolspec.MDNA3-0041-2014.
Nourse J, Mellentin JD, Galili N, Wilkinson J, Stanbridge E, Smith SD, et al. Chromosomal translocation t(1;19) results in synthesis of a homeobox fusion mRNA that codes for a potential chimeric transcription factor. Cell. 1990;60:535–45.
Hirabayashi S, Ohki K, Nakabayashi K, Ichikawa H, Momozawa Y, Okamura K, et al. ZNF384-related fusion genes define a subgroup of childhood B-cell precursor acute lymphoblastic leukemia with a characteristic immunotype. Haematologica. 2017;102:118–29.
Irving J, Matheson E, Minto L, Blair H, Case M, Halsey C, et al. Ras pathway mutations are prevalent in relapsed childhood acute lymphoblastic leukemia and confer sensitivity to MEK inhibition. Blood. 2014;124:3420–30.
Gamis AS, Alonzo TA, Meshinchi S, Sung L, Gerbing RB, Raimondi SC, et al. Gemtuzumab ozogamicin in children and adolescents with de novo acute myeloid leukemia improves event-free survival by reducing relapse risk: results from the randomized phase III Children’s Oncology Group trial AAML0531. J Clin Oncol. 2014;32:3021–32.
Laszlo GS, Gudgeon CJ, Harrington KH, Dell’Aringa J, Newhall KJ, Means GD, et al. Cellular determinants for preclinical activity of a novel CD33/CD3 bispecific T-cell engager (BiTE) antibody, AMG 330, against human AML. Blood. 2014;123:554–61.
Ravandi F, Subklewe M, Walter RB, Vachhani P, Ossenkoppele G, Buecklein V, et al. Safety and tolerability of AMG 330 in adults with relapsed/refractory AML: a phase 1a dose-escalation study. Leuk Lymphoma. 2024;65:1281–91.
Wan X, Wang T, Yang J, Yang X, Li W, Ding L, et al. Durable remission achieved in pediatric patients with TCF3-HLF-positive relapsed/refractory B-cell acute lymphoblastic leukemia by dual CD19 and CD22-targeted chimeric antigen receptor T-cell therapy. Haematologica. 2025; https://doi.org/10.3324/haematol.2024.287023. Epub ahead of print.
Lu W, Kang Y. Epithelial-mesenchymal plasticity in cancer progression and metastasis. Dev Cell. 2019;49:361–74.
Dudas J, Ladanyi A, Ingruber J, Steinbichler TB, Riechelmann H. Epithelial to mesenchymal transition: a mechanism that fuels cancer radio/chemoresistance. Cells. 2020;9:428.
Acknowledgements
The authors express their sincere gratitude to the patients and their families for their participation in this study. This work was supported by the Langfang Science and Technology Research and Development Program (Grant No. 2023013154).
Author information
Authors and Affiliations
Contributions
HL designed and supervised the study and revised the manuscript. XC and XM conducted comprehensive data analysis and drafted the article. LY, FW, YZ, and JC analyzed clinical data. JF, PC, XZ (Zhou), and SL performed molecular and bioinformatic analyses. TW conducted cytogenetic analysis. MX, JY, and XZ (Zhang) were responsible for patient management and provided clinical specimens. All authors reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, X., Ma, X., Yuan, L. et al. TCF3::HLF-positive B-ALL: integrated clinical and molecular characterization of 34 cases from a single-center cohort. Br J Cancer (2026). https://doi.org/10.1038/s41416-026-03370-9
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41416-026-03370-9