Abstract
Background
The gut microbiome is increasingly recognized as a modulator of cancer therapy outcomes and a potential predictive biomarker. This systematic review synthesizes current evidence on microbial biomarkers associated with neoadjuvant treatment (NT) response in rectal cancer (RC).
Methods
PubMed, Embase, and Ovid Medline databases were searched through March 2025. Eligible studies included RC patients treated with NT with baseline microbial analysis stratified by treatment response. Two reviewers independently performed screening, data extraction, and quality assessment (NIH and STORMS tools). Due to substantial heterogeneity, a structured qualitative synthesis without meta-analysis was conducted following SWiM guidelines, using a direction-of-effect vote-counting approach.
Results
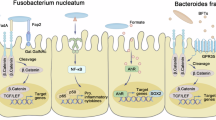
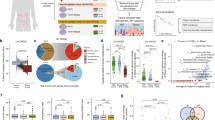
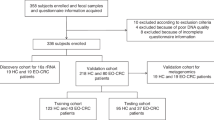
Sixteen observational studies (842 patients) were included, covering chemoradiotherapy (nCRT), total neoadjuvant therapy, chemotherapy, and immunochemoradiotherapy. Microbiota composition was investigated by 16S rRNA sequencing, metagenomics, or metatranscriptomics on fecal or tissue samples. While microbial diversity showed inconsistent associations, specific taxa -notably Bacteroides, Fusobacterium and Akkermansia- emerged as recurrent biomarkers of poor response to nCRT. Twelve predictive models reported AUROC values from 0.73 to 0.97, with limited external validation.
Conclusions
Specific microbial taxa show a consistent association with nCRT resistance across independent cohorts. However, methodological heterogeneity and limited reproducibility warrant standardized prospective validation before clinical implementation. PROSPERO: CRD42023433704.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The data used for this systematic review were extracted from articles that were all public and available from PubMed, Ovid MEDLINE and EMBASE.
References
Benson AB, Venook AP, Adam M, Chang G, Chen Y-J, Ciombor KK, et al. NCCN Guidelines® Insights: Rectal Cancer, Version 3.2024. J Natl Compr Canc Netw. 2024;22:366–75. https://doi.org/10.6004/jnccn.2024.0041.
van Gijn W, Marijnen CAM, Nagtegaal ID, Kranenbarg EM-K, Putter H, Wiggers T, et al. Preoperative radiotherapy combined with total mesorectal excision for resectable rectal cancer: 12-year follow-up of the multicentre, randomised controlled TME trial. Lancet Oncol. 2011;12:575–82. https://doi.org/10.1016/S1470-2045(11)70097-3.
Keller DS, Berho M, Perez RO, Wexner SD, Chand M. The multidisciplinary management of rectal cancer. Nat Rev Gastroenterol Hepatol. 2020;17:414–29. https://doi.org/10.1038/s41575-020-0275-y.
Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer. 2009;45:228–47. https://doi.org/10.1016/j.ejca.2008.10.026.
Rödel C, Hofheinz R, Fokas E. Rectal cancer: neoadjuvant chemoradiotherapy. Best Pract Res Clin Gastroenterol. 2016;30:629–39. https://doi.org/10.1016/j.bpg.2016.06.004.
van der Sluis FJ, Couwenberg AM, de Bock GH, Intven MPW, Reerink O, van Leeuwen BL, et al. Population-based study of morbidity risk associated with pathological complete response after chemoradiotherapy for rectal cancer. Br J Surg. 2020;107:131–9. https://doi.org/10.1002/bjs.11324.
Pucciarelli S, Del Bianco P, Efficace F, Serpentini S, Capirci C, De Paoli A, et al. Patient-reported outcomes after neoadjuvant chemoradiotherapy for rectal cancer: a multicenter prospective observational study. Ann Surg. 2011;253:71–7. https://doi.org/10.1097/SLA.0b013e3181fcb856.
Alexander JL, Wilson ID, Teare J, Marchesi JR, Nicholson JK, Kinross JM. Gut microbiota modulation of chemotherapy efficacy and toxicity. Nat Rev Gastroenterol Hepatol. 2017;14:356–65. https://doi.org/10.1038/nrgastro.2017.20.
Cheng WY, Wu C-Y, Yu J. The role of gut microbiota in cancer treatment: friend or foe? Gut. 2020;69:1867–76. https://doi.org/10.1136/gutjnl-2020-321153.
Wong CC, Yu J. Gut microbiota in colorectal cancer development and therapy. Nat Rev Clin Oncol. 2023;20:429–52. https://doi.org/10.1038/s41571-023-00766-x.
Chrysostomou D, Roberts LA, Marchesi JR, Kinross JM. Gut microbiota modulation of efficacy and toxicity of cancer chemotherapy and immunotherapy. Gastroenterology. 2023;164:198–213. https://doi.org/10.1053/j.gastro.2022.10.018.
Zhao R, Coker OO, Wu J, Zhou Y, Zhao L, Nakatsu G, et al. Aspirin reduces colorectal tumor development in mice and gut microbes reduce its bioavailability and chemopreventive effects. Gastroenterology. 2020;159:969–983.e4. https://doi.org/10.1053/j.gastro.2020.05.004.
Hou X-Y, Zhang P, Du H-Z, Gao Y-Q, Sun R-Q, Qin S-Y, et al. Prevotella contributes to individual response of FOLFOX in colon cancer. Clin Transl Med. 2021;11:e512. https://doi.org/10.1002/ctm2.512.
Zhang S, Yang Y, Weng W, Guo B, Cai G, Ma Y, et al. Fusobacterium nucleatum promotes chemoresistance to 5-fluorouracil by upregulation of BIRC3 expression in colorectal cancer. J Exp Clin Cancer Res. 2019;38:14 https://doi.org/10.1186/s13046-018-0985-y.
Yu T, Guo F, Yu Y, Sun T, Ma D, Han J, et al. Fusobacterium nucleatum promotes chemoresistance to colorectal cancer by modulating autophagy. Cell. 2017;170:548–563.e16. https://doi.org/10.1016/j.cell.2017.07.008.
Zheng D-W, Dong X, Pan P, Chen K-W, Fan J-X, Cheng S-X, et al. Phage-guided modulation of the gut microbiota of mouse models of colorectal cancer augments their responses to chemotherapy. Nat Biomed Eng. 2019;3:717–28. https://doi.org/10.1038/s41551-019-0423-2.
Roberti MP, Yonekura S, Duong CPM, Picard M, Ferrere G, Tidjani Alou M, et al. Chemotherapy-induced ileal crypt apoptosis and the ileal microbiome shape immunosurveillance and prognosis of proximal colon cancer. Nat Med. 2020;26:919–31. https://doi.org/10.1038/s41591-020-0882-8.
Iida N, Dzutsev A, Stewart CA, Smith L, Bouladoux N, Weingarten RA, et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science. 2013;342:967–70. https://doi.org/10.1126/science.1240527.
McQuade JL, Daniel CR, Helmink BA, Wargo JA. Modulating the microbiome to improve therapeutic response in cancer. Lancet Oncol. 2019;20:e77–91. https://doi.org/10.1016/S1470-2045(18)30952-5.
Tonneau M, Elkrief A, Pasquier D, Paz Del Socorro T, Chamaillard M, Bahig H, et al. The role of the gut microbiome on radiation therapy efficacy and gastrointestinal complications: a systematic review. Radiother Oncol. 2021;156:1–9. https://doi.org/10.1016/j.radonc.2020.10.033.
Liu J, Liu C, Yue J. Radiotherapy and the gut microbiome: facts and fiction. Radiat Oncol. 2021;16:9. https://doi.org/10.1186/s13014-020-01735-9.
Mima K, Nishihara R, Qian ZR, Cao Y, Sukawa Y, Nowak JA, et al. Fusobacterium nucleatum in colorectal carcinoma tissue and patient prognosis. Gut. 2016;65:1973–80. https://doi.org/10.1136/gutjnl-2015-310101.
Nakatsu G, Zhou H, Wu WKK, Wong SH, Coker OO, Dai Z, et al. Alterations in enteric virome are associated with colorectal cancer and survival outcomes. Gastroenterology. 2018;155:529–541.e5. https://doi.org/10.1053/j.gastro.2018.04.018.
Li X, Saxena D. The tumor mycobiome: a paradigm shift in cancer pathogenesis. Cell. 2022;185:3648–51. https://doi.org/10.1016/j.cell.2022.09.013.
Zitvogel L, Derosa L, Routy B, Loibl S, Heinzerling L, de Vries IJM, et al. Impact of the ONCOBIOME network in cancer microbiome research. Nat Med. 2025;31:1085–98. https://doi.org/10.1038/s41591-025-03608-8.
Schupack DA, Mars RAT, Voelker DH, Abeykoon JP, Kashyap PC. The promise of the gut microbiome as part of individualized treatment strategies. Nat Rev Gastroenterol Hepatol. 2022;19:7–25. https://doi.org/10.1038/s41575-021-00499-1.
Mirzayi C, Renson A, Genomic Standards Consortium, Massive Analysis and Quality Control Society, Zohra F, Elsafoury S, Geistlinger L, Kasselman LJ, et al. Reporting guidelines for human microbiome research: the STORMS checklist. Nat Med. 2021;27:1885–92. https://doi.org/10.1038/s41591-021-01552-x.
Jang B-S, Chang JH, Chie EK, Kim K, Park JW, Kim MJ, et al. Gut microbiome composition is associated with a pathologic response after preoperative chemoradiation in patients with rectal cancer. Int J Radiat Oncol Biol Phys. 2020;107:736–46. https://doi.org/10.1016/j.ijrobp.2020.04.015.
Shi W, Shen L, Zou W, Wang J, Yang J, Wang Y, et al. The gut microbiome is associated with therapeutic responses and toxicities of neoadjuvant chemoradiotherapy in rectal cancer patients-a pilot study. Front Cell Infect Microbiol. 2020;10:562463. https://doi.org/10.3389/fcimb.2020.562463.
Li J, Li J, Lyu N, Ma Y, Liu F, Feng Y, et al. Composition of fecal microbiota in low-set rectal cancer patients treated with FOLFOX. Ther Adv Chronic Dis. 2020;11:2040622320904293. https://doi.org/10.1177/2040622320904293
Yi Y, Shen L, Shi W, Xia F, Zhang H, Wang Y, et al. Gut microbiome components predict response to neoadjuvant chemoradiotherapy in patients with locally advanced rectal cancer: a prospective, longitudinal study. Clin Cancer Res. 2021;27:1329–40. https://doi.org/10.1158/1078-0432.CCR-20-3445.
Takenaka IKTM, Bartelli TF, Defelicibus A, Sendoya JM, Golubicki M, Robbio J, et al. Exome and tissue-associated microbiota as predictive markers of response to neoadjuvant treatment in locally advanced rectal cancer. Front Oncol. 2022;12:809441. https://doi.org/10.3389/fonc.2022.809441.
Teng H, Wang Y, Sui X, Fan J, Li S, Lei X, et al. Gut microbiota-mediated nucleotide synthesis attenuates the response to neoadjuvant chemoradiotherapy in rectal cancer. Cancer Cell. 2023;41:124–138.e6. https://doi.org/10.1016/j.ccell.2022.11.013.
Sun L, Qu J, Ke X, Zhang Y, Xu H, Lv N, et al. Interaction between intratumoral microbiota and tumor mediates the response of neoadjuvant therapy for rectal cancer. Front Microbiol. 2023;14:1229888. https://doi.org/10.3389/fmicb.2023.1229888.
Huang X, Chen C, Xie W, Zhou C, Tian X, Zhang Z, et al. Metagenomic analysis of intratumoral microbiome linking to response to neoadjuvant chemoradiotherapy in rectal cancer. Int J Radiat Oncol Biol Phys. 2023;S0360-3016(23)07591-0. https://doi.org/10.1016/j.ijrobp.2023.06.2515
White MG, Damania A, Alshenaifi J, Sahasrabhojane P, Peacock O, Losh J, et al. Young-onset rectal cancer: unique tumoral microbiome and correlation with response to neoadjuvant therapy. Ann Surg. 2023;278:538–48. https://doi.org/10.1097/SLA.0000000000006015.
Sun Y, Zhang X, Jin C, Yue K, Sheng D, Zhang T, et al. Prospective, longitudinal analysis of the gut microbiome in patients with locally advanced rectal cancer predicts response to neoadjuvant concurrent chemoradiotherapy. J Transl Med. 2023;21:221. https://doi.org/10.1186/s12967-023-04054-1.
Sulit AK, Wilson K, Pearson J, Silander OK, Sampurno S, Michael M, et al. Human gene and microbial analyses in rectal cancer complete responses to radiotherapy. BJS Open. 2023;7:zrad035. https://doi.org/10.1093/bjsopen/zrad035.
Zhou H, Wang L, Lin Z, Jiang C, Chen X, Wang K, et al. Methylglyoxal from gut microbes boosts radiosensitivity and radioimmunotherapy in rectal cancer by triggering endoplasmic reticulum stress and cGAS-STING activation. J Immunother Cancer. 2023;11:e007840 https://doi.org/10.1136/jitc-2023-007840.
Hein DM, Coughlin LA, Poulides N, Koh AY, Sanford NN. Assessment of distinct gut microbiome signatures in a diverse cohort of patients undergoing definitive treatment for rectal cancer. J Immunother Precis Oncol. 2024;7:150–8. https://doi.org/10.36401/JIPO-23-30.
Yang Z, Ma J, Han J, Li A, Liu G, Sun Y, et al. Gut microbiome model predicts response to neoadjuvant immunotherapy plus chemoradiotherapy in rectal cancer. Med. 2024;5:1293–1306.e4. https://doi.org/10.1016/j.medj.2024.07.002.
Roesel R, Strati F, Basso C, Epistolio S, Spina P, Djordjevic J, et al. Combined tumor-associated microbiome and immune gene expression profiling predict response to neoadjuvant chemo-radiotherapy in locally advanced rectal cancer. Oncoimmunology. 2025;14:2465015. https://doi.org/10.1080/2162402X.2025.2465015.
Lee HI, Jang B-S, Chang JH, Kim E, Lee TH, Park JH, et al. Relationships between the microbiome and response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer. Cancer Res Treat. 2024. https://doi.org/10.4143/crt.2024.521
Kokaine L, Gardovskis A, Gardovskis J. Evaluation and predictive factors of complete response in rectal cancer after neoadjuvant chemoradiation therapy. Medicina. 2021;57:1044 https://doi.org/10.3390/medicina57101044.
Roy S, Trinchieri G. Microbiota: a key orchestrator of cancer therapy. Nat Rev Cancer. 2017;17:271–85. https://doi.org/10.1038/nrc.2017.13.
Galeano Niño JL, Wu H, LaCourse KD, Kempchinsky AG, Baryiames A, Barber B, et al. Effect of the intratumoral microbiota on spatial and cellular heterogeneity in cancer. Nature. 2022;611:810–7. https://doi.org/10.1038/s41586-022-05435-0.
Hurst R, Brewer DS, Gihawi A, Wain J, Cooper CS. Cancer invasion and anaerobic bacteria: new insights into mechanisms. J Med Microbiol. 2024;73:001817 https://doi.org/10.1099/jmm.0.001817.
Li B, Wei Z, Wang Z, Xu F, Yang J, Lin B, et al. Fusobacterium nucleatum induces oxaliplatin resistance by inhibiting ferroptosis through E-cadherin/β-catenin/GPX4 axis in colorectal cancer. Free Radic Biol Med. 2024;220:125–38. https://doi.org/10.1016/j.freeradbiomed.2024.04.226.
Fan L, Xu C, Ge Q, Lin Y, Wong CC, Qi Y, et al. A. Muciniphila suppresses colorectal tumorigenesis by inducing TLR2/NLRP3-mediated M1-Like TAMs. Cancer Immunol Res. 2021;9:1111–24. https://doi.org/10.1158/2326-6066.CIR-20-1019.
Derosa L, Routy B, Thomas AM, Iebba V, Zalcman G, Friard S, et al. Intestinal Akkermansia muciniphila predicts clinical response to PD-1 blockade in advanced non-small cell lung cancer patients. Nat Med. 2022;28:315–24. https://doi.org/10.1038/s41591-021-01655-5.
Chen S, Fan L, Lin Y, Qi Y, Xu C, Ge Q, et al. Bifidobacterium adolescentis orchestrates CD143+ cancer-associated fibroblasts to suppress colorectal tumorigenesis by Wnt signaling-regulated GAS1. Cancer Commun. 2023;43:1027–47. https://doi.org/10.1002/cac2.12469.
Procaccianti G, Roggiani S, Conti G, Brigidi P, Turroni S, D’Amico F. Bifidobacterium in anticancer immunochemotherapy: friend or foe? Microbiome Res Rep. 2023;2:24 https://doi.org/10.20517/mrr.2023.23.
Li J, Chu R, Wang C, Li Y, Wu B, Wan J. Microbiome characteristics and Bifidobacterium longum in colorectal cancer patients pre- and post-chemotherapy. Transl Cancer Res. 2020;9:2178–90. https://doi.org/10.21037/tcr.2020.03.33.
Yang R, Liu W, Cai S, Feng X, Chen Y, Cheng X, et al. Evaluation of the efficacy of probiotics in the chemoradiotherapy of colorectal cancer: a meta-analysis of Randomized Controlled Trials. BMC Gastroenterol. 2025;25:312 https://doi.org/10.1186/s12876-025-03914-y.
Ugai S, Liu LI, Kosumi K, Kawamura H, Hamada T, Mima,K, et al. Long-term yogurt intake and colorectal cancer incidence subclassified by Bifidobacterium abundance in tumor. Gut Microbes. 2025;17:2452237. https://doi.org/10.1080/19490976.2025.2452237.
Little MS, Ervin SM, Walton WG, Tripathy A, Xu Y, Liu J, et al. Active site flexibility revealed in crystal structures of Parabacteroides merdae β-glucuronidase from the human gut microbiome. Protein Sci. 2018;27:2010–22. https://doi.org/10.1002/pro.3507.
Wallace BD, Wang H, Lane KT, Scott JE, Orans J, Koo JS, et al. Alleviating cancer drug toxicity by inhibiting a bacterial enzyme. Science. 2010;330:831–5. https://doi.org/10.1126/science.1191175.
Huo R-X, Wang Y-J, Hou S-B, Wang W, Zhang C-Z, Wan X-H. Gut mucosal microbiota profiles linked to colorectal cancer recurrence. World J Gastroenterol. 2022;28:1946–64. https://doi.org/10.3748/wjg.v28.i18.1946.
Weir TL, Manter DK, Sheflin AM, Barnett BA, Heuberger AL, Ryan EP. Stool microbiome and metabolome differences between colorectal cancer patients and healthy adults. PLoS ONE. 2013;8:e70803 https://doi.org/10.1371/journal.pone.0070803.
Tito RY, Verbandt S, Aguirre Vazquez M, Lahti L, Verspecht C, Lloréns-Rico V, et al. Microbiome confounders and quantitative profiling challenge predicted microbial targets in colorectal cancer development. Nat Med. 2024;30:1339–48. https://doi.org/10.1038/s41591-024-02963-2.
d’Humières C, Delavy M, Alla L, Ichou F, Gauliard E, Ghozlane A, et al. Perturbation and resilience of the gut microbiome up to 3 months after β-lactams exposure in healthy volunteers suggest an important role of microbial β-lactamases. Microbiome. 2024;12:50. https://doi.org/10.1186/s40168-023-01746-0.
Lam SY, Ioannou A, Konstanti P, Visseren T, Doukas M, Peppelenbosch MP, et al. Technical challenges regarding the use of formalin-fixed paraffin embedded (FFPE) tissue specimens for the detection of bacterial alterations in colorectal cancer. BMC Microbiol. 2021;21:297 https://doi.org/10.1186/s12866-021-02359-z.
Enderlin D, Bieri U, Gadient J, Morsy Y, Scharl M, Rüschoff JH, et al. Towards reliable methodology: microbiome analysis of fresh frozen vs. formalin-fixed paraffin-embedded bladder tissue samples: a feasibility study. Microorganisms. 2024;12:2594 https://doi.org/10.3390/microorganisms12122594.
Mukhopadhya I, Martin JC, Shaw S, McKinley AJ, Gratz SW, Scott KP. Comparison of microbial signatures between paired faecal and rectal biopsy samples from healthy volunteers using next-generation sequencing and culturomics. Microbiome. 2022;10:171. https://doi.org/10.1186/s40168-022-01354-4.
Sinha R, Abu-Ali G, Vogtmann E, Fodor AA, Ren B, Amir A, et al. Assessment of variation in microbial community amplicon sequencing by the Microbiome Quality Control (MBQC) project consortium. Nat Biotechnol. 2017;35:1077–86. https://doi.org/10.1038/nbt.3981.
Warmbrunn MV, Attaye I, Herrema H, Nieuwdorp M. Protocol standardization of microbiome studies—daunting but necessary. Gastroenterology. 2022;162:1822–4. https://doi.org/10.1053/j.gastro.2022.03.017.
Allaband C, Lingaraju A, Flores Ramos S, Kumar T, Javaheri H, Tiu MD, et al. Time of sample collection is critical for the replicability of microbiome analyses. Nat Metab. 2024;6:1282–93. https://doi.org/10.1038/s42255-024-01064-1.
Huttenhower C, Finn RD, McHardy AC. Challenges and opportunities in sharing microbiome data and analyses. Nat Microbiol. 2023;8:1960–70. https://doi.org/10.1038/s41564-023-01484-x.
Isali I, Wong TR, Tian S. Best practice guidelines for collecting microbiome samples in research studies. Eur Urol Focus. 2024;10:909–13. https://doi.org/10.1016/j.euf.2024.12.007.
Wu G, Xu T, Zhao N, Lam YY, Ding X, Wei D, et al. A core microbiome signature as an indicator of health. Cell. 2024;187:6550–6565.e11. https://doi.org/10.1016/j.cell.2024.09.019.
Zhou X, Shen X, Johnson JS, Spakowicz DJ, Agnello M, Zhou W, et al. Longitudinal profiling of the microbiome at four body sites reveals core stability and individualized dynamics during health and disease. Cell Host Microbe. 2024;32:506–526.e9. https://doi.org/10.1016/j.chom.2024.02.012.
Funding
This work was supported by AIRC under Grant IG 2019 - ID. 23381 – P.I. Scarpa Marco.
Author information
Authors and Affiliations
Consortia
Contributions
AS: data curation, investigation, visualization, writing-review and editing; AK, AR, IC: writing-review and editing; MS: funding acquisition, supervision, writing-review and editing; MeS: conceptualization, investigation, methodology, writing-original draft, writing-review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Stepanyan, A., Kotsafti, A., Rosato, A. et al. Gut microbiota-associated predictors as biomarkers of neoadjuvant treatment response in rectal cancer-a systematic review. Br J Cancer (2026). https://doi.org/10.1038/s41416-026-03443-9
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41416-026-03443-9