Abstract
Background
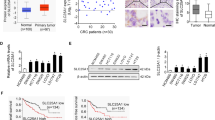
Colorectal cancer (CRC) is a highly vascularised tumour often characterised by elevated oxidative phosphorylation (OXPHOS) activity, positioning OXPHOS as a potential metabolic vulnerability for targeted therapy. SLIRP is an RNA-binding protein involved in the post-transcriptional regulation of mitochondrial gene expression. However, its specific function and underlying mechanism in CRC remain poorly understood.
Methods
Clinical specimens and public databases were utilised to analyse both the subcellular localisation and expression of SLIRP in CRC. The functional role of SLIRP in CRC progression was assessed through cell growth, apoptosis, and metabolic analyses. Post-transcriptional regulation of mitochondrial-encoded mRNAs by SLIRP was investigated using RNA immunoprecipitation and mRNA stability assays.
Results
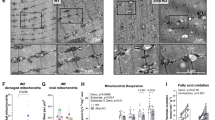
SLIRP expression was significantly elevated in CRC tissues compared to adjacent normal tissues, and high SLIRP expression correlated with poor patient survival. SLIRP knockdown induced an ATP crisis, leading to suppressed tumour growth and increased apoptosis in CRC cells. Mechanistically, SLIRP globally binds to mitochondrial-encoded mRNAs and maintains their stability, functioning as a key post-transcriptional regulator of mitochondrial gene expression.
Conclusions
These findings uncover a critical role for SLIRP in maintaining OXPHOS activity in CRC and highlight its potential as both a prognostic biomarker and a therapeutic metabolic target.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
References
Al Kamzari KAM & Constantinou C. Navigating the colorectal cancer maze: unveiling pathways to diagnosis, management, pathophysiology and prevention. Curr. Oncol. Rep. 2025 https://doi.org/10.1007/s11912-025-01707-w.
Teicher BA, Linehan WM, Helman LJ. Targeting cancer metabolism. Clin Cancer Res. 2012;18:5537–45.
Pang B, Wu H. Metabolic reprogramming in colorectal cancer: a review of aerobic glycolysis and its therapeutic implications for targeted treatment strategies. Cell Death Discov. 2025;11:321.
Qiu X, Wang A, Wang J, Zhang Z, Tao L. Mitochondrial metabolic reprogramming in colorectal cancer: mechanisms of resistance and future clinical interventions. Cell Death Discov. 2025;11:375.
Chao KSC, Chang HY, Huang WC, Yuan TT, Chen WTL, Ke TW, et al. Tilting immunometabolic balance of tumor microenvironment by targeting ENO1 in KRAS-mutated colorectal and pancreatic cancers. Int J Radiat Oncol, Biol, Phys. 2024;120:S97.
Molina JR, Sun Y, Protopopova M, Gera S, Bandi M, Bristow C, et al. An inhibitor of oxidative phosphorylation exploits cancer vulnerability. Nat Med. 2018;24:1036–46.
Yap TA, Daver N, Mahendra M, Zhang J, Kamiya-Matsuoka C, Meric-Bernstam F, et al. Complex I inhibitor of oxidative phosphorylation in advanced solid tumors and acute myeloid leukemia: phase I trials. Nat Med. 2023;29:115–26.
Vercellino I, Sazanov LA. The assembly, regulation and function of the mitochondrial respiratory chain. Nat Rev Mol Cell Biol. 2022;23:141–61.
Berner MJ, Wall SW, Echeverria GV. Deregulation of mitochondrial gene expression in cancer: mechanisms and therapeutic opportunities. Br J Cancer. 2024;131:1415–24.
Tan BG, Gustafsson CM, Falkenberg M. Mechanisms and regulation of human mitochondrial transcription. Nat Rev Mol Cell Biol. 2024;25:119–32.
Yan W, Xie C, Sun S, Zheng Q, Wang J, Wang Z, et al. SUCLG1 restricts POLRMT succinylation to enhance mitochondrial biogenesis and leukemia progression. EMBO J. 2024;43:2337–67.
Wang R, Chen B, Pan Y, Wang M, Xiao Y, Shi D, et al. POLRMT enhances lenvatinib resistance in hepatocellular carcinoma cells by maintaining mitochondrial ATP production. Life Sci. 2025;379:123876.
Wang H, Liu Y, Lu XS, Wu Y, Gu W, Yin G. Targeting POLRMT by IMT1 inhibits colorectal cancer cell growth. Cell Death Dis. 2024;15:643.
Li SP, Ou L, Zhang Y, Shen FR, Chen YG. A first-in-class POLRMT specific inhibitor IMT1 suppresses endometrial carcinoma cell growth. Cell Death Dis. 2023;14:152.
Bonekamp NA, Peter B, Hillen HS, Felser A, Bergbrede T, Choidas A, et al. Small-molecule inhibitors of human mitochondrial DNA transcription. Nature. 2020;588:712–6.
Li X, Ze X, Zhou S, Hu Z, He C, Jia Y, et al. Discovery of a novel, potent, orally active, and safe inhibitor targeting human mitochondrial RNA polymerase. J Med Chem. 2023;66:5118–53.
Schatton D, Rugarli EI. Post-transcriptional regulation of mitochondrial function. Curr Opin Physiol. 2018;3:6–15.
Reynaud K, Brothers M, Ly M, Ingolia NT. Dynamic post-transcriptional regulation by Mrn1 links cell wall homeostasis to mitochondrial structure and function. PLoS Genet. 2021;17:e1009521.
Chujo T, Ohira T, Sakaguchi Y, Goshima N, Nomura N, Nagao A, et al. LRPPRC/SLIRP suppresses PNPase-mediated mRNA decay and promotes polyadenylation in human mitochondria. Nucleic Acids Res. 2012;40:8033–47.
Singh V, Moran JC, Itoh Y, Soto IC, Fontanesi F, Couvillion M, et al. Structural basis of LRPPRC-SLIRP-dependent translation by the mitoribosome. Nat Struct Mol Biol. 2024;31:1838–47.
Spahr H, Rozanska A, Li X, Atanassov I, Lightowlers RN, Chrzanowska-Lightowlers ZM, et al. SLIRP stabilizes LRPPRC via an RRM-PPR protein interface. Nucleic Acids Res. 2016;44:6868–82.
Lagouge M, Mourier A, Lee HJ, Spahr H, Wai T, Kukat C, et al. SLIRP regulates the rate of mitochondrial protein synthesis and protects LRPPRC from degradation. PLoS Genet. 2015;11:e1005423.
Tang Z, Kang B, Li C, Chen T, Zhang Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019;47:W556–60.
Goldman MJ, Craft B, Hastie M, Repecka K, McDade F, Kamath A, et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat Biotechnol. 2020;38:675–8.
Lin A, Yang H, Shi Y, Cheng Q, Liu Z, Zhang J, et al. PanCanSurvPlot: a large-scale pan-cancer survival analysis web application. BioRxiv 2022 https://doi.org/10.1101/2022.12.25.521884.
Wang J, Song X, Wei M, Qin L, Zhu Q, Wang S, et al. PCAS: an integrated tool for multi-dimensional cancer research utilizing clinical proteomic tumor analysis consortium data. Int J Mol Sci 2024;25.
Zhang J, Zhang Q. Using Seahorse machine to measure OCR and ECAR in cancer cells. Methods Mol Biol. 2019;1928:353–63.
Caines JK, Barnes DA, Berry MD. The use of seahorse XF assays to interrogate real-time energy metabolism in cancer cell lines. Methods Mol Biol. 2022;2508:225–34.
Gou Q, Gao L, Nie X, Pu W, Zhu J, Wang Y, et al. Long noncoding RNA AB074169 inhibits cell proliferation via modulation of KHSRP-mediated CDKN1a expression in papillary thyroid carcinoma. Cancer Res. 2018;78:4163–74.
Ashton TM, McKenna WG, Kunz-Schughart LA, Higgins GS. Oxidative phosphorylation as an emerging target in cancer therapy. Clin Cancer Res. 2018;24:2482–90.
Bonnay F, Veloso A, Steinmann V, Kocher T, Abdusselamoglu MD, Bajaj S, et al. Oxidative metabolism drives immortalization of neural stem cells during tumorigenesis. Cell. 2020;182:1490–507.e1419.
El-Botty R, Morriset L, Montaudon E, Tariq Z, Schnitzler A, Bacci M, et al. Oxidative phosphorylation is a metabolic vulnerability of endocrine therapy and palbociclib resistant metastatic breast cancers. Nat Commun. 2023;14:4221.
Li K, Guo J, Ming Y, Chen S, Zhang T, Ma H, et al. A circular RNA activated by TGFbeta promotes tumor metastasis through enhancing IGF2BP3-mediated PDPN mRNA stability. Nat Commun. 2023;14:6876.
Chen S, Li K, Guo J, Chen HN, Ming Y, Jin Y, et al. circNEIL3 inhibits tumor metastasis through recruiting the E3 ubiquitin ligase Nedd4L to degrade YBX1. Proc Natl Acad Sci USA. 2023;120:e2215132120.
Guo J, Li K, Ming Y, Pan Y, Tan S, Ma H, et al. A circular RNA overcomes acquired resistance to BET inhibitors by antagonizing IGF2BP2-mediated c-MYC translation in TNBC. Proc Natl Acad Sci USA. 2025;122:e2504320122.
Sasarman F, Brunel-Guitton C, Antonicka H, Wai T, Shoubridge EA, Consortium L. LRPPRC and SLIRP interact in a ribonucleoprotein complex that regulates posttranscriptional gene expression in mitochondria. Mol Biol Cell. 2010;21:1315–23.
Yang M, Lin W, Huang J, Mannucci A, Luo H. Novel immunotherapeutic approaches in gastric cancer. Precis Clin Med. 2024;7:pbae020.
Pang H, Chen X, Yan M, Sun H. Dual anti-HER2 therapy combined with chemotherapy as a novel neoadjuvant treatment option for locally advanced HER2-positive and microsatellite stable colon cancer. Precis Clin Med. 2025;8:pbae033.
Tzeng SF, Yu YR, Park J, von Renesse J, Hsiao HW, Hsu CH, et al. PLT012, a humanized CD36-blocking antibody, is effective for unleashing antitumor immunity against liver cancer and liver metastasis. Cancer Discov. 2025;15:1676–96.
Veiga SR, Ge X, Mercer CA, Hernandez-Alvarez MI, Thomas HE, Hernandez-Losa J, et al. Phenformin-induced mitochondrial dysfunction sensitizes hepatocellular carcinoma for dual inhibition of mTOR. Clin Cancer Res. 2018;24:3767–80.
Schockel L, Glasauer A, Basit F, Bitschar K, Truong H, Erdmann G, et al. Targeting mitochondrial complex I using BAY 87-2243 reduces melanoma tumor growth. Cancer Metab. 2015;3:11.
Bendell JC, Patel MR, Infante JR, Kurkjian CD, Jones SF, Pant S, et al. Phase 1, open-label, dose escalation, safety, and pharmacokinetics study of ME-344 as a single agent in patients with refractory solid tumors. Cancer. 2015;121:1056–63.
Boland PM, Lenz HJ, Ciombor KK, Florou V, Pishvaian MJ, Cusnir M, et al. A Phase 1b study of the OxPhos inhibitor ME-344 with bevacizumab in refractory metastatic colorectal cancer. Invest N Drugs. 2025;43:60–8.
Daglish SCD, Fennell EMJ & Graves LM. Targeting mitochondrial DNA transcription by POLRMT inhibition or depletion as a potential strategy for cancer treatment. Biomedicines 2023;11.
Li X, Liu L, Feng D, Shi Y, Huang L, Yu M, et al. Design, optimization, and biological evaluation of a novel quinoline-based POLRMT inhibitor for prostate cancer therapy. J Med Chem 2025 https://doi.org/10.1021/acs.jmedchem.5c00130.
Mennuni M, Filograna R, Felser A, Bonekamp NA, Giavalisco P, Lytovchenko O, et al. Metabolic resistance to the inhibition of mitochondrial transcription revealed by CRISPR-Cas9 screen. EMBO Rep. 2022;23:e53054.
Giles RH, Lolkema MP, Snijckers CM, Belderbos M, van der Groep P, Mans DA, et al. Interplay between VHL/HIF1alpha and Wnt/beta-catenin pathways during colorectal tumorigenesis. Oncogene. 2006;25:3065–70.
Pham TCP, Raun SH, Havula E, Henriquez-Olguin C, Rubalcava-Gracia D, Frank E, et al. The mitochondrial mRNA-stabilizing protein SLIRP regulates skeletal muscle mitochondrial structure and respiration by exercise-recoverable mechanisms. Nat Commun. 2024;15:9826.
Fernando CD, Jayasekara WSN, Inampudi C, Kohonen-Corish MRJ, Cooper WA, Beilharz TH, et al. A STAT3 protein complex required for mitochondrial mRNA stability and cancer. Cell Rep. 2023;42:113033.
Bayona-Bafaluy MP, Sanchez-Cabo F, Fernandez-Silva P, Perez-Martos A, Enriquez JA. A genome-wide shRNA screen for new OxPhos-related genes. Mitochondrion. 2011;11:467–75.
Jourdain AA, Koppen M, Wydro M, Rodley CD, Lightowlers RN, Chrzanowska-Lightowlers ZM, et al. GRSF1 regulates RNA processing in mitochondrial RNA granules. Cell Metab. 2013;17:399–410.
Zhang X, Fryknas M, Hernlund E, Fayad W, De Milito A, Olofsson MH, et al. Induction of mitochondrial dysfunction as a strategy for targeting tumour cells in metabolically compromised microenvironments. Nat Commun. 2014;5:3295.
Vitiello GA, Medina BD, Zeng S, Bowler TG, Zhang JQ, Loo JK, et al. Mitochondrial inhibition augments the efficacy of Imatinib by resetting the metabolic phenotype of gastrointestinal stromal tumor. Clin Cancer Res. 2018;24:972–84.
Baughman JM, Nilsson R, Gohil VM, Arlow DH, Gauhar Z, Mootha VK. A computational screen for regulators of oxidative phosphorylation implicates SLIRP in mitochondrial RNA homeostasis. PLoS Genet. 2009;5:e1000590.
Hatchell EC, Colley SM, Beveridge DJ, Epis MR, Stuart LM, Giles KM, et al. SLIRP, a small SRA-binding protein, is a nuclear receptor corepressor. Mol Cell. 2006;22:657–68.
Li L, Miao W, Williams P, Guo C, Wang Y. SLIRP interacts with helicases to facilitate 2’-O-methylation of rRNA and to promote translation. J Am Chem Soc. 2019;141:10958–61.
Qi Z, Xue S, Chen J, Zhao W, Johnson K, Wen X, et al. Genome-wide mapping of RNA-protein associations through sequencing. Nat Biotechnol 2025 https://doi.org/10.1038/s41587-025-02780-z.
Acknowledgements
We thank Hao Wang (Laboratory of Dermatology, West China Hospital) for assistance with the mIHC assay; Yi Zhang and Yue Li (Core Facility, West China Hospital) for support with immunofluorescence; Selleck for providing antibodies against key energy metabolism-related proteins. Schematic diagrams were created using BioRender (https://BioRender.com).
Funding
This work was supported by the Science and Technology Foundation of Sichuan Province (2022NSFSC1296), National Natural Science Foundation of China (82102982), and the 1.3.5 Project for Disciplines of Excellence, West China Hospital, Sichuan University (ZYGD23027).
Author information
Authors and Affiliations
Contributions
J.W.G. and Y.P. designed experiments and supervised the study. C.Y.Y., J.W.G., Y.M., Z.X.Y., Y.Y.D., H.L.M., X.M.X., R.W., X.Y.Q., H.P.Y., and Y.Y. performed experiments. Q.B.W. collected clinical samples; C.Y.Y., J.W.G., and Y.P. analysed data and prepared the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All animal experiments were approved by the Ethics Committee of West China Hospital, Sichuan University (Approval no. 20250408004). Human CRC and matched adjacent normal tissues were obtained from West China Hospital, Sichuan University, with ethical approval (No. 2019(540)) and informed consent from all patients. All methods were performed in accordance with the relevant guidelines and regulations.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, C., Ming, Y., Wu, Q. et al. SLIRP maintains energy metabolism homeostasis in colorectal cancer by stabilizing mitochondrial-encoded mRNAs. Br J Cancer (2026). https://doi.org/10.1038/s41416-026-03453-7
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41416-026-03453-7