Abstract
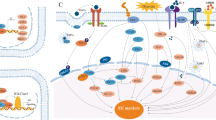
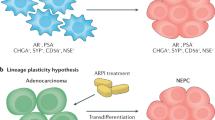
The treatment of prostate cancer (PCa) has made great progress in recent years, but treatment resistance always develops and can even lead to fatal disease. Exploring the mechanism of drug resistance is of great significance for improving treatment outcomes and developing biomarkers with predictive value. It is increasingly recognized that mechanism of drug resistance in advanced PCa is related to lineage plasticity and tissue differentiation. Specifically, one of the mechanisms by which castration-resistant prostate cancer (CRPC) cells acquire drug resistance and transform into neuroendocrine prostate cancer (NEPC) cells is lineage plasticity. NEPC is a subtype of PCa that is highly aggressive and lethal, with a median survival of only 7 months. With the development of high-throughput RNA sequencing technology, more and more non-coding RNAs have been identified, which play important roles in different diseases through different mechanisms. Several ncRNAs have shown great potential in PCa lineage plasticity and as biomarkers. In the review, the role of ncRNA in PCa lineage plasticity and its use as biomarkers were reviewed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Chen R, Dong X, Gleave M. Molecular model for neuroendocrine prostate cancer progression. BJU Int. 2018;122:560–70.
Crawford ED, Stanton W, Mandair D. Darolutamide: an evidenced-based review of its efficacy and safety in the treatment of prostate cancer. Cancer Manag Res. 2020;12:5667–76.
Chen Y, Zhou Q, Hankey W, Fang X, Yuan F. Second generation androgen receptor antagonists and challenges in prostate cancer treatment. Cell Death Dis. 2022;13:632.
Parimi V, Goyal R, Poropatich K, Yang XJ. Neuroendocrine differentiation of prostate cancer: a review. Am J Clin Exp Urol. 2014;2:273–85.
Beltran H, Prandi D, Mosquera JM, Benelli M, Puca L, Cyrta J, et al. Divergent clonal evolution of castration-resistant neuroendocrine prostate cancer. Nat Med. 2016;22:298–305.
Wang HT, Yao YH, Li BG, Tang Y, Chang JW, Zhang J. Neuroendocrine Prostate Cancer (NEPC) progressing from conventional prostatic adenocarcinoma: factors associated with time to development of NEPC and survival from NEPC diagnosis—a systematic review and pooled analysis. J Clin Oncol. 2014;32:3383–90.
Beltran H, Hruszkewycz A, Scher HI, Hildesheim J, Isaacs J, Yu EY, et al. The role of lineage plasticity in prostate cancer therapy resistance. Clin Cancer Res Off J Am Assoc Cancer Res. 2019;25:6916–24.
Davies AH, Beltran H, Zoubeidi A. Cellular plasticity and the neuroendocrine phenotype in prostate cancer. Nat Rev Urol. 2018;15:271–86.
Wang Y, Wang Y, Ci X, Choi SYC, Crea F, Lin D, et al. Molecular events in neuroendocrine prostate cancer development. Nat Rev Urol. 2021;18:581–96.
Liang X, Lin A, Wang Q, Zhang J, Luo P. Cell plasticity in patients with NSCLC: the controversial origins of transformed SCLC. Biomed Pharmacother. 2022;149:112909.
Conteduca V, Oromendia C, Eng KW, Bareja R, Sigouros M, Molina A, et al. Clinical features of neuroendocrine prostate cancer. Eur J Cancer. 2019;121:7–18.
Yamada Y, Beltran H. Clinical and biological features of neuroendocrine prostate cancer. Curr Oncol Rep. 2021;23:15.
Sreekumar A, Saini S. Role of microRNAs in neuroendocrine prostate cancer. Noncoding RNA. 2022;8:25.
Beltran H, Demichelis F. Therapy considerations in neuroendocrine prostate cancer: what next? Endocr Relat Cancer. 2021;28:T67–78.
Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, et al. Landscape of transcription in human cells. Nature. 2012;489:101–8.
Yang Y, Liu KY, Liu Q, Cao Q. Androgen receptor-related non-coding RNAs in prostate cancer. Front Cell Dev Biol. 2021;9:660853.
Esteller M. Non-coding RNAs in human disease. Nat Rev Genet. 2011;12:861–74.
Altschuler J, Stockert JA, Kyprianou N. Non-coding RNAs set a new phenotypic frontier in prostate cancer metastasis and resistance. Int J Mol Sci. 2021;22:2100.
Hayes J, Peruzzi PP, Lawler S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med. 2014;20:460–9.
Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer. 2015;15:321–33.
Khan S, Ayub H, Khan T, Wahid F. MicroRNA biogenesis, gene silencing mechanisms and role in breast, ovarian and prostate cancer. Biochimie. 2019;167:12–24.
Jafarzadeh A, Paknahad MH, Nemati M, Jafarzadeh S, Mahjoubin-Tehran M, Rajabi A, et al. Dysregulated expression and functions of microRNA-330 in cancers: a potential therapeutic target. Biomed Pharmacother. 2022;146:112600.
Aghdam SG, Ebrazeh M, Hemmatzadeh M, Seyfizadeh N, Shabgah AG, Azizi G, et al. The role of microRNAs in prostate cancer migration, invasion, and metastasis. J Cell Physiol. 2019;234:9927–42.
Nadiminty N, Tummala R, Lou W, Zhu Y, Zhang J, Chen X, et al. MicroRNA let-7c suppresses androgen receptor expression and activity via regulation of Myc expression in prostate cancer cells. J Biol Chem. 2012;287:1527–37.
Fernandes RC, Toubia J, Townley S, Hanson AR, Dredge BK, Pillman KA, et al. Post-transcriptional gene regulation by microRNA-194 promotes neuroendocrine transdifferentiation in prostate cancer. Cell Rep. 2021;34:108585.
Ding M, Lin B, Li T, Liu Y, Li Y, Zhou X, et al. A dual yet opposite growth-regulating function of miR-204 and its target XRN1 in prostate adenocarcinoma cells and neuroendocrine-like prostate cancer cells. Oncotarget. 2015;6:7686–700.
Natani S, Ramakrishna M, Nallavolu T, Ummanni R. MicroRNA-147b induces neuroendocrine differentiation of prostate cancer cells by targeting ribosomal protein RPS15A. Prostate. 2023;83:936–49.
Naidoo M, Levine F, Gillot T, Orunmuyi AT, Olapade-Olaopa EO, Ali T, et al. MicroRNA-1205 regulation of FRYL in prostate cancer. Front Cell Dev Biol. 2021;9:647485.
Ma S, Chan YP, Kwan PS, Lee TK, Yan M, Tang KH, et al. MicroRNA-616 induces androgen-independent growth of prostate cancer cells by suppressing expression of tissue factor pathway inhibitor TFPI-2. Cancer Res. 2011;71:583–92.
Yu J, Lu Y, Cui D, Li E, Zhu Y, Zhao Y, et al. miR-200b suppresses cell proliferation, migration and enhances chemosensitivity in prostate cancer by regulating Bmi-1. Oncol Rep. 2014;31:910–8.
Urabe F, Yamamoto Y, Kimura T. miRNAs in prostate cancer: intercellular and extracellular communications. Int J Urol. 2022;29:1429–38.
Guan H, Peng R, Fang F, Mao L, Chen Z, Yang S, et al. Tumor-associated macrophages promote prostate cancer progression via exosome-mediated miR-95 transfer. J Cell Physiol. 2020;235:9729–42.
Patel GK, Dutta S, Syed MM, Ramachandran S, Sharma M, Rajamanickam V, et al. TBX2 drives neuroendocrine prostate cancer through exosome-mediated repression of miR-200c-3p. Cancers. 2021;13:5020.
Bhagirath D, Liston M, Patel N, Akoto T, Lui B, Yang TL, et al. MicroRNA determinants of neuroendocrine differentiation in metastatic castration-resistant prostate cancer. Oncogene. 2020;39:7209–23.
Nabavi N, Saidy NRN, Venalainen E, Haegert A, Parolia A, Xue H, et al. miR-100-5p inhibition induces apoptosis in dormant prostate cancer cells and prevents the emergence of castration-resistant prostate cancer. Sci Rep. 2017;7:4079.
Takeshita F, Patrawala L, Osaki M, Takahashi RU, Yamamoto Y, Kosaka N, et al. Systemic delivery of synthetic microRNA-16 inhibits the growth of metastatic prostate tumors via downregulation of multiple cell-cycle genes. Mol Ther. 2010;18:181–7.
Xu YH, Deng JL, Wang G, Zhu YS. Long non-coding RNAs in prostate cancer: functional roles and clinical implications. Cancer Lett. 2019;464:37–55.
Bhan A, Soleimani M, Mandal SS. Long noncoding RNA and cancer: a new paradigm. Cancer Res. 2017;77:3965–81.
Misawa A, Takayama KI, Inoue S. Long non-coding RNAs and prostate cancer. Cancer Sci. 2017;108:2107–14.
Mirzaei S, Paskeh MDA, Okina E, Gholami MH, Hushmandi K, Hashemi M, et al. Molecular landscape of LncRNAs in prostate cancer: a focus on pathways and therapeutic targets for intervention. J Exp Clin Cancer Res. 2022;41:214.
Smolarz B, Zadrożna-Nowak A, Romanowicz H. The role of lncRNA in the development of tumors, including breast cancer. Int J Mol Sci. 2021;22:8427.
Fang Y, Fullwood MJ. Roles, functions, and mechanisms of long non-coding RNAs in cancer. Genomics Proteom Bioinforma. 2016;14:42–54.
Liu B, Jiang HY, Yuan T, Luo J, Zhou WD, Jiang QQ, et al. Enzalutamide-induced upregulation of PCAT6 promotes prostate cancer neuroendocrine differentiation by regulating miR-326/HNRNPA2B1 axis. Front Oncol. 2021;11:650054.
Singh N, Ramnarine VR, Song JH, Pandey R, Padi SKR, Nouri M, et al. The long noncoding RNA H19 regulates tumor plasticity in neuroendocrine prostate cancer. Nat Commun. 2021;12:7349.
Ghildiyal R, Sawant M, Renganathan A, Mahajan K, Kim EH, Luo J, et al. Loss of long noncoding RNA NXTAR in prostate cancer augments androgen receptor expression and enzalutamide resistance. Cancer Res. 2022;82:155–68.
Luo J, Wang K, Yeh S, Sun Y, Liang L, Xiao Y, et al. LncRNA-p21 alters the antiandrogen enzalutamide-induced prostate cancer neuroendocrine differentiation via modulating the EZH2/STAT3 signaling. Nat Commun. 2019;10:2571.
Zhang B, Zhang M, Shen C, Liu G, Zhang F, Hou J, et al. LncRNA PCBP1-AS1-mediated AR/AR-V7 deubiquitination enhances prostate cancer enzalutamide resistance. Cell Death Dis. 2021;12:856.
Liu B, Jiang HY, Yuan T, Zhou WD, Xiang ZD, Jiang QQ, et al. Long non-coding RNA AFAP1-AS1 facilitates prostate cancer progression by regulating miR-15b/IGF1R axis. Curr Pharm Des. 2021;27:4261–69.
Jiang X, Guo S, Xu M, Ma B, Liu R, Xu Y, et al. TFAP2C-mediated lncRNA PCAT1 inhibits ferroptosis in docetaxel-resistant prostate cancer through c-Myc/miR-25-3p/SLC7A11 signaling. Front Oncol. 2022;12:862015.
Mather RL, Parolia A, Carson SE, Venalainen E, Roig-Carles D, Jaber M, et al. The evolutionarily conserved long non-coding RNA LINC00261 drives neuroendocrine prostate cancer proliferation and metastasis via distinct nuclear and cytoplasmic mechanisms. Mol Oncol. 2021;15:1921–41.
Zhang D, Fang C, Li H, Lu C, Huang J, Pan J, et al. Long ncRNA MALAT1 promotes cell proliferation, migration, and invasion in prostate cancer via sponging miR-145. Transl Androl Urol. 2021;10:2307–19.
Wang ZH, Wang JH, Wang KQ, Zhou Y, Wang J. LncRNA FEZF1-AS1 promoted chemoresistance, autophagy and epithelial-mesenchymal transition (EMT) through regulation of miR-25-3p/ITGB8 axis in prostate cancer. Eur Rev Med Pharmacol Sci. 2020;24:2281–93.
Ye C, Chen YG, Qin SF, Tang SY, Li S, Shi MF, et al. Enzalutamide-resistant related lncRNA NONHSAT210528 promotes the proliferation and invasion of prostate cancer. Transl Androl Urol. 2022;11:643–58.
Hsu MT, Coca-Prados M. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature. 1979;280:339–40.
Lei M, Zheng G, Ning Q, Zheng J, Dong D. Translation and functional roles of circular RNAs in human cancer. Mol Cancer. 2020;19:30.
Yang L, Zou X, Zou J, Zhang G. Functions of circular RNAs in bladder, prostate and renal cell cancer (Review). Mol Med Rep. 2021;23:307.
Yang X, Ye T, Liu H, Lv P, Duan C, Wu X, et al. Expression profiles, biological functions and clinical significance of circRNAs in bladder cancer. Mol Cancer. 2021;20:4.
Zhang ZH, Wang Y, Zhang Y, Zheng SF, Feng T, Tian X, et al. The function and mechanisms of action of circular RNAs in urologic cancer. Mol Cancer. 2023;22:61.
Chao F, Wang S, Zhang C, Han D, Xu G, Chen G. The emerging role of circular RNAs in prostate cancer: a systematic review. Front Cell Dev Biol. 2021;9:681163.
Ding Y, Wang M, Yang J. Circular RNA midline-1 (circMID1) promotes proliferation, migration, invasion and glycolysis in prostate cancer. Bioengineered. 2022;13:6293–308.
Chen Y, Yang F, Fang E, Xiao W, Mei H, Li H, et al. Circular RNA circAGO2 drives cancer progression through facilitating HuR-repressed functions of AGO2-miRNA complexes. Cell Death Differ. 2019;26:1346–64.
Zhang Y, Liu F, Feng Y, Xu X, Wang Y, Zhu S, et al. CircRNA circ_0006156 inhibits the metastasis of prostate cancer by blocking the ubiquitination of S100A9. Cancer Gene Ther. 2022;29:1731–41.
Ding L, Wang R, Zheng Q, Shen D, Wang H, Lu Z, et al. circPDE5A regulates prostate cancer metastasis via controlling WTAP-dependent N6-methyladenisine methylation of EIF3C mRNA. J Exp Clin Cancer Res. 2022;41:187.
Zhou H, Zheng XD, Lin CM, Min J, Hu S, Hu Y, et al. Advancement and properties of circular RNAs in prostate cancer: an emerging and compelling frontier for discovering. Int J Biol Sci. 2021;17:651–69.
Luo G-C, Chen L, Fang J, Yan Z-J. Hsa_circ_0030586 promotes epithelial–mesenchymal transition in prostate cancer via PI3K-AKT signaling. Bioengineered. 2021;12:11089–107.
Mao S, Zhang W, Yang F, Guo Y, Wang H, Wu Y, et al. Hsa_circ_0004296 inhibits metastasis of prostate cancer by interacting with EIF4A3 to prevent nuclear export of ETS1 mRNA. J Exp Clin Cancer Res. 2021;40:336.
Shen Z, Zhou L, Zhang C, Xu J. Reduction of circular RNA Foxo3 promotes prostate cancer progression and chemoresistance to docetaxel. Cancer Lett. 2020;468:88–101.
Yu YZ, Lv DJ, Wang C, Song XL, Xie T, Wang T, et al. Hsa_circ_0003258 promotes prostate cancer metastasis by complexing with IGF2BP3 and sponging miR-653-5p. Mol Cancer. 2022;21:12.
Chen L, Song Y, Hou T, Li X, Cheng L, Li Y, et al. Circ_0004087 interaction with SND1 promotes docetaxel resistance in prostate cancer by boosting the mitosis error correction mechanism. J Exp Clin Cancer Res. 2022;41:194.
Jiang X, Guo S, Wang S, Zhang Y, Chen H, Wang Y, et al. EIF4A3-induced circARHGAP29 promotes aerobic glycolysis in docetaxel-resistant prostate cancer through IGF2BP2/c-Myc/LDHA signaling. Cancer Res. 2022;82:831–45.
Lim MCJ, Baird AM, Greene J, Mcnevin C, Ronan K, Podlesniy P, et al. hsa_circ_0001275 is one of a number of circRNAs dysregulated in enzalutamide resistant prostate cancer and confers enzalutamide resistance in vitro. Cancers. 2021;13:6383.
Yu Y, Song Y, Cheng L, Chen L, Liu B, Lu D, et al. CircCEMIP promotes anoikis-resistance by enhancing protective autophagy in prostate cancer cells. J Exp Clin Cancer Res. 2022;41:188.
Lu J, Zhong C, Luo J, Shu F, Lv D, Liu Z, et al. HnRNP-L-regulated circCSPP1/miR-520h/EGR1 axis modulates autophagy and promotes progression in prostate cancer. Mol Ther Nucleic Acids. 2021;26:927–44.
Wu X, Pan Y, Fang Y, Zhang J, Xie M, Yang F, et al. The biogenesis and functions of piRNAs in human diseases. Mol Ther Nucleic Acids. 2020;21:108–20.
Liu Y, Dou M, Song X, Dong Y, Liu S, Liu H, et al. The emerging role of the piRNA/piwi complex in cancer. Mol Cancer. 2019;18:123.
Tan L, Mai D, Zhang B, Jiang X, Zhang J, Bai R, et al. PIWI-interacting RNA-36712 restrains breast cancer progression and chemoresistance by interaction with SEPW1 pseudogene SEPW1P RNA. Mol Cancer. 2019;18:9.
Ou B, Liu Y, Gao Z, Xu J, Yan Y, Li Y, et al. Senescent neutrophils-derived exosomal piRNA-17560 promotes chemoresistance and EMT of breast cancer via FTO-mediated m6A demethylation. Cell Death Dis. 2022;13:905.
Zhang L, Meng X, Li D, Han X. piR-001773 and piR-017184 promote prostate cancer progression by interacting with PCDH9. Cell Signal. 2020;76:109780.
Dietrich D, Meller S, Uhl B, Ralla B, Stephan C, Jung K, et al. Nucleic acid-based tissue biomarkers of urologic malignancies. Crit Rev Clin Lab Sci. 2014;51:173–99.
Markert L, Holdmann J, Klinger C, Kaufmann M, Schork K, Turewicz M, et al. Small RNAs as biomarkers to differentiate benign and malign prostate diseases: an alternative for transrectal punch biopsy of the prostate? PLoS ONE. 2021;16:e0247930.
Otandault A, Anker P, Al Amir Dache Z, Guillaumon V, Meddeb R, Pastor B, et al. Recent advances in circulating nucleic acids in oncology. Ann Oncol. 2019;30:374–84.
Berchuck JE, Baca SC, Mcclure HM, Korthauer K, Tsai HK, Nuzzo PV, et al. Detecting neuroendocrine prostate cancer through tissue-informed cell-free DNA methylation analysis. Clin Cancer Res. 2022;28:928–38.
Yan H, Bu P. Non-coding RNA in cancer. Essays Biochem. 2021;65:625–39.
Pickl JM, Heckmann D, Ratz L, Klauck SM, Sültmann H. Novel RNA markers in prostate cancer: functional considerations and clinical translation. Biomed Res Int. 2014;2014:765207.
Romano G, Veneziano D, Acunzo M, Croce CM. Small non-coding RNA and cancer. Carcinogenesis. 2017;38:485–91.
Babayan A, Pantel K. Advances in liquid biopsy approaches for early detection and monitoring of cancer. Genome Med. 2018;10:21.
Zhao SG, Sperger JM, Schehr JL, Mckay RR, Emamekhoo H, Singh A, et al. A clinical-grade liquid biomarker detects neuroendocrine differentiation in prostate cancer. J Clin Investig. 2022;132:e161858.
Lin BB, Lei HQ, Xiong HY, Fu X, Shi F, Yang XW, et al. MicroRNA-regulated transcriptome analysis identifies four major subtypes with prognostic and therapeutic implications in prostate cancer. Comput Struct Biotechnol J. 2021;19:4941–53.
Fabris L, Ceder Y, Chinnaiyan AM, Jenster GW, Sorensen KD, Tomlins S, et al. The potential of microRNAs as prostate cancer biomarkers. Eur Urol. 2016;70:312–22.
Moustafa AA, Ziada M, Elshaikh A, Datta A, Kim H, Moroz K, et al. Identification of microRNA signature and potential pathway targets in prostate cancer. Exp Biol Med. 2017;242:536–46.
Ambs S, Prueitt RL, Yi M, Hudson RS, Howe TM, Petrocca F, et al. Genomic profiling of microRNA and messenger RNA reveals deregulated microRNA expression in prostate cancer. Cancer Res. 2008;68:6162–70.
Ostano P, Mello-Grand M, Sesia D, Gregnanin I, Peraldo-Neia C, Guana F, et al. Gene expression signature predictive of neuroendocrine transformation in prostate adenocarcinoma. Int J Mol Sci. 2020;21:1078.
Alahari SV, Eastlack SC, Alahari SK. Role of long noncoding RNAs in neoplasia: special emphasis on prostate cancer. Int Rev Cell Mol Biol. 2016;324:229–54.
Mouraviev V, Lee B, Patel V, Albala D, Johansen TE, Partin A, et al. Clinical prospects of long noncoding RNAs as novel biomarkers and therapeutic targets in prostate cancer. Prostate Cancer Prostatic Dis. 2016;19:14–20.
Martens-Uzunova ES, Böttcher R, Croce CM, Jenster G, Visakorpi T, Calin GA. Long noncoding RNA in prostate, bladder, and kidney cancer. Eur Urol. 2014;65:1140–51.
Li P, Shi Y, Guo M, Xu H, Zhan M, Wang Z, et al. MAFG-AS1 is a prognostic biomarker and facilitates prostate cancer progression. Front Oncol. 2022;12:856580.
Li W, Zhou R, Sun B, Jin X, Chen Y, Xu X. Prognostic significance of lncRNA AP004608.1 in prostate cancer. Front Oncol. 2022;12:1017635.
Li C, Hu J, Hu X, Zhao C, Mo M, Zu X, et al. LncRNA SNHG9 is a prognostic biomarker and correlated with immune infiltrates in prostate cancer. Transl Androl Urol. 2021;10:215–26.
Ramnarine VR, Alshalalfa M, Mo F, Nabavi N, Erho N, Takhar M, et al. The long noncoding RNA landscape of neuroendocrine prostate cancer and its clinical implications. Gigascience. 2018;7:giy050.
Crea F, Venalainen E, Ci X, Cheng H, Pikor L, Parolia A, et al. The role of epigenetics and long noncoding RNA MIAT in neuroendocrine prostate cancer. Epigenomics. 2016;8:721–31.
Zhang Z, Yang T, Xiao J. Circular RNAs: promising biomarkers for human diseases. EBioMedicine. 2018;34:267–74.
Fang J, Qi J, Dong X, Luo J. Perspectives on circular RNAs as prostate cancer biomarkers. Front Cell Dev Biol. 2020;8:594992.
Greene J, Baird AM, Lim M, Flynn J, Mcnevin C, Brady L, et al. Differential CircRNA expression signatures may serve as potential novel biomarkers in prostate cancer. Front Cell Dev Biol. 2021;9:605686.
Vo JN, Cieslik M, Zhang Y, Shukla S, Xiao L, Zhang Y, et al. The landscape of circular RNA in cancer. Cell. 2019;176:869–81.e13.
He T, Tao W, Zhang LL, Wang BY, Li K, Lu HM, et al. CircSCAF8 promotes growth and metastasis of prostate cancer through the circSCAF8-miR-140-3p/miR-335-LIF pathway. Cell Death Dis. 2022;13:517.
Li T, Sun X, Chen L. Exosome circ_0044516 promotes prostate cancer cell proliferation and metastasis as a potential biomarker. J Cell Biochem. 2020;121:2118–26.
Hansen EB, Fredsøe J, Okholm TLH, Ulhøi BP, Klingenberg S, Jensen JB, et al. The transcriptional landscape and biomarker potential of circular RNAs in prostate cancer. Genome Med. 2022;14:8.
Tao W, Luo ZH, He YD, Wang BY, Xia TL, Deng WM, et al. Plasma extracellular vesicle circRNA signature and resistance to abiraterone in metastatic castration-resistant prostate cancer. Br J cancer. 2023;128:1320–32.
Jia DD, Jiang H, Zhang YF, Zhang Y, Qian LL, Zhang YF. The regulatory function of piRNA/PIWI complex in cancer and other human diseases: the role of DNA methylation. Int J Biol Sci. 2022;18:3358–73.
Hanusek K, Poletajew S, Kryst P, Piekiełko-Witkowska A, Bogusławska J. piRNAs and PIWI proteins as diagnostic and prognostic markers of genitourinary cancers. Biomolecules. 2022;12:186.
Peng Q, Chiu PK, Wong CY, Cheng CK, Teoh JY, Ng CF. Identification of piRNA targets in urinary extracellular vesicles for the diagnosis of prostate cancer. Diagnostics. 2021;11:1828.
Zuo YL, Liang Y, Zhang JT, Hao YY, Li ML, Wen ZN, et al. Transcriptome analysis identifies Piwi-interacting RNAs as prognostic markers for recurrence of prostate cancer. Front Genet. 2019;10:1018.
Ge R, Wang Z, Montironi R, Jiang Z, Cheng M, Santoni M, et al. Epigenetic modulations and lineage plasticity in advanced prostate cancer. Ann Oncol. 2020;31:470–79.
Yan Z, Xiao Y, Chen Y, Luo G. Screening and identification of epithelial-to-mesenchymal transition-related circRNA and miRNA in prostate cancer. Pathol Res Pr. 2020;216:152784.
Malla B, Zaugg K, Vassella E, Aebersold DM, Dal Pra A. Exosomes and exosomal microRNAs in prostate cancer radiation therapy. Int J Radiat Oncol Biol Phys. 2017;98:982–95.
Konoshenko M, Laktionov P. The miRNAs involved in prostate cancer chemotherapy response as chemoresistance and chemosensitivity predictors. Andrology. 2022;10:51–71.
Juracek J, Madrzyk M, Stanik M, Slaby O. Urinary microRNAs and their significance in prostate cancer diagnosis: a 5-year update. Cancers. 2022;14:3157.
Acknowledgements
This work was supported by National Natural Science Foundation of China (No. 82303122), Natural Science Foundation of Hunan Province (Nos. 2021JJ30613, 2022JJ30106), Health Research Project of Hunan Provincial Health Commission (Nos. B20180192, D202304059395, C20180075), Hunan University students Innovation training program project (No. 2022X10555139).
Author information
Authors and Affiliations
Contributions
The idea of the review was conceived by EML, who also created the Figures. WHT took charge of writing the manuscript and creating the Table, significantly contributing to the manuscript’s composition. CKX and MM conducted the initial research or gathered the data used in the review. YHC and ZGH prepared figures or tables to illustrate the findings. XLW, RK and ZFL critically reviewed and edited drafts of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tan, W., Xiao, C., Ma, M. et al. Role of non-coding RNA in lineage plasticity of prostate cancer. Cancer Gene Ther 32, 1–10 (2025). https://doi.org/10.1038/s41417-024-00834-z
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41417-024-00834-z