Abstract
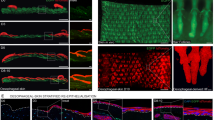
Androgen deprivation induces extensive tissue remodelling in the prostate, characterized by epithelial cell attrition and acquisition of a progenitor-like state in residual luminal epithelial cells. The mechanisms driving differentiated epithelial cells toward a progenitor fate remain unclear. Here, we identify that prostate regression is mediated by the engulfment of apoptotic neighbours by epithelial cells and that efferocytosis promotes acquisition of a progenitor-like state. We show that epithelial cells are the predominant phagocytes during regression, engaging in temporally coordinated waves of apoptotic cell clearance. This process is accompanied by marked metabolic reprogramming, including increased aerobic glycolysis and lactate production. This coincides with enhanced histone lysine-lactylation at promoters of genes involved in autophagy, apoptosis regulation, and luminal progenitor identity. Blockade of efferocytosis in vivo via epithelial-specific expression of the dominant negative phosphatidylserine binding protein MFGE8-D89E impaired prostate regression and compromised the induction of the luminal progenitor marker Tacstd2. These findings reveal that epithelial efferocytosis is an essential mechanism that couples cell clearance and epithelial plasticity. This work establishes epithelial efferocytosis as a determinant of cell state transitions in the prostate, with implications for a direct role in castration-resistant prostate cancer and other regenerative or remodelling processes.
Similar content being viewed by others
Data availability
All sequencing data (ChIP-seq) has been uploaded to the GEO repository under the accession number GSE311917.
References
Coffey D, Ichinose R, Shimazaki J, Williams-Ashman H. Effects of testosterone on adenosine triphosphate and nicotinamide adenine dinucleotide levels, and on nicotinamide mononucleotide adenylytransferse activity, in the ventral prostate of castrated rats. Mol Pharmacol. 1968;4:580–90.
Reiss AB, Gulkarov S, Pinkhasov A, Sheehan KM, Srivastava A, De Leon J. et al. Androgen deprivation therapy for prostate cancer: focus on cognitive function and mood. Medicina (Kaunas). 2023;60:77.
Maru S, Uchino H, Osawa T, Chiba S, Mouri G, Sazawa A. Long-term treatment outcomes of intermittent androgen deprivation therapy for relapsed prostate cancer after radical prostatectomy. PLoS One. 2018;13:e0197252.
Wang X, Kruithof-de Julio M, Economides KD, Walker D, Yu H, et al. A luminal epithelial stem cell that is a cell of origin for prostate cancer. Nature. 2009;461:495–500.
Choi N, Zhang B, Zhang L, Ittmann M, Xin L. Adult murine prostate basal and luminal cells are self-sustained lineages that can both serve as targets for prostate cancer initiation. Cancer Cell. 2012;21:253–65.
Ousset M, Van Keymeulen A, Bouvencourt G, Sharma N, Achouri Y, Simons BD, et al. Multipotent and unipotent progenitors contribute to prostate postnatal development. Nat Cell Biol. 2012;14:1131–8.
Karthaus WR, Hofree M, Choi D, Linton EL, Turkekul M, Bejnood A, et al. Regenerative potential of prostate luminal cells revealed by single-cell analysis. Science. 2020;368:497–505.
Kirk JS, Wang J, Long M, Rosario S, Tracz A, Ji Y, et al. Integrated single-cell analysis defines the epigenetic basis of castration-resistant prostate luminal cells. Cell Stem Cell. 2024;31:1203–21.e7.
Kerr JFR, Searle J. Deletion of cells by apoptosis during castration-induced involution of the rat prostate. Virchows Arch B Cell Pathol. 1973;13:87–102.
Bruni-Cardoso A, Augusto TM, Pravatta H, Damas-Souza DM, Carvalho HF. Stromal remodelling is required for progressive involution of the rat ventral prostate after castration: identification of a matrix metalloproteinase-dependent apoptotic wave. Int J Androl. 2010;33:686–95.
Fogarty CE, Bergmann A. The sound of silence: signaling by apoptotic cells. Curr Top Dev Biol. 2015;114:241–65.
Fadok VA, de Cathelineau A, Daleke DL, Henson PM, Bratton DL. Loss of phospholipid asymmetry and surface exposure of phosphatidylserine is required for phagocytosis of apoptotic cells by macrophages and fibroblasts. J Biol Chem. 2001;276:1071–7.
Colegio OR, Chu NQ, Szabo AL, Chu T, Rhebergen AM, Jairam V, et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature. 2014;513:559–63.
Morioka S, Perry JSA, Raymond MH, Medina CB, Zhu Y, Zhao L, et al. Efferocytosis induces a novel SLC program to promote glucose uptake and lactate release. Nature. 2018;563:714–8.
Zhang S, Weinberg S, DeBerge M, Gainullina A, Schipma M, Kinchen JM, et al. Efferocytosis fuels requirements of fatty acid oxidation and the electron transport chain to polarize macrophages for tissue repair. Cell Metab. 2019;29:443–56.e5.
Ngai D, Schilperoort M, Tabas I. Efferocytosis-induced lactate enables the proliferation of pro-resolving macrophages to mediate tissue repair. Nat Metab. 2023;5:2206–19.
English HF, Santen RJ. Isaacs JT. Response of glandular versus basal rat ventral prostatic epithelial cells to androgen withdrawal and replacement. Prostate. 1987;11:229–42.
Kerr JF, Searle J. Deletion of cells by apoptosis during castration-induced involution of the rat prostate. Virchows Arch B Cell Pathol. 1973;13:87–102.
Garcia-Florez M, Oliveira CA, Carvalho HF. Early effects of estrogen on the rat ventral prostate. Braz J Med Biol Res. 2005;38:487–97.
Isaacs JT. Antagonistic effect of androgen on prostatic cell death. Prostate. 1984;5:545–57.
English HF, Kyprianou N, Isaacs JT. Relationship between DNA fragmentation and apoptosis in the programmed cell death in the rat prostate following castration. Prostate. 1989;15:233–50.
Kyprianou N, Isaacs JT. Activation of programmed cell death in the rat ventral prostate after castration. Endocrinology. 1988;122:552–62.
Crowley L, Cambuli F, Aparicio L, Shibata M, Robinson BD, Xuan S. et al. A single-cell atlas of the mouse and human prostate reveals heterogeneity and conservation of epithelial progenitors. Elife. 2020;9:e59465.
Rosa-Ribeiro R, Barbosa GO, Kuhne F, Carvalho HF. Desquamation is a novel phenomenon for collective prostate epithelial cell deletion after castration. Histochem Cell Biol. 2014;141:213–20.
Merkuri F, Rothstein M, Simoes-Costa M. Histone lactylation couples cellular metabolism with developmental gene regulatory networks. Nat Commun. 2024;15:90.
Asano K, Miwa M, Miwa K, Hanayama R, Nagase H, Nagata S, et al. Masking of phosphatidylserine inhibits apoptotic cell engulfment and induces autoantibody production in mice. J Exp Med. 2004;200:459–67.
Tang YC, Ponsin K, Graham-Paquin AL, Luthold C, Homsy K, Schindler M, et al. Coordination of non-professional efferocytosis and actomyosin contractility during epithelial tissue morphogenesis. Cell Rep. 2023;42:112202.
Greenhalgh DG. The role of apoptosis in wound healing. Int J Biochem Cell Biol. 1998;30:1019–30.
Wu YS, Chen SN. Apoptotic cell: linkage of inflammation and wound healing. Front Pharm. 2014;5:1.
Tseng AS, Adams DS, Qiu D, Koustubhan P, Levin M. Apoptosis is required during early stages of tail regeneration in Xenopus laevis. Dev Biol. 2007;301:62–9.
Guicciardi ME, Malhi H, Mott JL, Gores GJ. Apoptosis and necrosis in the liver. Compr Physiol. 2013;3:977–1010.
Yan L, Wang J, Cai X, Liou YC, Shen HM, Hao J, et al. Macrophage plasticity: signaling pathways, tissue repair, and regeneration. MedComm (2020). 2024;5:e658.
Sihombing M, Safitri M, Zhou T, Wang L, McGinty S, Zhang HJ, et al. Unexpected Role of Nonimmune Cells: Amateur Phagocytes. DNA Cell Biol. 2021;40:157–71.
Silva JAF, Bruni-Cardoso A, Augusto TM, Damas-Souza DM, Barbosa GO, Felisbino SL, et al. Macrophage roles in the clearance of apoptotic cells and control of inflammation in the prostate gland after castration. Prostate. 2018;78:95–103.
Beltran H, Tomlins S, Aparicio A, Arora V, Rickman D, Ayala G, et al. Aggressive variants of castration-resistant prostate cancer. Clin Cancer Res. 2014;20:2846–50.
Madisen L, Zwingman TA, Sunkin SM, Oh SW, Zariwala HA, Gu H, et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat Neurosci. 2010;13:133–40.
Tremblay M, Viala S, Shafer ME, Graham-Paquin AL, Liu C, Bouchard M. Regulation of stem/progenitor cell maintenance by BMP5 in prostate homeostasis and cancer initiation. Elife. 2020;9:e54542.
Viala S, Hadjadj C, Nathan V, Guiot MC, McCaffrey L, Cockburn K, et al. LGN loss randomizes spindle orientation and accelerates tumorigenesis in PTEN-deficient epidermis. Mol Biol Cell. 2024;35:br5.
Acknowledgements
We thank the Metabolomics Innovation Resource (MIR), the McGill University Advanced BioImaging Facility (ABIF), and the Rosalind and Morris Goodman Cancer Institute Histology Innovation Platform and Centre for Applied Genomics at SickKids Hospital for technical assistance. We thank Mitra Cowan and the McGill Integrated Core for Animal Modelling for assistance in generating mice. LM is a Fonds Recherche du Québec—Santé Research Scholar. ALGP was supported by a Canadian Institutes of Health Research Doctoral Fellowship. MKMW was supported by a FRQS Doctoral Fellowship. This work is dedicated to the memory of Maxime Bouchard. This work was supported by a Canadian Institutes of Health Research grant (PJT-159706) to MB and LM.
Author information
Authors and Affiliations
Contributions
ALGP and MB conceived the project. ALGP, MB, and LM directed the work, designed experiments, and interpreted data. ALGP performed the experiments. ALGP, SV, and MT generated mice used for experiments and performed mouse surgeries. MKMW, WAP, and DS supported data analysis. The manuscript was written by ALGP and edited by LM, SV, MKMW, MT, WAP, and DS. All authors discussed the data and contributed to the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics
All experimental procedures performed with animals were conducted in compliance with the Canadian Council of Animal Care (CACC) requirements for research using mice and overseen and approved by the McGill University Animal Care Committee.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Hans-Uwe Simon
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Graham-Paquin, AL., Saini, D., Viala, S. et al. Apoptotic cell clearance triggers epithelial fate reprogramming during prostate regression. Cell Death Dis (2026). https://doi.org/10.1038/s41419-026-08565-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-026-08565-9