Abstract
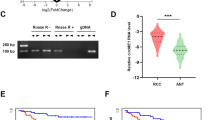
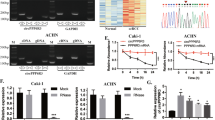
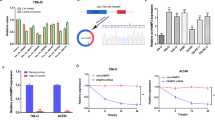
Renal cell carcinoma (RCC) relies heavily on aerobic glycolysis for rapid proliferation and metastasis; however, the role of circular RNAs (circRNAs) in this process remains unclear. This study identified circIQGAP1 as a key regulator; its expression was upregulated under glucose deficiency, with U2AF2 promoting its biosynthesis. Functional assays showed that circIQGAP1 enhances RCC cell proliferation, motility, invasion, and glycolytic flux. Mechanistically, circIQGAP1 binds to CARM1, inhibiting its K48-linked ubiquitination and prolonging its half-life. Elevated CARM1 then catalyses COL5A1 promoter dimethylation, driving COL5A1 transcription. Rescue experiments confirmed that both CARM1 and COL5A1 were essential for circIQGAP1-mediated metabolic reprogramming and malignant phenotypes. Clinically, circIQGAP1 is overexpressed in RCC tissues and is correlated with poor outcomes. These findings revealed that circIQGAP1 promotes glycolysis-dependent RCC progression by stabilizing CARM1 to activate COL5A1, highlighting that this regulatory axis may provide an innovative strategy for RCC treatment.
Similar content being viewed by others
Data availability
The data underlying the results presented in this paper are available from the corresponding author upon reasonable request. Full and uncropped blots are provided with this paper (Supplemental Material).
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Hsieh JJ, Purdue MP, Signoretti S, Swanton C, Albiges L, Schmidinger M, et al. Renal cell carcinoma. Nat Rev Dis Prim. 2017;3:17009.
Brassier M, Khene ZE, Bernhard JC, Dang VT, Ouzaid I, Nouhaud FX, et al. Percutaneous ablation versus surgical resection for local recurrence following partial nephrectomy for renal cell cancer: a propensity score analysis (REPART Study-UroCCR 71). Eur Urol Focus. 2022;8:210–6.
Santoni M, Mollica V, Rizzo A, Massari F. Dynamics of resistance to immunotherapy and TKI in patients with advanced renal cell carcinoma. Cancer Treat Rev. 2025;133:102881.
Rose TL, Kim WY. Renal cell carcinoma: a review. JAMA. 2024;332:1001–10.
Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science. 2009;324:1029–33.
Mirzaei H, Hamblin MR. Regulation of glycolysis by non-coding RNAs in cancer: switching on the Warburg effect. Mol Ther Oncolytics. 2020;19:218–39.
Fukushi A, Kim HD, Chang YC, Kim CH. Revisited metabolic control and reprogramming cancers by means of the Warburg effect in tumor cells. Int J Mol Sci. 2022;23:10037.
Singer K, Kastenberger M, Gottfried E, Hammerschmied CG, Büttner M, Aigner M, et al. Warburg phenotype in renal cell carcinoma: high expression of glucose-transporter 1 (GLUT-1) correlates with low CD8(+) T-cell infiltration in the tumor. Int J Cancer. 2011;128:2085–95.
Dong Y, Eskandari R, Ray C, Granlund KL, Santos-Cunha LD, Miloushev VZ, et al. Hyperpolarized MRI visualizes Warburg effects and predicts treatment response to mTOR inhibitors in patient-derived ccRCC xenograft models. Cancer Res. 2019;79:242–50.
Wettersten HI, Hakimi AA, Morin D, Bianchi C, Johnstone ME, Donohoe DR, et al. Grade-dependent metabolic reprogramming in kidney cancer revealed by combined proteomics and metabolomics analysis. Cancer Res. 2015;75:2541–52.
Jeck WR, Sharpless NE. Detecting and characterizing circular RNAs. Nat Biotechnol. 2014;32:453–61.
Li H, Su B, Jiang Y, Zhang B, Du R, Song C, et al. Circular RNA circDCUN1D4 suppresses hepatocellular carcinoma development via targeting the miR-590-5p/ TIMP3 axis. Mol Cancer. 2025;24:95.
Peng M, Zhang S, Wu P, Hou X, Wang D, Ge J, et al. Circular RNA circCLASP2 promotes nasopharyngeal carcinoma progression through binding to DHX9 to enhance PCMT1 translation. Mol Cancer. 2025;24:67.
Song X, Wei Z, Zhang C, Han D, Liu J, Song Y, et al. CircAKT3 promotes prostate cancer proliferation and metastasis by enhancing the binding of RPS27A and RPL11. Mol Cancer. 2025;24:53.
Yu T, Wang Y, Fan Y, Fang N, Wang T, Xu T, et al. CircRNAs in cancer metabolism: a review. J Hematol Oncol. 2019;12:90.
Wang S, Zhang Y, Cai Q, Ma M, Jin LY, Weng M, et al. Circular RNA FOXP1 promotes tumor progression and Warburg effect in gallbladder cancer by regulating PKLR expression. Mol Cancer. 2019;18:145.
Zheng J, Yan X, Lu T, Song W, Li Y, Liang J, et al. CircFOXK2 promotes hepatocellular carcinoma progression and leads to a poor clinical prognosis via regulating the Warburg effect. J Exp Clin Cancer Res. 2023;42:63.
Li T, Gu Y, Xu B, Kuca K, Zhang J, Wu W. CircZBTB44 promotes renal carcinoma progression by stabilizing HK3 mRNA structure. Mol Cancer. 2023;22:77.
Gui CP, Liao B, Luo CG, Chen YH, Tan L, Tang YM, et al. circCHST15 is a novel prognostic biomarker that promotes clear cell renal cell carcinoma cell proliferation and metastasis through the miR-125a-5p/EIF4EBP1 axis. Mol Cancer. 2021;20:169.
Cen J, Liang Y, Huang Y, Pan Y, Shu G, Zheng Z, et al. Circular RNA circSDHC serves as a sponge for miR-127-3p to promote the proliferation and metastasis of renal cell carcinoma via the CDKN3/E2F1 axis. Mol Cancer. 2021;20:19.
Kiss L, James LC, Schulman BA. UbiREAD deciphers proteasomal degradation code of homotypic and branched K48 and K63 ubiquitin chains. Mol Cell. 2025;85:1467–76.e6.
Suresh S, Huard S, Dubois T. CARM1/PRMT4: making its mark beyond its function as a transcriptional coactivator. Trends Cell Biol. 2021;31:402–17.
Vaupel P, Schmidberger H, Mayer A. The Warburg effect: essential part of metabolic reprogramming and central contributor to cancer progression. Int J Radiat Biol. 2019;95:912–9.
Ganapathy-Kanniappan S. Linking tumor glycolysis and immune evasion in cancer: Emerging concepts and therapeutic opportunities. Biochim Biophys Acta Rev Cancer. 2017;1868:212–20.
Zhao J, Jin D, Huang M, Ji J, Xu X, Wang F, et al. Glycolysis in the tumor microenvironment: a driver of cancer progression and a promising therapeutic target. Front Cell Dev Biol. 2024;12:1416472.
Nakajima R, Matsuo Y, Kondo T, Abe K, Sakai S. Prognostic value of metabolic tumor volume and total lesion glycolysis on preoperative 18F-FDG PET/CT in patients with renal cell carcinoma. Clin Nucl Med. 2017;42:e177–e82.
Rathmell WK, Rathmell JC, Linehan WM. Metabolic pathways in kidney cancer: current therapies and future directions. J Clin Oncol. 2018;36:3540–46.
Li J, Zhang Q, Guan Y, Liao D, Jiang D, Xiong H, et al. Circular RNA circVAMP3 promotes aerobic glycolysis and proliferation by regulating LDHA in renal cell carcinoma. Cell Death Dis. 2022;13:443.
Zhang MX, Wang JL, Mo CQ, Mao XP, Feng ZH, Li JY, et al. CircME1 promotes aerobic glycolysis and sunitinib resistance of clear cell renal cell carcinoma through cis-regulation of ME1. Oncogene. 2022;41:3979–90.
Glasser E, Maji D, Biancon G, Puthenpeedikakkal AMK, Cavender CE, Tebaldi T, et al. Pre-mRNA splicing factor U2AF2 recognizes distinct conformations of nucleotide variants at the center of the pre-mRNA splice site signal. Nucleic Acids Res. 2022;50:5299–312.
Huang Y, Zhu Q. Mechanisms regulating abnormal circular RNA biogenesis in cancer. Cancers. 2021;13:4185.
Pisignano G, Michael DC, Visal TH, Pirlog R, Ladomery M, Calin GA. Going circular: history, present, and future of circRNAs in cancer. Oncogene. 2023;42:2783–800.
Jiang Y, Zhou J, Zhao J, Zhang H, Li L, Li H, et al. The U2AF2 /circRNA ARF1/miR-342-3p/ISL2 feedback loop regulates angiogenesis in glioma stem cells. J Exp Clin Cancer Res. 2020;39:182.
Li H, Jiang Y, Hu J, Xu J, Chen L, Zhang G, et al. The U2AF65/circNCAPG/RREB1 feedback loop promotes malignant phenotypes of glioma stem cells through activating the TGF-β pathway. Cell Death Dis. 2023;14:23.
Wang Z, Sun A, Yan A, Yao J, Huang H, Gao Z, et al. Circular RNA MTCL1 promotes advanced laryngeal squamous cell carcinoma progression by inhibiting C1QBP ubiquitin degradation and mediating beta-catenin activation. Mol Cancer. 2022;21:92.
Dong G, Wang X, Wang X, Jia Y, Jia Y, Zhao W, et al. Circ_0084653 promotes the tumor progression and immune escape in triple-negative breast cancer via the deubiquitination of MYC and upregulation of SOX5. Int J Biol Macromol. 2024;280:135655.
Lu J, Wu H, Zhan P, Lu Y, Fang Q, Luo C, et al. PSMD14-mediated deubiquitination of CARM1 facilitates the proliferation and metastasis of hepatocellular carcinoma by inducing the transcriptional activation of FERMT1. Cell Death Dis. 2025;16:141.
Wang F, Zhang J, Tang H, Pang Y, Ke X, Peng W, et al. Nup54-induced CARM1 nuclear importation promotes gastric cancer cell proliferation and tumorigenesis through transcriptional activation and methylation of Notch2. Oncogene. 2022;41:246–59.
Feng S, Rao Z, Zhang J, She X, Chen Y, Wan K, et al. Inhibition of CARM1-mediated methylation of ACSL4 promotes ferroptosis in colorectal cancer. Adv Sci. 2023;10:e2303484.
Wang L, Zhao Z, Meyer MB, Saha S, Yu M, Guo A, et al. CARM1 methylates chromatin remodeling factor BAF155 to enhance tumor progression and metastasis. Cancer Cell. 2014;25:21–36.
Davis MB, Liu X, Wang S, Reeves J, Khramtsov A, Huo D, et al. Expression and sub-cellular localization of an epigenetic regulator, co-activator arginine methyltransferase 1 (CARM1), is associated with specific breast cancer subtypes and ethnicity. Mol Cancer. 2013;12:40.
Karakashev S, Zhu H, Wu S, Yokoyama Y, Bitler BG, Park PH, et al. CARM1-expressing ovarian cancer depends on the histone methyltransferase EZH2 activity. Nat Commun. 2018;9:631.
Li S, Cheng D, Zhu B, Yang Q. The overexpression of CARM1 promotes human osteosarcoma cell proliferation through the pGSK3β/β-Catenin/cyclinD1 signaling pathway. Int J Biol Sci. 2017;13:976–84.
Xie Z, Tian Y, Guo X, Xie N. The emerging role of CARM1 in cancer. Cell Oncol. 2024;47:1503–22.
Qiu Y, Wang H, Liao P, Xu B, Hu R, Yang Y, et al. Systematic pan-cancer landscape identifies CARM1 as a potential prognostic and immunological biomarker. BMC Genom Data. 2022;23:7.
Liu F, Ma F, Wang Y, Hao L, Zeng H, Jia C, et al. PKM2 methylation by CARM1 activates aerobic glycolysis to promote tumorigenesis. Nat Cell Biol. 2017;19:1358–70.
Abeywardana T, Oh M, Jiang L, Yang Y, Kong M, Song J, et al. CARM1 suppresses de novo serine synthesis by promoting PKM2 activity. J Biol Chem. 2018;293:15290–303.
Lin L, Yuan Q, Gu J, Bai G, Cong X, Hu Q, et al. CARM1-mediated OGT arginine methylation promotes non-small cell lung cancer glycolysis by stabilizing OGT. Cell Death Dis. 2024;15:927.
Zhong XY, Yuan XM, Xu YY, Yin M, Yan WW, Zou SW, et al. CARM1 methylates GAPDH to regulate glucose metabolism and is suppressed in liver cancer. Cell Rep. 2018;24:3207–23.
Covic M, Hassa PO, Saccani S, Buerki C, Meier NI, Lombardi C, et al. Arginine methyltransferase CARM1 is a promoter-specific regulator of NF-kappaB-dependent gene expression. EMBO J. 2005;24:85–96.
An W, Kim J, Roeder RG. Ordered cooperative functions of PRMT1, p300, and CARM1 in transcriptional activation by p53. Cell. 2004;117:735–48.
Wang X, Roberts CW. CARMA: CARM1 methylation of SWI/SNF in breast cancer. Cancer Cell. 2014;25:3–4.
Tsai HF, Chang YC, Li CH, Chan MH, Chen CL, Tsai WC, et al. Type V collagen alpha 1 chain promotes the malignancy of glioblastoma through PPRC1-ESM1 axis activation and extracellular matrix remodeling. Cell Death Discov. 2021;7:313.
Zhang Y, Jing Y, Wang Y, Tang J, Zhu X, Jin W-L, et al. NAT10 promotes gastric cancer metastasis via N4-acetylated COL5A1. Signal Transduct Target Ther. 2021;6:173.
Boguslawska J, Kedzierska H, Poplawski P, Rybicka B, Tanski Z, Piekielko-Witkowska A. Expression of genes involved in cellular adhesion and extracellular matrix remodeling correlates with poor survival of patients with renal cancer. J Urol. 2016;195:1892–902.
Feng G, Ma HM, Huang HB, Li YW, Zhang P, Huang JJ, et al. Overexpression of COL5A1 promotes tumor progression and metastasis and correlates with poor survival of patients with clear cell renal cell carcinoma. Cancer Manag Res. 2019;11:1263–74.
Yao J, Li R, Liu X, Zhou X, Li J, Liu T, et al. Prognostic implication of glycolysis related gene signature in non-small cell lung cancer. J Cancer. 2021;12:885–98.
Xiao Y, Huang W, Zhang L, Wang H. Identification of glycolysis genes signature for predicting prognosis in malignant pleural mesothelioma by bioinformatics and machine learning. Front Endocrinol. 2022;13:1056152.
Park Y, Park M, Kim J, Ahn J, Sim J, Bang JI, et al. NOX2-induced high glycolytic activity contributes to the gain of COL5A1-mediated mesenchymal phenotype in GBM. Cancers. Cancers. 2022;14:516.
Na J, Yang Z, Shi Q, Li C, Liu Y, Song Y, et al. Extracellular matrix stiffness as an energy metabolism regulator drives osteogenic differentiation in mesenchymal stem cells. Bioact Mater. 2024;35:549–63.
Xie Q, Qin F, Luo L, Deng S, Zeng K, Wu Y, et al. hsa_circ_0003596, as a novel oncogene, regulates the malignant behavior of renal cell carcinoma by modulating glycolysis. Eur J Med Res. 2023;28:315.
Santos M, Hwang JW, Bedford MT. CARM1 arginine methyltransferase as a therapeutic target for cancer. J Biol Chem. 2023;299:105124.
Drew AE, Moradei O, Jacques SL, Rioux N, Boriack-Sjodin AP, Allain C, et al. Identification of a CARM1 inhibitor with potent in vitro and in vivo activity in preclinical models of multiple myeloma. Sci Rep. 2017;7:17993.
Inoue F, Sone K, Toyohara Y, Tanimoto S, Takahashi Y, Kusakabe M, et al. Histone arginine methyltransferase CARM1 selective inhibitor TP-064 induces apoptosis in endometrial cancer. Biochem Biophys Res Commun. 2022;601:123–8.
Kumar S, Zeng Z, Bagati A, Tay RE, Sanz LA, Hartono SR, et al. CARM1 inhibition enables immunotherapy of resistant tumors by dual action on tumor cells and T cells. Cancer Discov. 2021;11:2050–71.
Zhang LX, Gao J, Long X, Zhang PF, Yang X, Zhu SQ, et al. The circular RNA circHMGB2 drives immunosuppression and anti-PD-1 resistance in lung adenocarcinomas and squamous cell carcinomas via the miR-181a-5p/CARM1 axis. Mol Cancer. 2022;21:110.
Zhao X, Qin X, Wang R, Wang Y, He Y, Wang Y, et al. An epigenetic nanoagonist facilitates T cell priming, recruitment, and reinvigoration in tumors resistant to PD-L1 therapy. Adv Mater. 2025;37:e02800.
Li Z, Lu H, Zhang Y, Lv J, Zhang Y, Xu T, et al. Blocking CXCR4-CARM1-YAP axis overcomes osteosarcoma doxorubicin resistance by suppressing aerobic glycolysis. Cancer Sci. 2024;115:3305–19.
Acknowledgements
We would like to express our gratitude to all members of our laboratory for their participation in this experiment. This work was supported by the National Natural Science Foundation of China (No. 82203815), the Natural Science Foundation of Shandong Province (No. ZR2020LZL011, ZR2022MH308, ZR2022MH003, and ZR2024QH467), Research Funds of the Shandong Provincial Key Laboratory of Animal Cell and Developmental Biology (No. SDKLACDB-2019010), and Key Scientific and Medical Project of Shandong (No. 2011QZ016 and 2016GSF201042).
Author information
Authors and Affiliations
Contributions
RJ and BZ contributed equally to this work. RG, RJ, BZ, and YL conceived of and designed the experiments. RG, RJ, BZ, YL, QC, TC, CZ, FP, and XZ performed the experiments. RG and BZ collected the clinical samples. RG, BZ, XL, and JB analyzed the data. RJ, BZ, YL, and RG wrote and revised the manuscript. All the authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Ethics Review Committee of Shandong Cancer Hospital and Institute (Approval no. SDTHEC2023001023). All experimental procedures were conducted in strict accordance with relevant guidelines and regulations, and informed consent was obtained from all patients.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Dr George Calin
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jia, R., Zou, B., Liang, Y. et al. CircIQGAP1-CARM1 axis promotes renal cell carcinoma progression through glycolytic reprogramming. Cell Death Dis (2026). https://doi.org/10.1038/s41419-026-08661-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-026-08661-w