Abstract
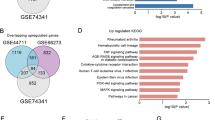
The dysregulation of growth factors is associated with defective trophoblast invasion, which leads to uteroplacental malperfusion due to the inadequate remodeling of spiral arteries. Pregnancy disorders, including preeclampsia, particularly early-onset preeclampsia, are closely related to compromised placental function caused by aberrant trophoblast invasion. Activin A, a growth factor detectable in serum that belongs to the transforming growth factor-β (TGF-β) superfamily, has been implicated in the development of preeclampsia, as evidenced by its elevated serum levels and its role in regulating trophoblast invasion. However, the existing research on its regulatory mechanisms in trophoblast invasion has focused mainly on intracellular, nonsecretory epithelial-mesenchymal transition (EMT) markers in conventional trophoblast cell lines, which limits its translational relevance to clinical applications. In this study, we performed small RNA sequencing combined with cell biology studies in primary human trophoblast and 2D human trophoblast stem cell models and revealed that the upregulation of the SOX4 (SRY-box transcription factor 4) and miR-103a-3p induced by activin A contributes to trophoblast invasion and potential extravillous differentiation. The bioinformatic analysis of proteomic and microRNA profiles from public databases revealed increased expression of the activin A protein and exosomal miR-103a-3p in maternal blood during the second trimester of pregnancy complicated by preeclampsia. Overall, our integrated approach reveals the regulatory mechanism by which activin A, SOX4, and miR-103a-3p regulate human trophoblast invasion and EVT differentiation, highlighting their potential as early diagnostic biomarkers for preeclampsia.
Similar content being viewed by others
Data availability
The data are available upon reasonable request. The raw small RNA sequencing data reported in this paper have been deposited in the Genome Sequence Archive for Human (GSA-Human) under accession No. HRA008074 and are available upon reasonable request.
References
Turco MY, Moffett A. Development of the human placenta. Development. 2019;146:dev163428.
Soares MJ, Varberg KM, Iqbal K. Hemochorial placentation: development, function, and adaptations. Biol Reprod. 2018;99:196–211.
Abbas Y, Turco MY, Burton GJ, Moffett A. Investigation of human trophoblast invasion in vitro. Hum Reprod Update. 2020;26:501–13.
Burton GJ, Woods AW, Jauniaux E, Kingdom JCP. Rheological and physiological consequences of conversion of the maternal spiral arteries for uteroplacental blood flow during human pregnancy. Placenta. 2009;30:473–482.
Ji L, Brkić J, Liu M, Fu G, Peng C, Wang YL. Placental trophoblast cell differentiation: physiological regulation and pathological relevance to preeclampsia. Mol Asp Med. 2013;34:981–1023.
Burton GJ, Redman CW, Roberts JM, Moffett A. Pre-eclampsia: pathophysiology and clinical implications. BMJ. 2019;366:l2381.
Ackerman WET, Buhimschi CS, Brown TL, Zhao G, Summerfield TL, Buhimschi IA. Transcriptomics-based subphenotyping of the human placenta enabled by weighted correlation network analysis in early-onset preeclampsia with and without fetal growth restriction. Hypertension. 2023;80:1363–74.
Xie L, Mouillet JF, Chu T, Parks WT, Sadovsky E, Knöfler M, et al. C19MC microRNAs regulate the migration of human trophoblasts. Endocrinology. 2014;155:4975–85.
Fu G, Ye G, Nadeem L, Ji L, Manchanda T, Wang Y, et al. MicroRNA-376c impairs transforming growth factor-β and nodal signaling to promote trophoblast cell proliferation and invasion. Hypertension. 2013;61:864–72.
Frazier S, McBride MW, Mulvana H, Graham D. From animal models to patients: the role of placental microRNAs, miR-210, miR-126, and miR-148a/152 in preeclampsia. Clin Sci. 2020;134:1001–25.
Luo R, Shao X, Xu P, Liu Y, Wang Y, Zhao Y, et al. MicroRNA-210 contributes to preeclampsia by downregulating potassium channel modulatory factor 1. Hypertension. 2014;64:839–45.
Torres-Torres J, Espino YSS, Martinez-Portilla R, Borboa-Olivares H, Estrada-Gutierrez G, Acevedo-Gallegos S. et al. A narrative review on the pathophysiology of preeclampsia. Int J Mol Sci. 2024;25:7569.
Kamrani A, Alipourfard I, Ahmadi-Khiavi H, Yousefi M, Rostamzadeh D, Izadi M, et al. The role of epigenetic changes in preeclampsia. Biofactors. 2019;45:712–24.
Bai Y, Yang W, Yang HX, Liao Q, Ye G, Fu G, et al. Downregulated miR-195 detected in preeclamptic placenta affects trophoblast cell invasion via modulating ActRIIA expression. PLoS ONE. 2012;7:e38875.
Jones RL, Findlay JK, Farnworth PG, Robertson DM, Wallace E, Salamonsen LA. Activin A and inhibin A differentially regulate human uterine matrix metalloproteinases: potential interactions during decidualization and trophoblast invasion. Endocrinology. 2006;147:724–32.
Li Y, Klausen C, Zhu H, Leung PC. Activin A increases human trophoblast invasion by inducing SNAIL-mediated MMP2 up-regulation through ALK4. J Clin Endocrinol Metab. 2015;100:E1415–27.
Muttukrishna S, Knight PG, Groome NP, Redman CW, Ledger WL. Activin A and inhibin A as possible endocrine markers for pre-eclampsia. Lancet. 1997;349:1285–8.
Barrero JA, Villamil-Camargo LM, Imaz JN, Arciniegas-Villa K, Rubio-Romero JA. Maternal serum activin A, inhibin A and follistatin-related proteins across preeclampsia: insights into their role in pathogenesis and prediction. J Mother Child. 2023;27:119–33.
Lee CQ, Gardner L, Turco M, Zhao N, Murray MJ, Coleman N, et al. What is trophoblast? A combination of criteria define human first-trimester trophoblast. Stem Cell Rep. 2016;6:257–72.
Irving JA, Lysiak JJ, Graham CH, Hearn S, Han VK, Lala PK. Characteristics of trophoblast cells migrating from first trimester chorionic villus explants and propagated in culture. Placenta. 1995;16:413–33.
Okae H, Toh H, Sato T, Hiura H, Takahashi S, Shirane K, et al. Derivation of human trophoblast stem cells. Cell Stem Cell. 2018;22:50–63.e6.
Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10:R25.
Wang L, Feng Z, Wang X, Wang X, Zhang X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics. 2010;26:136–8.
Rahmati S, Abovsky M, Pastrello C, Jurisica I. pathDIP: an annotated resource for known and predicted human gene-pathway associations and pathway enrichment analysis. Nucleic Acids Res. 2017;45:D419–26.
Chang L, Zhou G, Soufan O, Xia J. miRNet 2.0: network-based visual analytics for miRNA functional analysis and systems biology. Nucleic Acids Res. 2020;48:W244–51.
Xie J, Zhu H, Chang H-M, Klausen C, Dong M, Leung PCK. GDF8 promotes the cell invasiveness in human trophoblasts by upregulating the expression of follistatin-like 3 through the ALK5-SMAD2/3 signaling pathway. Front Cell Dev Biol. 2020;8:573781.
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, et al. The human genome browser at UCSC. Genome Res. 2002;12:996–1006.
Arancio W, Sciaraffa N, Coronnello C. MBS: a genome browser annotation track for high-confident microRNA binding sites in whole human transcriptome. Database. 2023;2023:baad015.
Vervoort SJ, de Jong OG, Roukens MG, Frederiks CL, Vermeulen JF, Lourenço AR. et al. Global transcriptional analysis identifies a novel role for SOX4 in tumor-induced angiogenesis. Elife. 2018;7:e27706.
Mehta GA, Angus SP, Khella CA, Tong K, Khanna P, Dixon SAH, et al. SOX4 and SMARCA4 cooperatively regulate PI3k signaling through transcriptional activation of TGFBR2. NPJ Breast Cancer. 2021;7:40.
Yoney A, Etoc F, Ruzo A, Carroll T, Metzger JJ, Martyn I, et al. WNT signaling memory is required for ACTIVIN to function as a morphogen in human gastruloids. eLife. 2018;7:e38279.
Varberg KM, Dominguez EM, Koseva B, Varberg JM, McNally RP, Moreno-Irusta A, et al. Extravillous trophoblast cell lineage development is associated with active remodeling of the chromatin landscape. Nat Commun. 2023;14:4826.
Vento-Tormo R, Efremova M, Botting RA, Turco MY, Vento-Tormo M, Meyer KB, et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature. 2018;563:347–53.
Shannon MJ, Baltayeva J, Castellana B, Wachter J, McNeill GL, Yoon JS. et al. Cell trajectory modeling identifies a primitive trophoblast state defined by BCAM enrichment. Development. 2022;149:dev199840.
Arutyunyan A, Roberts K, Troulé K, Wong FCK, Sheridan MA, Kats I, et al. Spatial multiomics map of trophoblast development in early pregnancy. Nature. 2023;616:143–51.
França GS, Vibranovski MD, Galante PA. Host gene constraints and genomic context impact the expression and evolution of human microRNAs. Nat Commun. 2016;7:11438.
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A. Identification of mammalian microRNA host genes and transcription units. Genome Res. 2004;14:1902–10.
Huang P, Deng W, Bao H, Lin Z, Liu MA-O, Wu J. et al. SOX4 facilitates PGR protein stability and FOXO1 expression conducive for human endometrial decidualization. Elife. 2022;11:e72073.
Morgan WD, Williams GT, Morimoto RI, Greene J, Kingston RE, Tjian R. Two transcriptional activators, CCAAT-box-binding transcription factor and heat shock transcription factor, interact with a human hsp7O gene promoter. Mol Cell Biol. 1987;7:1129–38.
Polster BJ, Yoon MY, Hayflick SJ. Characterization of the human PANK2 promoter. Gene. 2010;465:53–60.
Rabinovici J, Goldsmith PC, Librach CL, Jaffe RB. Localization and regulation of the activin-A dimer in human placental cells. J Clin Endocrinol Metab. 1992;75:571–6.
Liu Y, Fan X, Wang R, Lu X, Dang YL, Wang H, et al. Single-cell RNA-seq reveals the diversity of trophoblast subtypes and patterns of differentiation in the human placenta. Cell Res. 2018;28:819–32.
Jones RL, Salamonsen LA, Critchley HOD, Rogers PAW, Affandi B, Findlay JK. Inhibin and activin subunits are differentially expressed in endometrial cells and leukocytes during the menstrual cycle, in early pregnancy and in women using progestin-only contraception. Mol Hum Reprod. 2000;6:1107–17.
Barber CV, Yo JH, Rahman RA, Wallace EM, Palmer KR, Marshall SA. Activin A and pathologies of pregnancy: a review. Placenta. 2023;136:35–41.
Zhu S, Li Z, Cui L, Ban Y, Leung PCK, Li Y, et al. Activin A increases human trophoblast invasion by upregulating integrin β1 through ALK4. FASEB J. 2021;35:e21220.
Li Y, Klausen C, Cheng JC, Zhu H, Leung PC. Activin A, B, and AB increase human trophoblast cell invasion by up-regulating N-cadherin. J Clin Endocrinol Metab. 2014;99:E2216–25.
Abou-Kheir W, Barrak J, Hadadeh O, Daoud G. HTR-8/SVneo cell line contains a mixed population of cells. Placenta. 2017;50:1–7.
Martello G, Rosato A, Ferrari F, Manfrin A, Cordenonsi M, Dupont S, et al. A MicroRNA targeting dicer for metastasis control. Cell. 2010;141:1195–207.
Peng H, Park JK, Katsnelson J, Kaplan N, Yang W, Getsios S, et al. microRNA-103/107 family regulates multiple epithelial stem cell characteristics. Stem Cells. 2015;33:1642–56.
Moshkovsky AR, Kirschner MW. The nonredundant nature of the Axin2 regulatory network in the canonical Wnt signaling pathway. Proc Natl Acad Sci USA. 2022;119:e2108408119.
Karakis V, Jabeen M, Britt JW, Cordiner A, Mischler A, Li F, et al. Laminin switches terminal differentiation fate of human trophoblast stem cells under chemically defined culture conditions. J Biol Chem. 2023;299:104650.
Scharer CD, McCabe CD, Ali-Seyed M, Berger MF, Bulyk ML, Moreno CS. Genome-wide promoter analysis of the SOX4 transcriptional network in prostate cancer cells. Cancer Res. 2009;69:709–17.
Braun AE, Mitchel OR, Gonzalez TL, Sun T, Flowers AE, Pisarska MD, et al. Sex at the interface: the origin and impact of sex differences in the developing human placenta. Biol Sex Differ. 2022;13:50.
Buckberry S, Bianco-Miotto T, Bent SJ, Dekker GA, Roberts CT. Integrative transcriptome meta-analysis reveals widespread sex-biased gene expression at the human fetal-maternal interface. Mol Hum Reprod. 2014;20:810–9.
Tarca AL, Romero R, Benshalom-Tirosh N, Than NG, Gudicha DW, Done B, et al. The prediction of early preeclampsia: results from a longitudinal proteomics study. PLoS ONE. 2019;14:e0217273.
Salomon C, Guanzon D, Scholz-Romero K, Longo S, Correa P, Illanes SE, et al. Placental exosomes as early biomarker of preeclampsia: potential role of exosomal microRNAs across gestation. J Clin Endocrinol Metab. 2017;102:3182–94.
Acknowledgements
The authors extend their sincere gratitude to the hard work of the staff at British Columbia’s Women’s Hospital’s CARE Program for recruiting participants for our study. Additionally, we would like to thank Dr. Jingsong Wang at the Imaging Core Team of British Columbia’s Children’s Hospital Research Institute for helping with the microscopy imaging.
Funding
This work was supported by a Canadian Institutes of Health Research operating grant (PJT-183796 to PCKL and AGB), a Canadian Institutes of Health Research and a British Columbia Children’s Hospital graduate studentship (to MJS), and a graduate studentship of the Chinese Scholarship Council (to JMX).
Author information
Authors and Affiliations
Contributions
Conceptualization: JMX, MYD, and PCKL; Methodology: JMX, MYD, HZ, MJS, AGB, and PCKL; Formal analysis: JMX, MJS, and MYD; Investigation: JMX and HZ; Resources: PCKL and MYD; Writing—original draft: JMX; Writing—review and editing: HZ, MJS, AGB, MYD, and PCKL; Supervision: PCKL; Project administration: PCKL; Funding acquisition: PCKL and AGB.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Mauro Piacentini
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xie, J., Shannon, M.J., Zhu, H. et al. Elucidating the roles of microRNA-103a-3p in trophoblast invasion and SOX4-mediated extravillous differentiation induced by activin A. Cell Death Dis (2026). https://doi.org/10.1038/s41419-026-08665-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-026-08665-6