Abstract
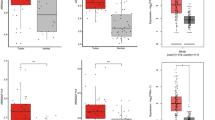
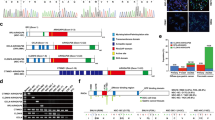
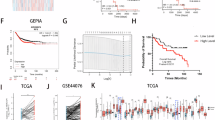
Rho GTPase-activating protein 21 (ARHGAP21) plays a role in the occurrence and development of certain cancers, but its function in hepatocellular carcinoma (HCC) remains unclear. In this study, elevated ARHGAP21 expression was observed in both HCC cell lines and tissues and correlated with poor patient prognosis. Knockdown of ARHGAP21 suppressed HCC cell migration and invasion in vitro by regulating the actin cytoskeleton, while overexpression of ARHGAP21 had the opposite effect. In vivo, knockdown of ARHGAP21 inhibited HCC tumorigenesis and metastasis. Mechanistically, we demonstrated that ARHGAP21 directly binds to FLNA, and the PDZ domain of ARHGAP21 functions as a potential mediator of its binding to the 1–1200 aa fragment of FLNA. ARHGAP21 also directly binds to and recruits HSP90α to stabilize FLNA by inhibiting its ubiquitination and degradation. Overexpression of FLNA reversed the cytoskeleton remodeling-related suppression of tumor metastasis in ARHGAP21-knockdown HCC cells. These results revealed that ARHGAP21 promotes cytoskeleton remodeling and stimulates HCC metastasis by inhibiting FLNA ubiquitination and degradation via HSP90α recruitment. Our results position ARHGAP21 as both a potential prognostic marker and a promising therapeutic target in HCC.
Similar content being viewed by others
Data availability
All relevant data are available from the authors.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer J Clin. 2021;71:209–49.
Chang M-H, You S-L, Chen C-J, Liu C-J, Lee C-M, Lin S-M, et al. Decreased incidence of hepatocellular carcinoma in hepatitis B vaccinees: a 20-year follow-up study. J Natl Cancer Inst. 2009;101:1348–55.
Llovet JM, Montal R, Sia D, Finn RS. Molecular therapies and precision medicine for hepatocellular carcinoma. Nat Rev Clin Oncol. 2018;15:599–616.
Kudo M, Finn RS, Qin S, Han KH, Ikeda K, Piscaglia F, et al. Lenvatinib versus sorafenib in first-line treatment of patients with unresectable hepatocellular carcinoma: a randomised phase 3 non-inferiority trial. Lancet. 2018;391:1163–73.
Foerster F, Gairing SJ, Ilyas SI, Galle PR. Emerging immunotherapy for HCC: a guide for hepatologists. Hepatology. 2022;75:1604–26.
Reig M, Forner A, Rimola J, Ferrer-Fábrega J, Burrel M, Garcia-Criado Á, et al. BCLC strategy for prognosis prediction and treatment recommendation: the 2022 update. J Hepatol. 2022;76:681–93.
Rhee H, Park YN, Choi JY. Advances in understanding hepatocellular carcinoma vasculature: implications for diagnosis, prognostication, and treatment. Korean J Radiol. 2024;25:887–901.
Sj H, Aj R. Mammalian rho GTPases: new insights into their functions from in vivo studies. Nat Rev Mol Cell Biol. 2008;9:690–701.
Smithers CC, Overduin M. Structural mechanisms and drug discovery prospects of rho GTPases. Cells. 2016;5:26
Schaefer A, Der CJ. RHOA takes the RHOad less traveled to cancer. Trends Cancer. 2022;8:655–69.
Morstein J, Bowcut V, Fernando M, Yang Y, Zhu L, Jenkins ML, et al. Targeting Ras-, Rho-, and Rab-family GTPases via a conserved cryptic pocket. Cell. 2024;187:6379–92.e17.
Rosa LRO, Soares GM, Silveira LR, Boschero AC, Barbosa-Sampaio HCL. ARHGAP21 as a master regulator of multiple cellular processes. J Cell Physiol. 2018;233:8477–81.
Bigarella CL, Borges L, Costa FF, Saad ST. ARHGAP21 modulates FAK activity and impairs glioblastoma cell migration. Biochim Biophys Acta. 2009;1793:806–16.
Ménétrey J, Perderiset M, Cicolari J, Dubois T, Elkhatib N, El Khadali F, et al. Structural basis for ARF1-mediated recruitment of ARHGAP21 to Golgi membranes. EMBO J. 2007;26:1953–62.
Lazarini M, Traina F, Machado-Neto JA, Barcellos KS, Moreira YB, Brandão MM, et al. ARHGAP21 is a RhoGAP for RhoA and RhoC with a role in proliferation and migration of prostate adenocarcinoma cells. Biochim Biophys Acta. 2013;1832:365–74.
Lazarini M, Assis-Mendonça GR, Machado-Neto JA, Latuf-Filho P, Bezerra SM, Vieira KP, et al. Silencing of ARHGAP21, a Rho GTPase activating protein (RhoGAP), reduces the growth of prostate cancer xenografts in NOD/SCID mice. Biochim Biophys Acta Mol Cell Res. 2023;1870:119439.
Wang Z, Wu S, Wang G, Yang Z, Zhang Y, Zhu C. et al. ARHGAP21 is involved in the carcinogenic mechanism of cholangiocarcinoma: a study based on bioinformatic analyses and experimental validation. Medicina. 2023;59:139
Mylvaganam S, Freeman SA, Grinstein S. The cytoskeleton in phagocytosis and macropinocytosis. Curr Biol. 2021;31:R619–r32.
Kouchi Z, Kojima M. A Structural network analysis of neuronal ArhGAP21/23 interactors by computational modeling. ACS Omega. 2023;8:19249–64.
Lüchow S, Sundell GN, Ivarsson Y. Identification of PDZ interactions by proteomic peptide phage display. Methods Mol Biol. 2021;2256:41–60.
Healy MD, Collins BM. The PDLIM family of actin-associated proteins and their emerging role in membrane trafficking. Biochem Soc Trans. 2023;51:2005–16.
Sui Y, Shen Z, Pan R, Ma R, Si R, Feng J, et al. AHSA1-HSP90AA1 complex stabilized IFI6 and TGFB1 promotes mitochondrial stability and EMT in EGFR-mutated lung adenocarcinoma under Osimertinib pressure. Cell Death Dis. 2025;16:298.
Wan S, Liu C, Li C, Wang Z, Zhao G, Li J, et al. AKIP1 accelerates glioblastoma progression through stabilizing EGFR expression. Oncogene. 2023;42:2905–18.
Ahn JC, Teng PC, Chen PJ, Posadas E, Tseng HR, Lu SC, et al. Detection of circulating tumor cells and their implications as a biomarker for diagnosis, prognostication, and therapeutic monitoring in hepatocellular carcinoma. Hepatology. 2021;73:422–36.
Carles A, Millon R, Cromer A, Ganguli G, Lemaire F, Young J, et al. Head and neck squamous cell carcinoma transcriptome analysis by comprehensive validated differential display. Oncogene. 2006;25:1821–31.
Barcellos KSA, Bigarella CL, Wagner MV, Vieira KP, Lazarini M, Langford PR, et al. ARHGAP21 protein, a new partner of α-tubulin involved in cell-cell adhesion formation and essential for epithelial-mesenchymal transition. J Biol Chem. 2013;288:2179–89.
Novosedlik S, Reichel F, van Veldhuisen T, Li Y, Wu H, Janssen H, et al. Cytoskeleton-functionalized synthetic cells with life-like mechanical features and regulated membrane dynamicity. Nat Chem. 2025;17:356–64.
Park J, Wu Y, Suk Kim J, Byun J, Lee J, Oh YK. Cytoskeleton-modulating nanomaterials and their therapeutic potentials. Adv Drug Deliv Rev. 2024;211:115362.
Iwamoto DV, Huehn A, Simon B, Huet-Calderwood C, Baldassarre M, Sindelar CV, et al. Structural basis of the filamin A actin-binding domain interaction with F-actin. Nat Struct Mol Biol. 2018;25:918–27.
Robertson SP. Filamin A: phenotypic diversity. Curr Opin Genet Dev. 2005;15:301–7.
Treppiedi D, Catalano R, Mangili F, Mantovani G, Peverelli E. Role of filamin A in the pathogenesis of neuroendocrine tumors and adrenal cancer. Endocr Oncol. 2022;2:R143–r52.
Shead KD, Salyahetdinova V, Baillie GS. Charting the importance of filamin A posttranslational modifications. Biochem J. 2024;481:865–81.
Chiang TS, Wu HF, Lee FS. ADP-ribosylation factor-like 4C binding to filamin-A modulates filopodium formation and cell migration. Mol Biol Cell. 2017;28:3013–28.
Li C, Yu S, Nakamura F, Yin S, Xu J, Petrolla AA, et al. Binding of pro-prion to filamin A disrupts cytoskeleton and correlates with poor prognosis in pancreatic cancer. J Clin Invest. 2009;119:2725–36.
Donadon M, Di Tommaso L, Soldani C, Franceschini B, Terrone A, Mimmo A, et al. Filamin A expression predicts early recurrence of hepatocellular carcinoma after hepatectomy. Liver Int. 2018;38:303–11.
Maki T, Kidoaki S, Usui K, Suzuki H, Ito M, Ito F, et al. Dynamic force spectroscopy of the specific interaction between the PDZ domain and its recognition peptides. Langmuir. 2007;23:2668–73.
Zhang Y, Zhao G, Yu L, Wang X, Meng Y, Mao J, et al. Heat-shock protein 90α protects NME1 against degradation and suppresses metastasis of breast cancer. Br J Cancer. 2023;129:1679–91.
Nakasone MA, Majorek KA, Gabrielsen M, Sibbet GJ, Smith BO, Huang DT. Structure of UBE2K-Ub/E3/polyUb reveals mechanisms of K48-linked Ub chain extension. Nat Chem Biol. 2022;18:422–31.
Ohtake F, Tsuchiya H, Saeki Y, Tanaka K. K63 ubiquitylation triggers proteasomal degradation by seeding branched ubiquitin chains. Proc Natl Acad Sci USA. 2018;115:E1401–e8.
Aalto A, Martínez-Chacón G, Kietz C, Tsyganova N, Kreutzer J, Kallio P, et al. M1-linked ubiquitination facilitates NF-κB activation and survival during sterile inflammation. FEBS J. 2022;289:5180–97.
Zhou Q, Zhang J. K27-linked noncanonic ubiquitination in immune regulation. J Leukoc Biol. 2022;111:223–35.
Michel MA, Elliott PR, Swatek KN, Simicek M, Pruneda JN, Wagstaff JL, et al. Assembly and specific recognition of k29- and k33-linked polyubiquitin. Mol Cell. 2015;58:95–109.
Funding
The work was supported by the National Natural Science Foundation of China (Grant No.81872251); the Natural Science Foundation of Guangdong Province (Grant Nos. 2024A1515013279, 2021A1515012104, 2020A1515010093); China Foundation for Youth Entrepreneurship and Employment; Project of Guangdong Medical Science and Technology Research Foundation (A2025087).
Author information
Authors and Affiliations
Contributions
HY and ZX designed the study, completed the experiment, and collated the data. HY produced the initial draft of the manuscript and contributed to modifying the manuscript. FL, JL, and FC designed the study, provided research funds, and were responsible for the revision of the entire manuscript. XT, XZ, XW, KX, ZZ, and YZ participated in the course of the experiment. All authors have read and approved the final submitted manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
All animal studies were approved by Shenzhen Top Biotech Co., Ltd., Institutional Animal Care and Use Committee (ethical approval code: TOP-IACUC-2022-0230), and all methods were performed in accordance with the relevant guidelines and regulations.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yao, H., Xie, Z., Tao, X. et al. ARHGAP21 enhances metastasis in hepatocellular carcinoma by inhibiting ubiquitination of filamin A. Cell Death Discov. (2026). https://doi.org/10.1038/s41420-026-03103-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-026-03103-0