Abstract
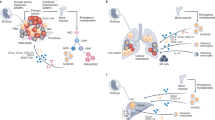
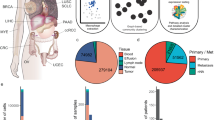
Tumor-associated macrophages (TAMs) play crucial roles in tumor progression. However, the mechanisms underlying the posttranscriptional regulation of TAMs remain largely unknown. Here, we demonstrated that Trmt61a, the "writer" enzyme of tRNA N1-methyladenosine (m1A) modification, is highly expressed in proinflammatory macrophages in tumor microenvironment. We generated conditional knockout (KO) mice for Trmt61a and observed that Trmt61a deletion in macrophages significantly promoted tumor growth. Mechanistically, we identified that m1A maintains the translation of STING, enhances STING-TBK1-IFN-β signaling in macrophages and therefore suppresses tumor cell growth. We further generated TRMT61A-overexpressing human iPSC-derived CAR-macrophage and demonstrated that human TRMT61A effectively promoted antitumor CAR-macrophage therapy in vivo. Collectively, our findings reveal a novel regulatory mechanism of tRNA m1A modification in macrophages, highlighting the antitumor therapeutic potential of targeting tRNA m1A modification in macrophages.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kloosterman DJ, Akkari L. Macrophages at the interface of the coevolving cancer ecosystem. Cell. 2023;186:1627–51.
Cassetta L, Pollard JW. Targeting macrophages: therapeutic approaches in cancer. Nat Rev Drug Discov. 2018;17:887–904.
Pathria P, Louis TL, Varner JA. Targeting Tumor-Associated Macrophages in Cancer. Trends Immunol. 2019;40:310–27.
Snell LM, McGaha TL, Brooks DG. Type I Interferon in Chronic Virus Infection and Cancer. Trends Immunol. 2017;38:542–57.
Yu R, Zhu B, Chen D. Type I interferon-mediated tumor immunity and its role in immunotherapy. Cell Mol Life Sci. 2022;79:191.
von Locquenghien M., Rozalén C, Celià-Terrassa T. Interferons in cancer immunoediting: sculpting metastasis and immunotherapy response. J Clin Investig. 131, (2021)
Woo SR, Fuertes MB, Corrales L, Spranger S, Furdyna MJ, Leung MY, et al. STING-dependent cytosolic DNA sensing mediates innate immune recognition of immunogenic tumors. Immunity. 2014;41:830–42.
Andzinski L, Spanier J, Kasnitz N, Kröger A, Jin L, Brinkmann MM, et al. Growing tumors induce a local STING dependent Type I IFN response in dendritic cells. Int J Cancer. 2016;139:1350–7.
Dégut C, Ponchon L, Folly-Klan M, Barraud P, Tisné C. The m1A(58) modification in eubacterial tRNA: An overview of tRNA recognition and mechanism of catalysis by TrmI. Biophys Chem. 2016;210:27–34.
Oerum S., Dégut C, Barraud P., Tisné C. m1A Post-Transcriptional Modification in tRNAs. Biomolecules. 7, (2017)
Barraud P, Golinelli-Pimpaneau B, Atmanene C, Sanglier S, Van Dorsselaer A, Droogmans L, et al. Crystal structure of Thermus thermophilus tRNA m1A58 methyltransferase and biophysical characterization of its interaction with tRNA. J Mol Biol. 2008;377:535–50.
Liu F, Clark W, Luo G, Wang X, Fu Y, Wei J, et al. ALKBH1-Mediated tRNA Demethylation Regulates Translation. Cell. 2016;167:816–28.e816.
Zhang C, Jia G. Reversible RNA Modification N(1)-methyladenosine (m(1)A) in mRNA and tRNA. Genomics Proteomics Bioinformatics. 2018;16:155–61.
Chen Y, Oh MH, Flavell R, Li HB. RNA methylation in immune cells. Adv Immunol. 2022;155:39–94.
Liu Y, Zhou J, Li X, Zhang X, Shi J, Wang X, et al. tRNA-m(1)A modification promotes T-cell expansion via efficient MYC protein synthesis. Nat Immunol. 2022;23:1433–44.
Miao S., Li H, Song X., Liu Y., Wang G., Kan C., et al. tRNA m1A modification regulates cholesterol biosynthesis to promote antitumor immunity of CD8+ T cells. J. Exp. Med. 222, (2025)
Zhao Y, Zhao Q, Kaboli PJ, Shen J, Li M, Wu X, et al. m1A Regulated Genes Modulate PI3K/AKT/mTOR and ErbB Pathways in Gastrointestinal Cancer. Transl Oncol. 2019;12:1323–33.
Li Y, Zheng D, Wang F, Xu Y, Yu H, Zhang H. Expression of Demethylase Genes, FTO and ALKBH1, Is Associated with Prognosis of Gastric Cancer. Dig Dis Sci. 2019;64:1503–13.
Yamato I, Sho M, Shimada K, Hotta K, Ueda Y, Yasuda S, et al. PCA-1/ALKBH3 contributes to pancreatic cancer by supporting apoptotic resistance and angiogenesis. Cancer Res. 2012;72:4829–39.
Woo HH, Chambers SK. Human ALKBH3-induced m(1)A demethylation increases the CSF-1 mRNA stability in breast and ovarian cancer cells. Biochim Biophys Acta Gene Regul Mech. 2019;1862:35–46.
Tasaki M, Shimada K, Kimura H, Tsujikawa K, Konishi N. ALKBH3, a human AlkB homolog, contributes to cell survival in human non-small cell lung cancer. Br J Cancer. 2011;104:700–6.
Shi L, Yang XM, Tang DD, Liu G, Yuan P, Yang Y, et al. Expression and significance of m1A transmethylase, hTrm6p/hTrm61p and its related gene hTrm6/hTrm61 in bladder urothelial carcinoma. Am J Cancer Res. 2015;5:2169–79.
Wang Y, Wang J, Li X, Xiong X, Wang J, Zhou Z, et al. N(1)-methyladenosine methylation in tRNA drives liver tumorigenesis by regulating cholesterol metabolism. Nat Commun. 2021;12:6314.
Tao EW, Wang Y, Tan J, Chen Y, Sun TY, Hao Y, et al. TRMT6-mediated tRNA m(1)A modification acts as a translational checkpoint of histone synthesis and facilitates colorectal cancer progression. Nat Cancer. 2025;6:1458–76.
Chen H, Yao J, Bao R, Dong Y, Zhang T, Du Y, et al. Cross-talk of four types of RNA modification writers defines tumor microenvironment and pharmacogenomic landscape in colorectal cancer. Mol Cancer. 2021;20:29.
Dong L, Chen C, Zhang Y, Guo P, Wang Z, Li J, et al. The loss of RNA N(6)-adenosine methyltransferase Mettl14 in tumor-associated macrophages promotes CD8(+) T cell dysfunction and tumor growth. Cancer Cell. 2021;39:945–57.e910.
Wang Z, Fang Y, Yu Y, Pan H. ALKBH1-mediated N6-methyladenosine methylation of TRAF1 promotes osteosarcoma proliferation and metastasis. Am J Cancer Res. 2025;15:375–89.
Zhang M, Yang S, Nelakanti R, Zhao W, Liu G, Li Z, et al. Mammalian ALKBH1 serves as an N(6)-mA demethylase of unpairing DNA. Cell Res. 2020;30:197–210.
Xiao CL, Zhu S, He M, Chen D, Zhang Q, Chen Y, et al. N(6)-Methyladenine DNA Modification in the Human Genome. Mol Cell. 2018;71:306–18.e307.
Wollen KL, Hagen L, Vågbø CB, Rabe R, Iveland TS, Aas PA, et al. ALKBH3 partner ASCC3 mediates P-body formation and selective clearance of MMS-induced 1-methyladenosine and 3-methylcytosine from mRNA. J Transl Med. 2021;19:287.
Chen Z, Qi M, Shen B, Luo G, Wu Y, Li J, et al. Transfer RNA demethylase ALKBH3 promotes cancer progression via induction of tRNA-derived small RNAs. Nucleic Acids Res. 2019;47:2533–45.
Zuo H, Wu A, Wang M, Hong L, Wang H. tRNA m(1)A modification regulate HSC maintenance and self-renewal via mTORC1 signaling. Nature commun. 2024;15:5706.
Lei A, Yu H, Lu S, Lu H, Ding X, Tan T, et al. A second-generation M1-polarized CAR macrophage with antitumor efficacy. Nat Immunol. 2023;25:102–16.
Zhang L, Tian L, Dai X, Yu H, Wang J, Lei A, et al. Pluripotent stem cell-derived CAR-macrophages with antigen-dependent anticancer cell functions. J Hematol Oncol. 2020;13:153.
Liu Y, You Y, Lu Z, Yang J, Li P, Liu L, et al. N (6)-methyladenosine RNA modification-mediated cellular metabolism rewiring inhibits viral replication. Science. 2019;365:1171–6.
Du J, Liao W, Liu W, Deb DK, He L, Hsu PJ, et al. N(6)-Adenosine Methylation of Socs1 mRNA Is Required to Sustain the Negative Feedback Control of Macrophage Activation. Dev Cell. 2020;55:737–53.e737.
Qiu W, Zhang Q, Zhang R, Lu Y, Wang X, Tian H, et al. N(6)-methyladenosine RNA modification suppresses antiviral innate sensing pathways by reshaping double-stranded RNA. Nat Commun. 2021;12:1582.
Qin Y, Li B, Arumugam S, Lu Q, Mankash SM, Li J, et al. m(6)A mRNA methylation-directed myeloid cell activation controls progression of NAFLD and obesity. Cell Rep. 2021;37:109968.
Xiong J, He J, Zhu J, Pan J, Liao W, Ye H, et al. Lactylation-driven METTL3-mediated RNA m(6)A modification promotes immunosuppression of tumor-infiltrating myeloid cells. Mol Cell. 2022;82:1660–77.e1610.
Tong J., Wang X, Liu Y., Ren X., Wang A., Chen Z., et al. Pooled CRISPR screening identifies m(6)A as a positive regulator of macrophage activation. Sci. Adv. 7, (2021).
McNab F, Mayer-Barber K, Sher A, Wack A, O'Garra A. Type I interferons in infectious disease. Nat Rev Immunol. 2015;15:87–103.
Gan Y, Li X, Han S, Liang Q, Ma X, Rong P, et al. The cGAS/STING Pathway: A Novel Target for Cancer Therapy. Front Immunol. 2021;12:795401.
Shen Z, Wei L, Yu Z, Yao Z, Cheng J, Wang Y, et al. The Roles of TRIMs in Antiviral Innate Immune Signaling. Front Cell Infect Microbiol. 2021;11:628275.
Chen Y, Yu Z, Tan X, Jiang H, Xu Z, Fang Y, et al. CAR-macrophage: A new immunotherapy candidate against solid tumors. Biomed Pharmacother. 2021;139:111605.
Pan K, Farrukh H, Chittepu V, Xu H, Pan CX, Zhu Z. CAR race to cancer immunotherapy: from CAR T, CAR NK to CAR macrophage therapy. J Exp Clin Cancer Res. 2022;41:119.
Behrens A, Rodschinka G, Nedialkova DD. High-resolution quantitative profiling of tRNA abundance and modification status in eukaryotes by mim-tRNAseq. Mol Cell. 2021;81:1802–15.e1807.
Acknowledgements
We thank K. Mao, M. Yang, and J. Zhao for their technical and administrative assistance and all other members of the Hua-Bing Li laboratory for their discussions and comments. This work was supported by the National Natural Science Foundation of China (82341017/82541015/82441048/82461160323/82325024/82350112 to H.-B.L., 32400747 to X. W.), the National Key R&D Plan of China (2025YFC3410100 to H-B.L.), the Ministry of Science and Technology of China (2021YFA1100800 to H-B.L.), the Chongqing Municipal Science and Technology Bureau (CSTB2025TIAD-STX0043/CSTB2025TIAD-STX0045), the Science and Technology Research Program of Chongqing Municipal Education Commission (Grant No. KJZD-M202500402), the Chongqing Municipal Finance Bureau and Chongqing Municipal Education Commission (010405000001), the Chongqing Science and Technology Bureau (CSTB2025YCJH-KYXM0002). We thank Ning Xu and Yue Yin of the mass spectrometry system at the National Facility for Protein Science in Shanghai (NFPS), Shanghai Advanced Research Institute, Chinese Academy of Science, China, for MS sample preparation, data collection and data analysis.
Author information
Authors and Affiliations
Contributions
H-B.L. and X.W. conceived the project, designed the research and wrote the manuscript. X.W. designed and performed the experiments, analyzed and interpreted the data, and drafted the manuscript. X.W. and S.S. performed the CAR-iMAC experiments. H.L. performed and analyzed the mass spectrometry data and tRNA-seq data. S.L. and Y.L. performed part of the tumor model and Western blotting experiments. H.C. and Y.Y. helped with the analysis of the scRNA-seq, bulk RNA-seq and ChIP-seq data. B.L., R.L., W.H., X.Z. and Y.Y. contributed to the discussion and suggestions. This study was supervised by H-B.L. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing interests. B.L. is an editorial board member of Cellular & Molecular Immunology, but he has not been involved in the peer review or the decision-making of the article.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, X., Wang, X., Li, H. et al. tRNA m1A modification orchestrates STING translation in macrophages to enhance antitumor immunity and CAR-macrophage immunotherapy. Cell Mol Immunol 23, 261–272 (2026). https://doi.org/10.1038/s41423-025-01383-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41423-025-01383-7